Search Count: 73,598
All
Selected
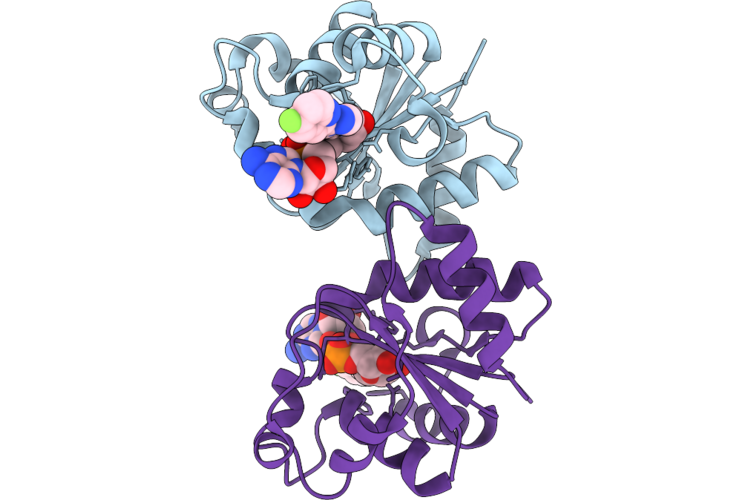 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: A1C9A |
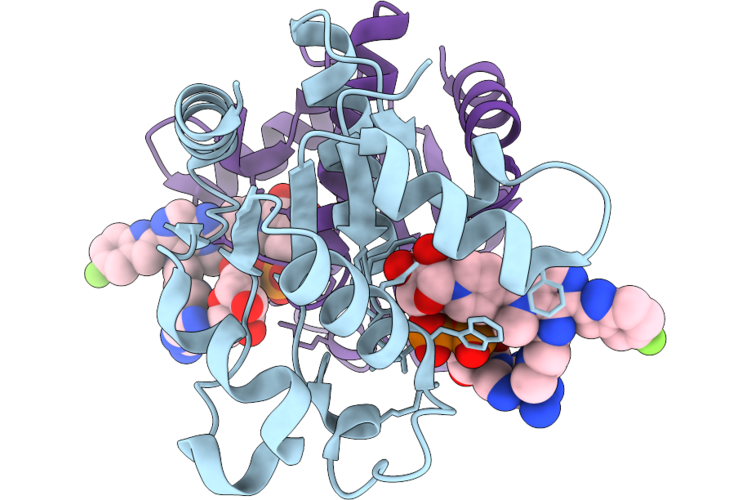 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: A1C9H |
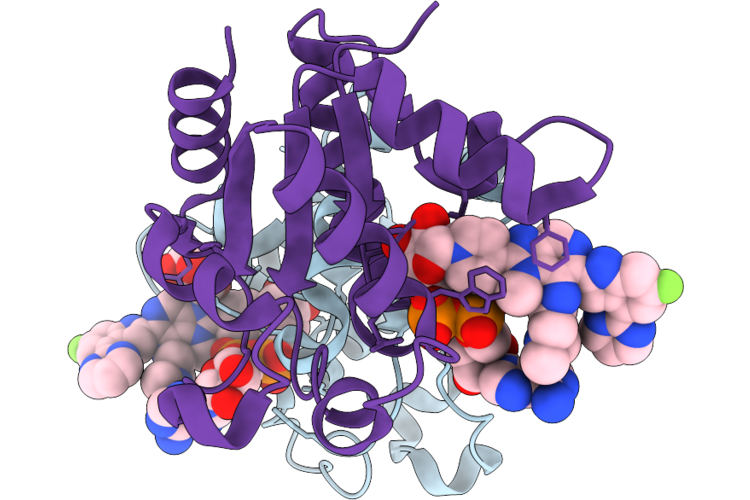 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: GOL, A1C9I |
 |
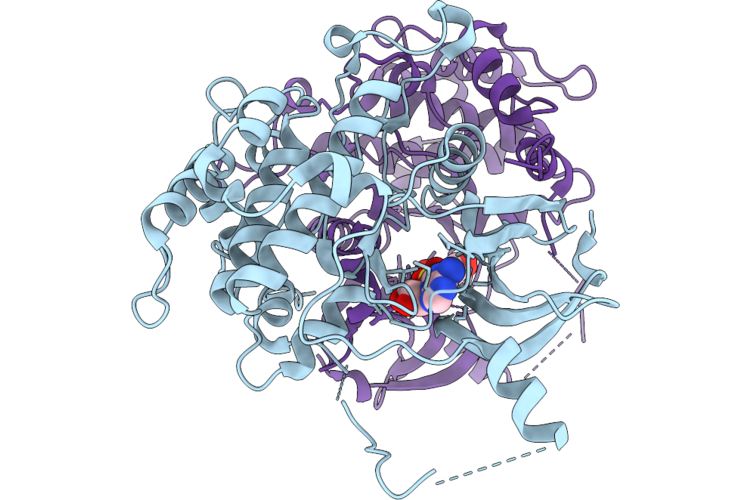
Structure Of Bacillus Subtilis Nitric Oxide Synthase In Complex With 7-((3-(((3-(6-Aminopyridin-2-Yl)Propyl)Amino)Methyl)Phenoxy)Methyl)Quinolin-2-Amine
