Search Count: 3,832
All
Selected
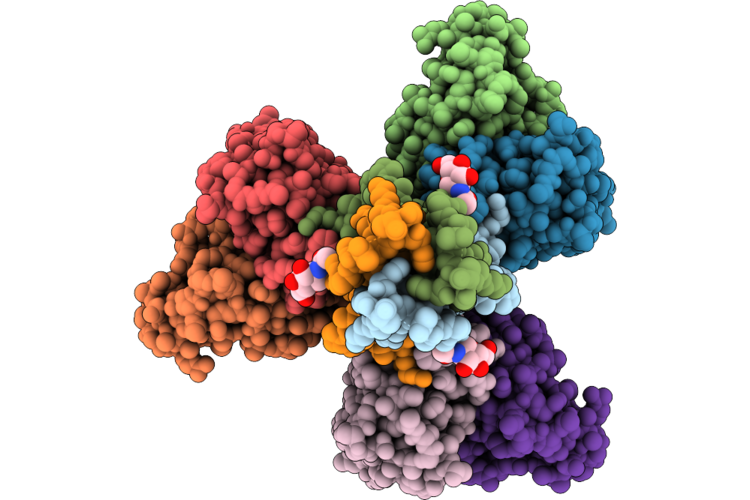 |
Sars-Cov-2 Spike S2 Trimer Stabilized In The Early Fusion Intermediate Conformation (E-Fics-V3) Bound To The Vn01H1 Fab (Fab Local Refinement)
Organism: Saccharomyces cerevisiae s288c, Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
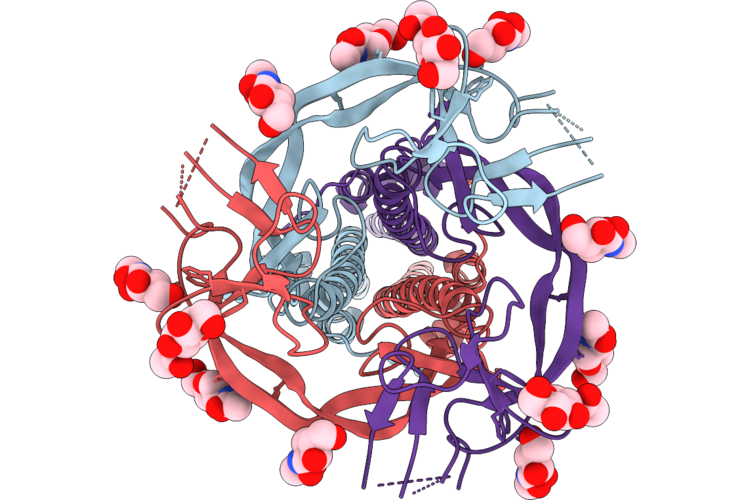 |
Sars-Cov-2 Spike S2 Trimer Stabilized In The Early Fusion Intermediate Conformation (E-Fics-V3) Bound To The Vn01H1 Fab (S2 Local Refinement)
Organism: Saccharomyces cerevisiae s288c, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
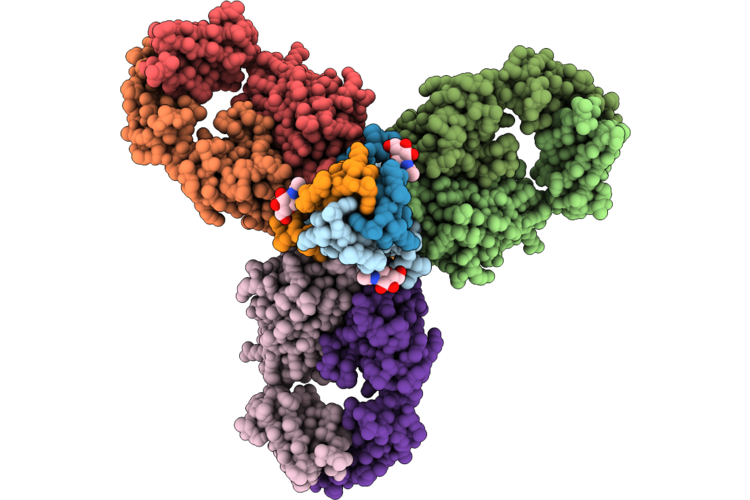 |
Sars-Cov-2 Spike S2 Trimer Stabilized In The Early Fusion Intermediate Conformation (E-Fics-V3) Bound To C77G12 (Fab Local Refinement)
Organism: Saccharomyces cerevisiae s288c, Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
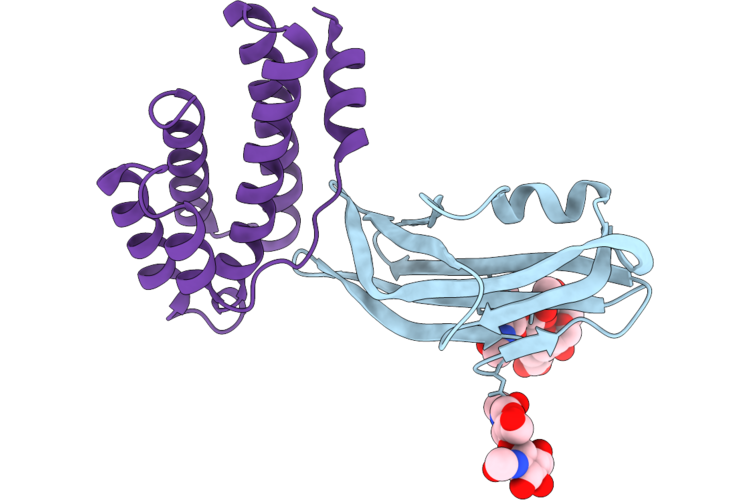 |