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, V5D, CL, PEG, GOL, POL |
 |
Crystal Structure Of Serine/Threonine-Protein Kinase (Aek1) From Trypanosoma Cruzi In Complex With Lms
Organism: Trypanosoma cruzi
Method: X-RAY DIFFRACTION Resolution:2.82 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: LMS |
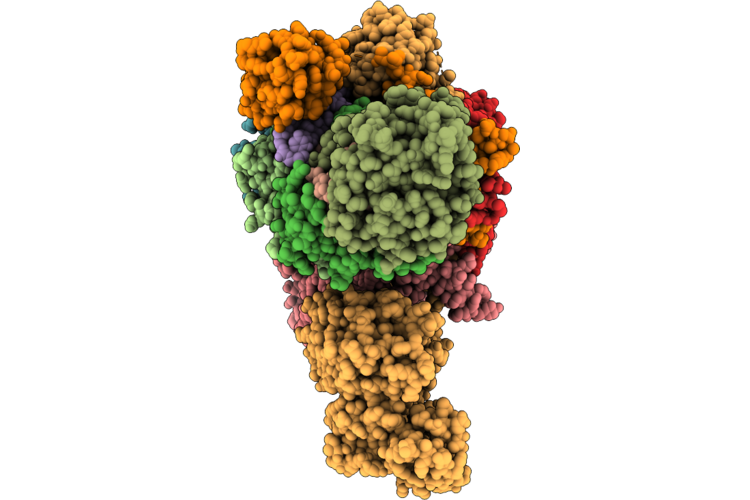 |
Cryo-Em Structure Of Type Vii Crispr-Cas Complex At The Target Engagement State
Organism: Metagenome
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: ANTIVIRAL PROTEIN/RNA Ligands: ZN |
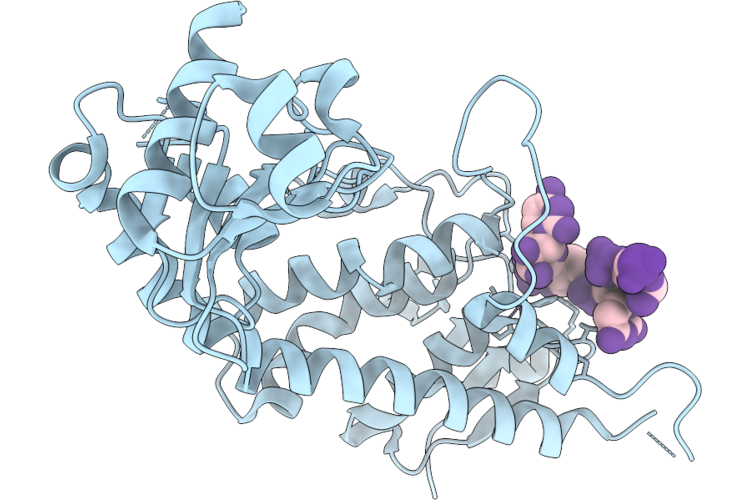 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-22 Classification: HYDROLASE/RNA Ligands: MN |
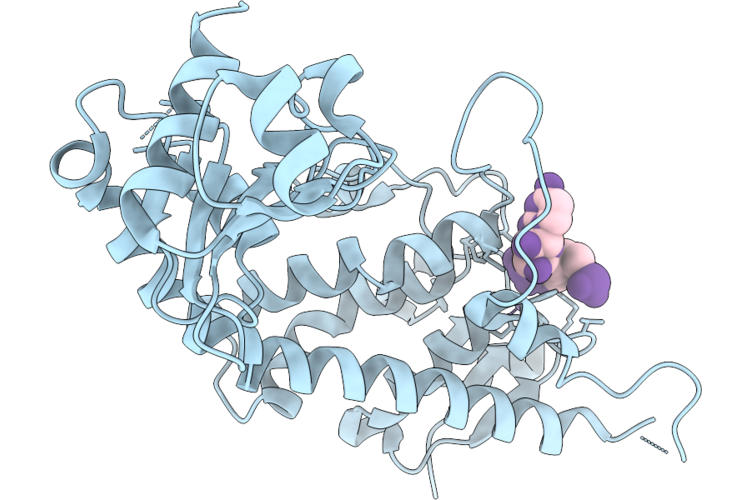 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2026-04-22 Classification: HYDROLASE/RNA Ligands: MN |
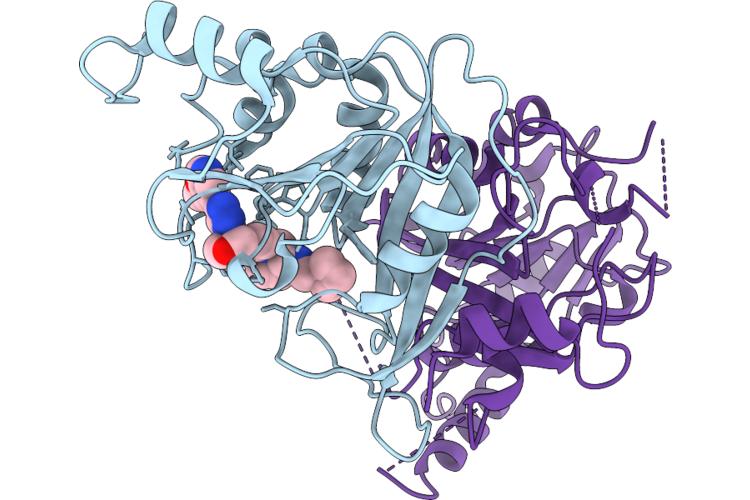 |
Crystal Structure Of The Human Mettl3-Mettl14 In Complex With Small Molecule Inhibitor Compound 8
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-04-22 Classification: RNA BINDING PROTEIN Ligands: A1I4E |