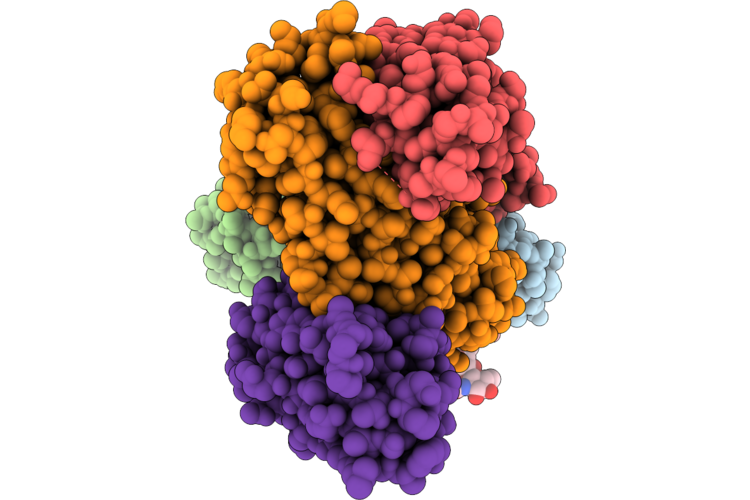
Structure Of The Porcine Deltacoronavirus (Pdcov) Receptor-Binding Domain Bound To The Rbd Minibinder 11, The Pd3 Fab, And The Kappa Light Chain Nanobody (Local Refinement)
Organism: Porcine deltacoronavirus, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-13 Classification: VIRAL PROTEIN |
 |
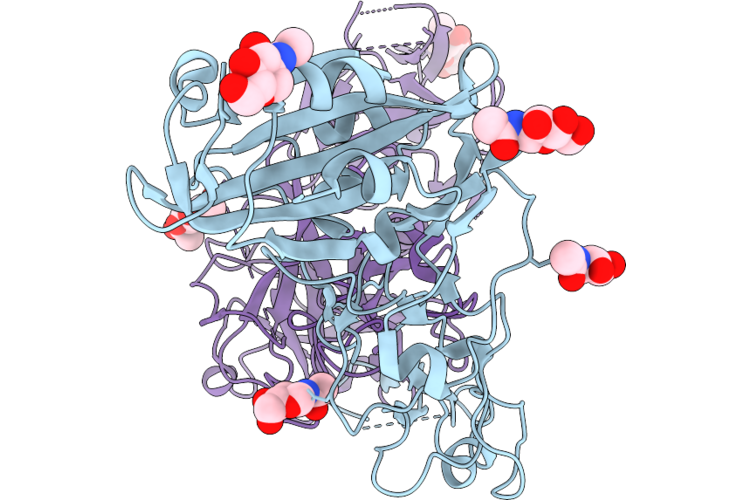
Structure Of The Porcine Deltacoronavirus (Pdcov) Receptor-Binding Domain Bound To The Rbd Minibinder 11, The Pd3 Fab, And The Kappa Light Chain Nanobody
Organism: Porcine deltacoronavirus, Synthetic construct, Mus musculus, Lama glama
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-05-13 Classification: VIRAL PROTEIN |
 |
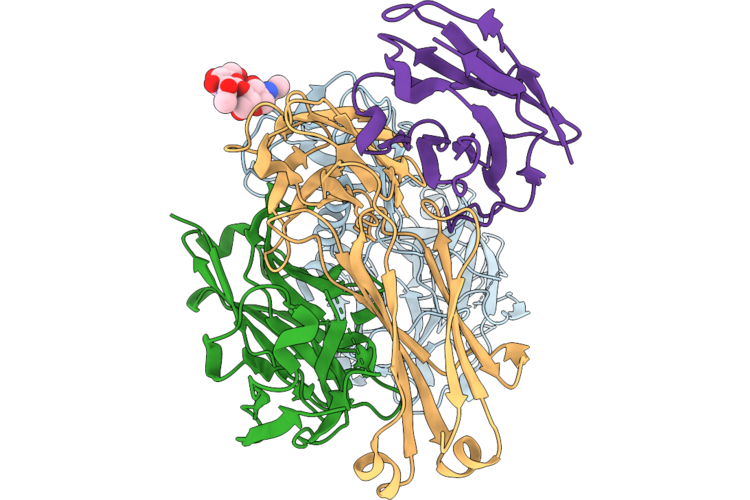
Tmprss2 (S441A) Bound To The Hcov-Nl63 S2'Region Genetically Fused To The Hcov-Hku1 Rbd