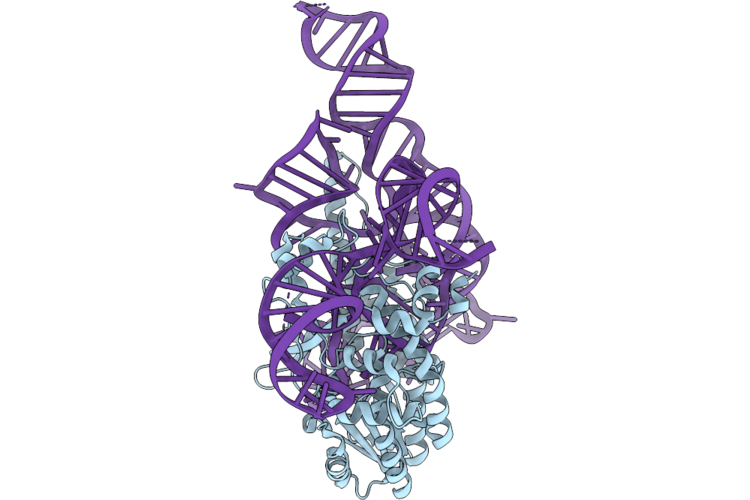 |
Organism: Klebsiella pneumoniae
Method: ELECTRON MICROSCOPY Resolution:3.49 Å Release Date: 2026-04-22 Classification: ANTIVIRAL PROTEIN Ligands: MG |
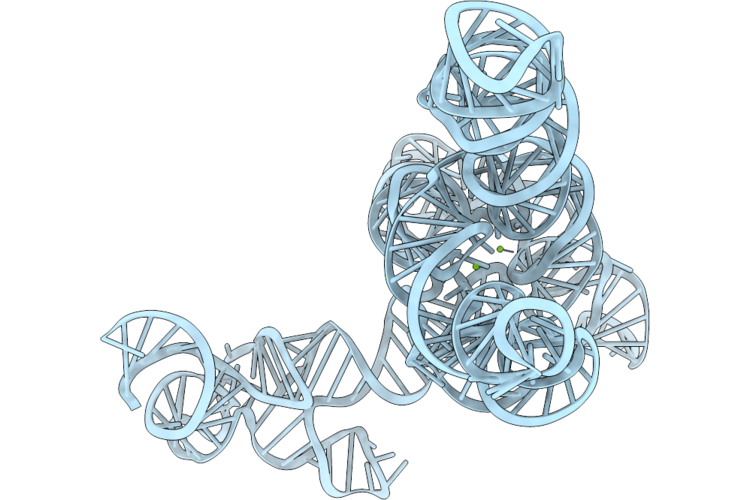 |
Organism: Anabaena
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: RNA Ligands: MG |
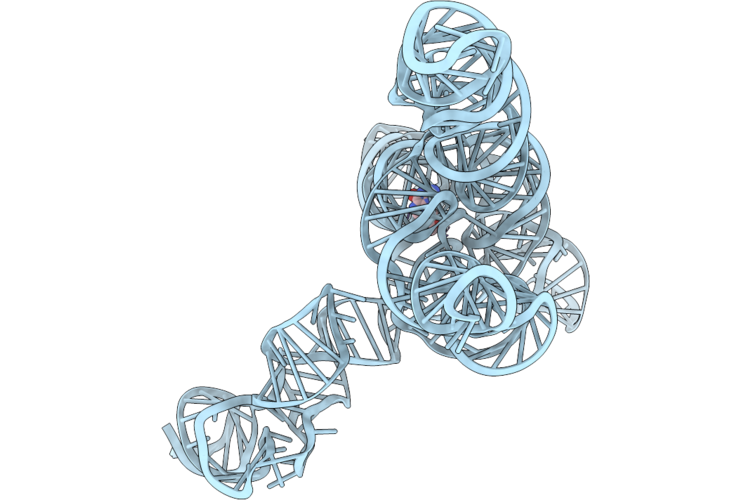 |
Organism: Anabaena
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: RNA Ligands: MG, GMP |
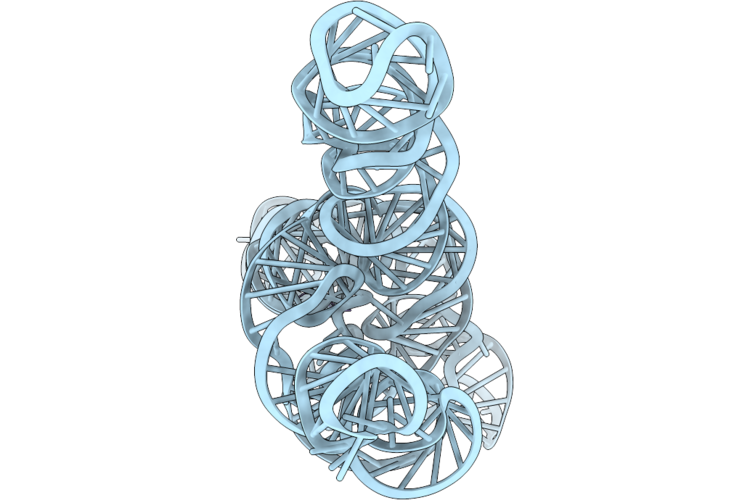 |
Organism: Anabaena
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: RNA Ligands: MG |
 |
Organism: Anabaena
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: RNA Ligands: MG |
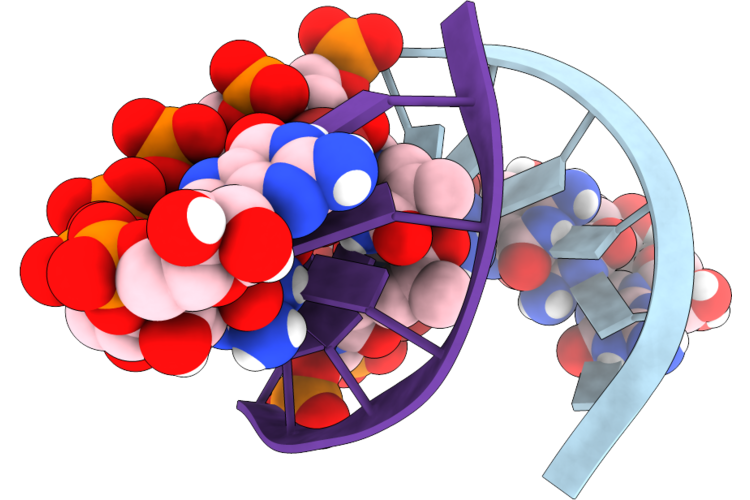 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-04-22 Classification: RNA Ligands: A1B9T, 5BU, 5GP, C5P, C |
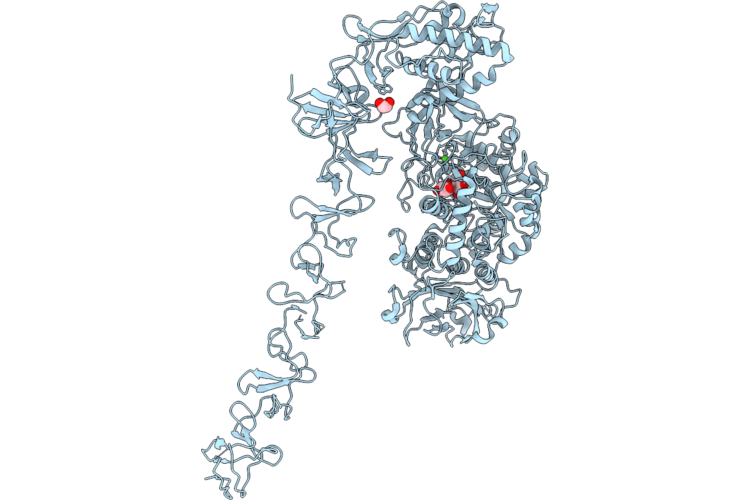 |
Structure Of Full-Length Streptococcus Mutans Gtfd With Inhibitor Maltose In Active Site
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: CA, EDO |
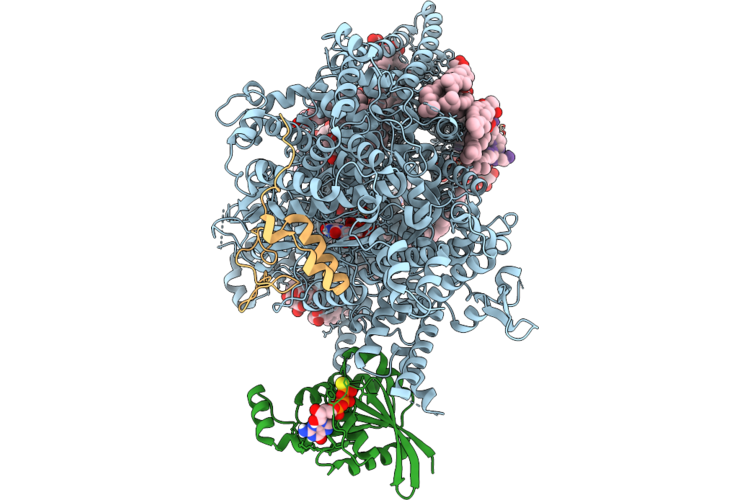 |