Organism: Human coronavirus hku1, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-05-13 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG |
 |
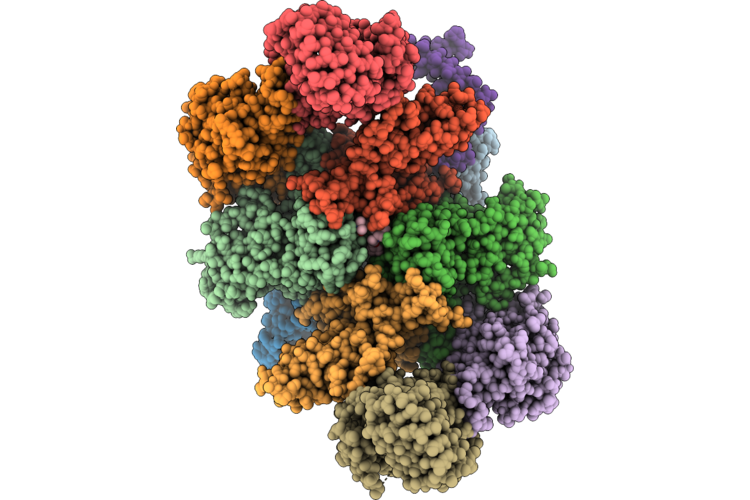
Tmprss2 S441A In Complex With The H1H7 Fab And Anti-Kappa Light Chain Nanobody
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-05-13 Classification: HYDROLASE/IMMUNE SYSTEM |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.83 Å Release Date: 2026-05-13 Classification: TRANSLATION Ligands: A1I5O |
 |
Organism: Homo sapiens
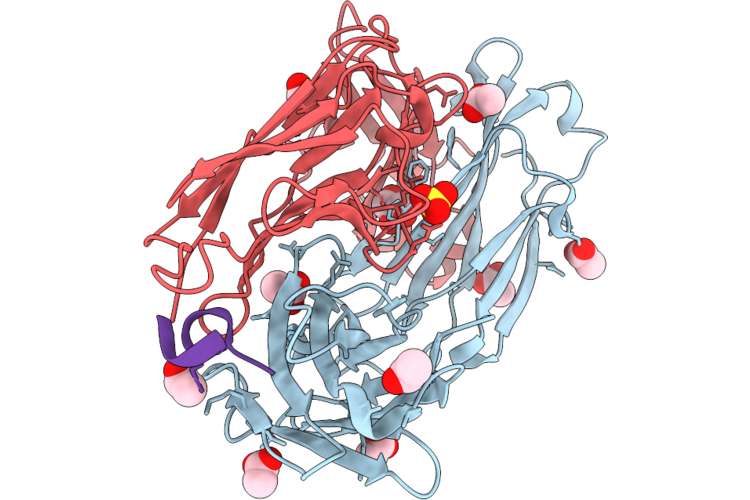
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-05-13 Classification: IMMUNE SYSTEM Ligands: GOL, ACT, SO4 |
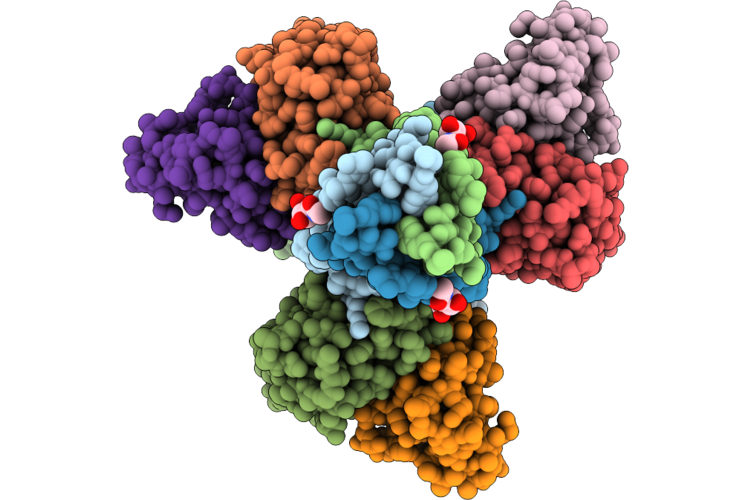 |
Sars-Cov-2 Spike Trimer In The Early Fusion Intermediate Conformation Bound To The Vn01H1 Fab (Fab Local Refinement)
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
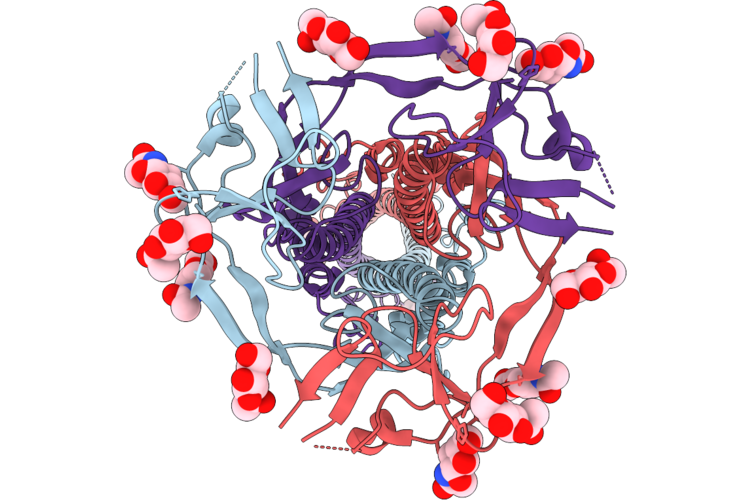 |
Sars-Cov-2 Spike Trimer In The Early Fusion Intermediate Conformation Bound To The Vn01H1 Fab (S2 Local Refinement)
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
 |
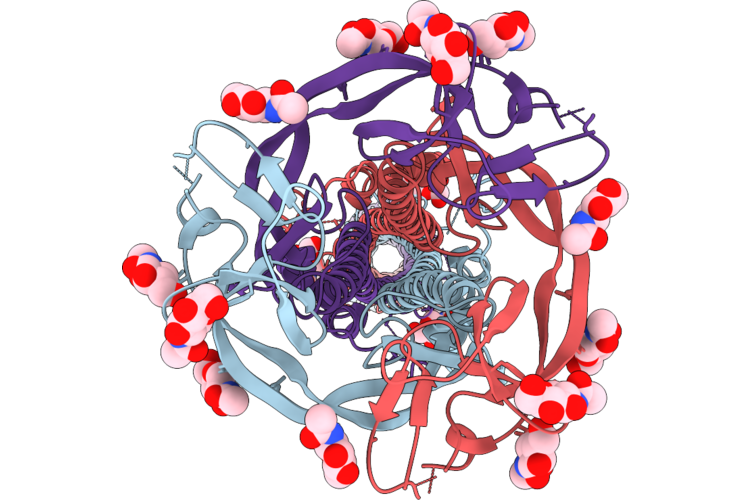
Hcov-Nl63 S2' Peptide Bound To Tmprss2 S441A (Complexed With The H1H7 Fab And An Anti-Kappa-Nanobody)
Organism: Human coronavirus nl63, Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-05-13 Classification: VIRAL PROTEIN/Immune System |
 |
Sars-Cov-2 S2 Trimer Stabilized In The Early Fusion Intermediate Conformation By Circular Permutation And Clamping By Gp41 (E-Fics-V1)
Organism: Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
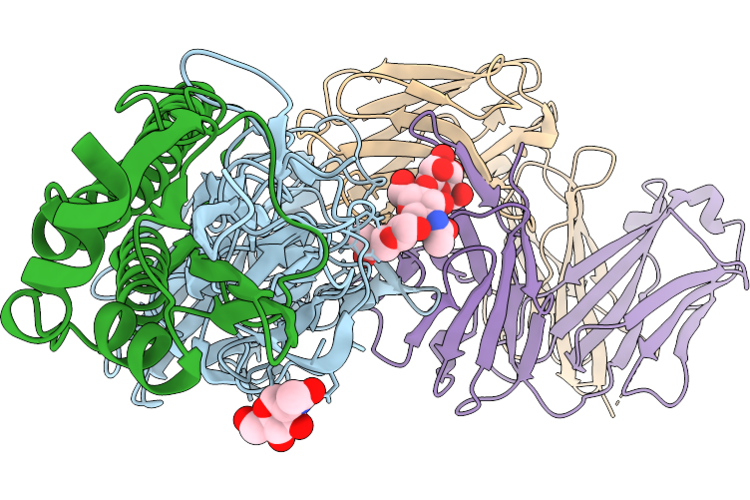 |
Crystal Structure Of Hemagglutinin From H1N1 Influenza A Virus A/California/04/2009 Bound To The 3_H2 Antibody
Organism: Influenza a virus (a/california/04/2009(h1n1)), Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.39 Å Release Date: 2026-05-06 Classification: ANTIVIRAL PROTEIN Ligands: NAG |
 |
Crystal Structure Of Hemagglutinin From H1N1 Influenza A Virus A/California/04/2009 Bound To The 49_C09 Antibody
Organism: Influenza a virus (a/california/04/2009(h1n1)), Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.80 Å Release Date: 2026-05-06 Classification: ANTIVIRAL PROTEIN Ligands: NAG |
 |
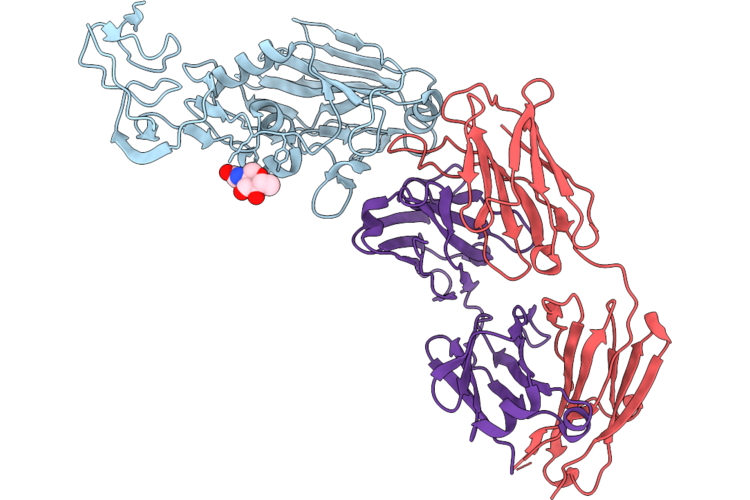
Crystal Structure Of Hemagglutinin Head Domain From H3N2 Influenza A Virus A/New York/631/1996 Bound To The 3_H2 Antibody
Organism: Influenza a virus (a/new york/631/1996(h3n2)), Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-05-06 Classification: ANTIVIRAL PROTEIN Ligands: NAG, PO4 |
 |
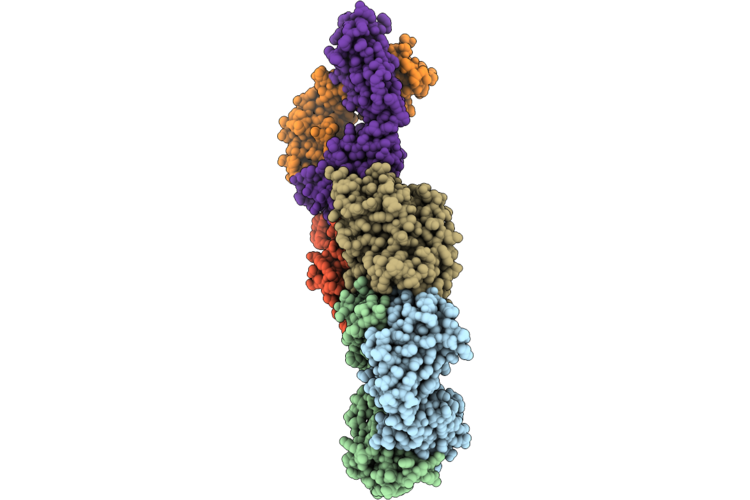
Crystal Structure Of Hemagglutinin Head Domain From H3N2 Influenza A Virus A/New York/631/1996 Bound To The 49_C09 Antibody
Organism: Influenza a virus (a/new york/631/1996(h3n2)), Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.71 Å Release Date: 2026-05-06 Classification: ANTIVIRAL PROTEIN Ligands: NAG |
 |
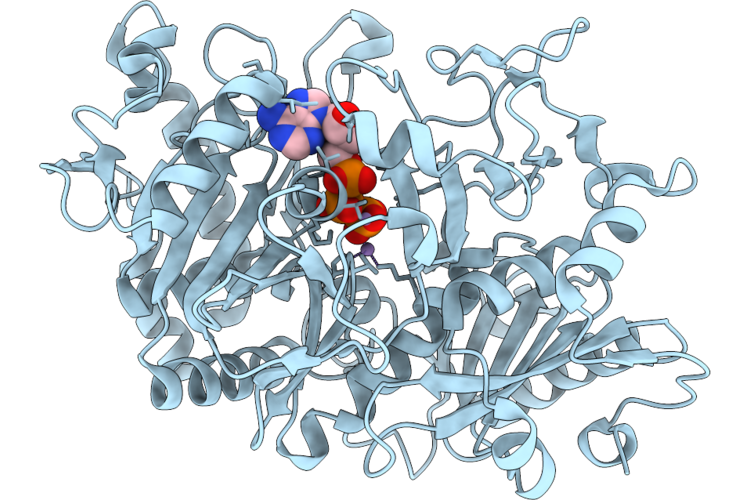
Crystal Structure Of Hemagglutinin Head Domain From H1N1 Influenza A Virus A/Victoria/2570/2019 Bound To The 49_C09 Antibody
Organism: Influenza a virus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.89 Å Release Date: 2026-05-06 Classification: ANTIVIRAL PROTEIN |
 |
Structure Of Escherichia Coli Phosphoenolpyruvate Carboxykinase With Bound Atp, Mg2+, And Mn2+
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-04-29 Classification: LYASE Ligands: ATP, MN, MG |
 |
Organism: Homo sapiens, Chlorocebus aethiops
Method: ELECTRON MICROSCOPY Resolution:3.70 Å Release Date: 2026-04-29 Classification: VIRUS Ligands: SPH |