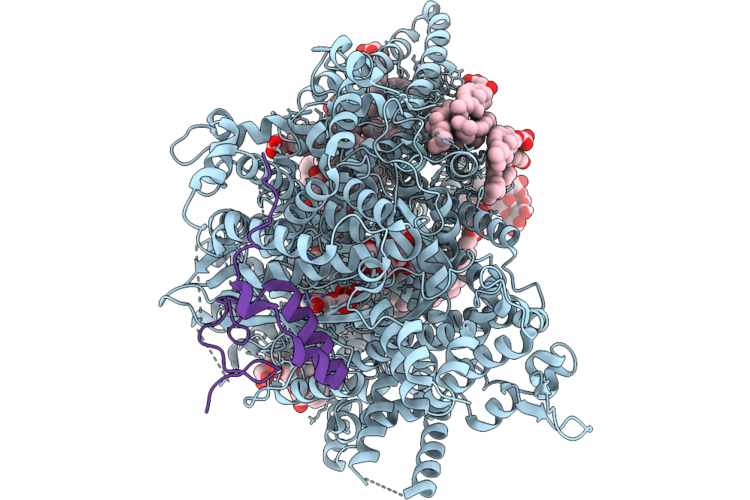
Structure Of Beta-1,3-Glucan Synthase In Complex With Caspofungin, Rho1 And Long Glucan
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP, GSP, A1CHR |
 |
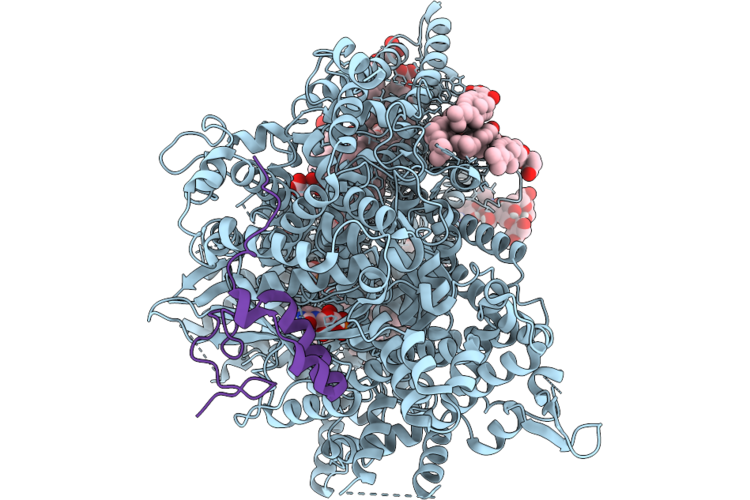
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) In Complex With Short Glucan
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP |
 |
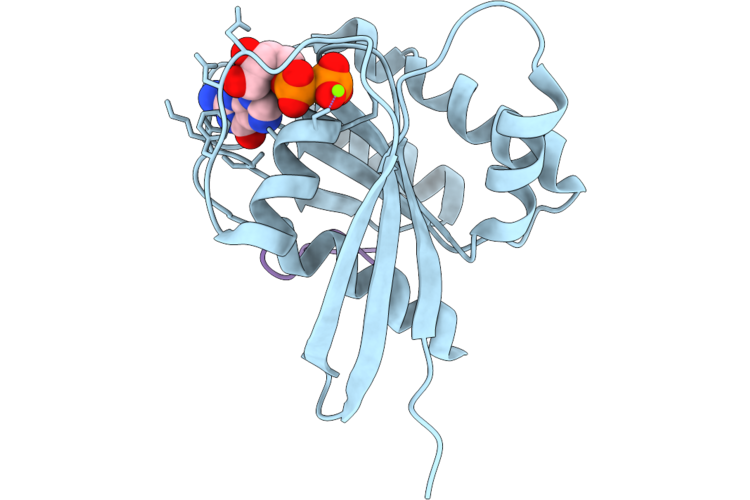
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Relevant Ground State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |

