Search Count: 1,10,007
All
Selected
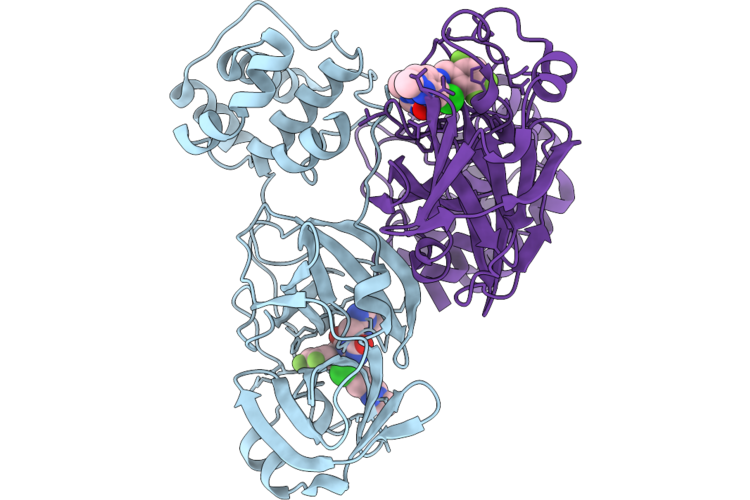 |
Room Temperature X-Ray Structure Of Sars Cov-2 Main Protease Intermediate Precursor With Ensitrelvir (Esv)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-05-06 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: 7YY |
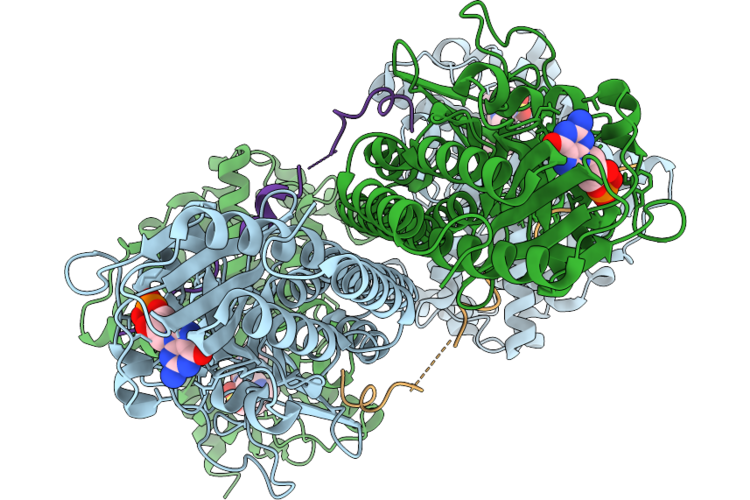 |
Organism: Bos taurus
Method: ELECTRON MICROSCOPY Resolution:3.06 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: VIA, PCG, MG, ZN |
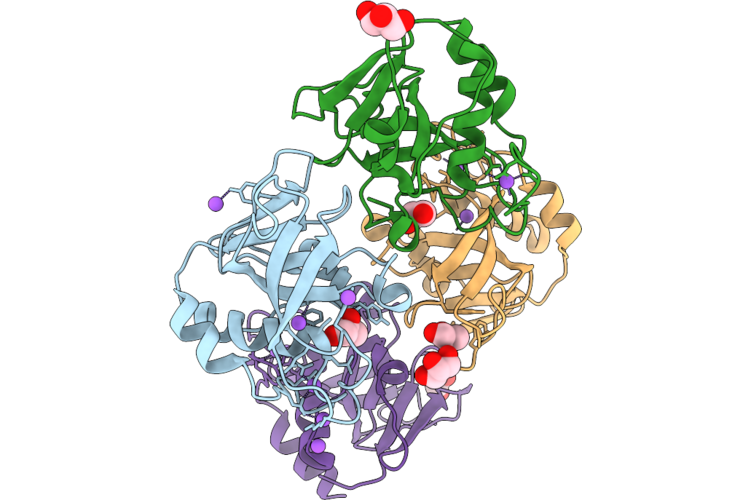 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.25 Å Release Date: 2026-05-06 Classification: HYDROLASE |
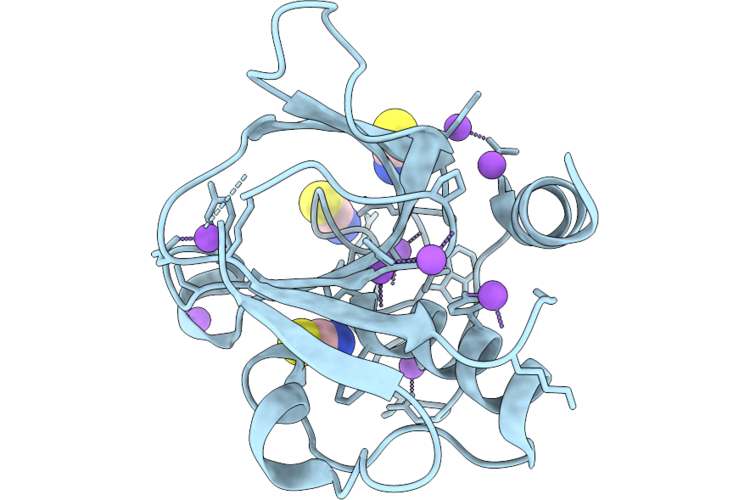 |
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.15 Å Release Date: 2026-05-06 Classification: HYDROLASE |
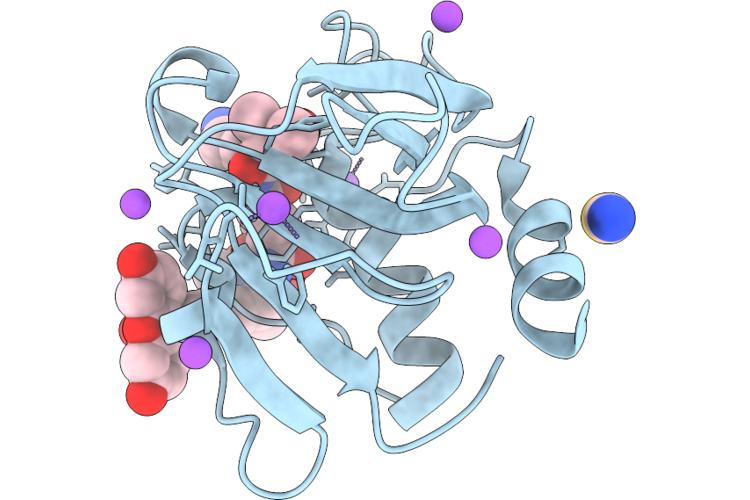 |
Chap Domain Of Staphylococcus Aureus-Specific Lysin L1 Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.09 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA, SCN |
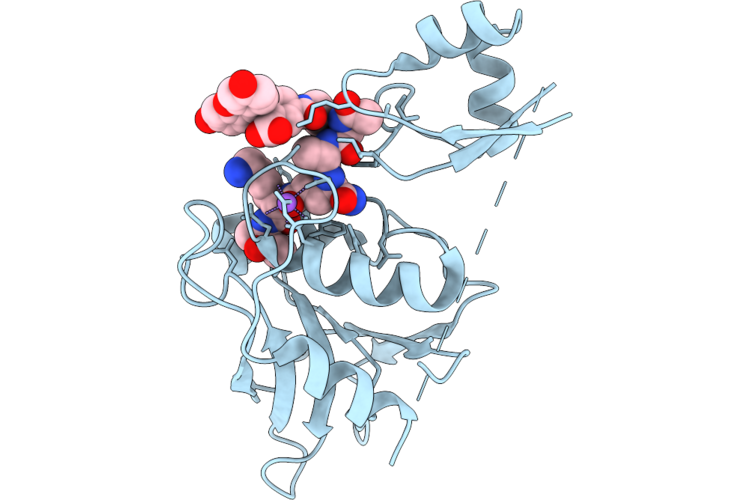 |
Staphylococcus Aureus-Specific Lysin L1-3 (Lysm-Chap) Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA |
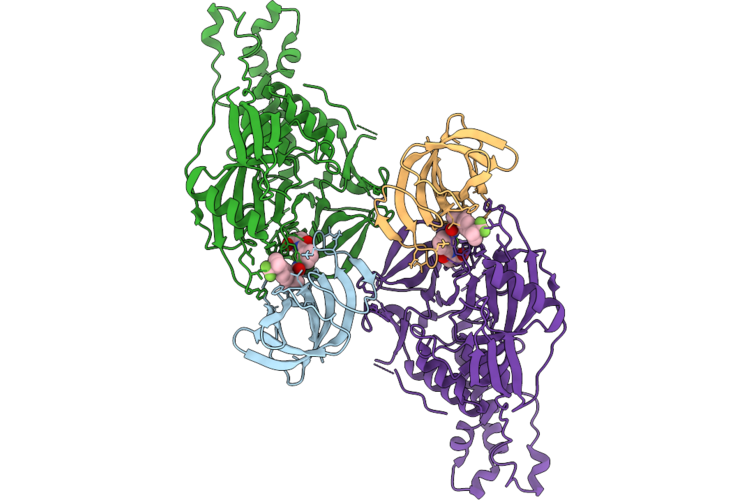 |
Cryo-Em Structure Of Crbn In Complex With Hbs1L And Tng-4857 (Focused Refinement)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: CYTOSOLIC PROTEIN Ligands: ZN, A1C9W |
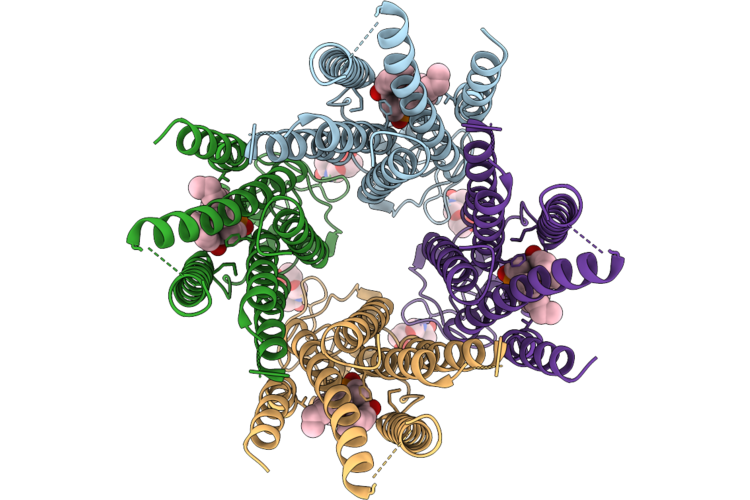 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: CPL, NAG |
 |
Organism: Homo sapiens, Mus musculus, Bos taurus, Escherichia coli, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: SIGNALING PROTEIN |
 |
Organism: Rattus norvegicus, Bos taurus, Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: SIGNALING PROTEIN/IMMUNE SYSTEM Ligands: CLR |
 |
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Composite Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.32 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
 |
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Consensus Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.91 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
 |
Organism: Homo sapiens
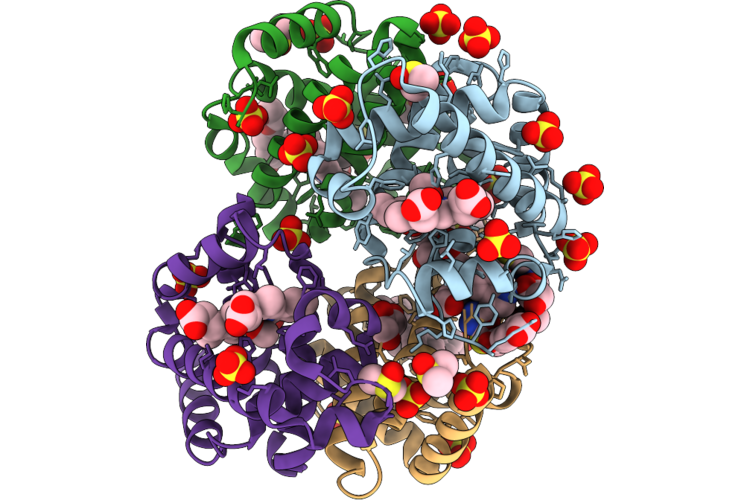
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: HEM, GOL, A1ITR, DMS, CMO, ACT, SO4, PGE |
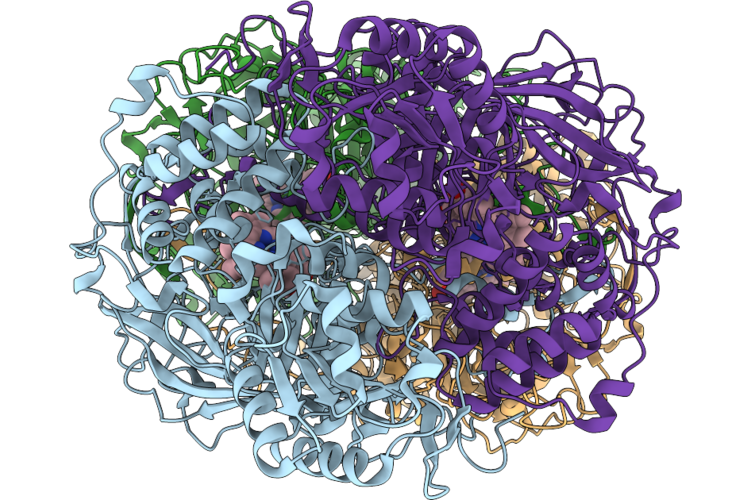 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: HEM |
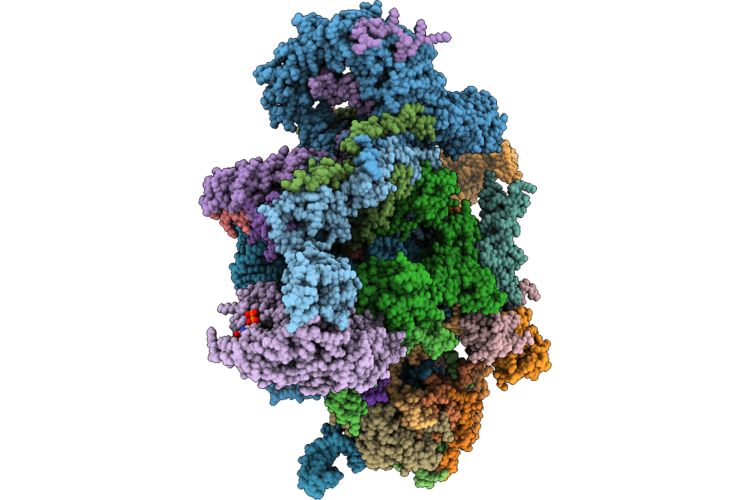 |
Assembly Intermediate Of Human Mitochondrial Ribosome Small Subunit In Complex With Noa1 And Partial Rbfa (State N2)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: RIBOSOME Ligands: MG, K, ZN, FES, ATP, GDP |
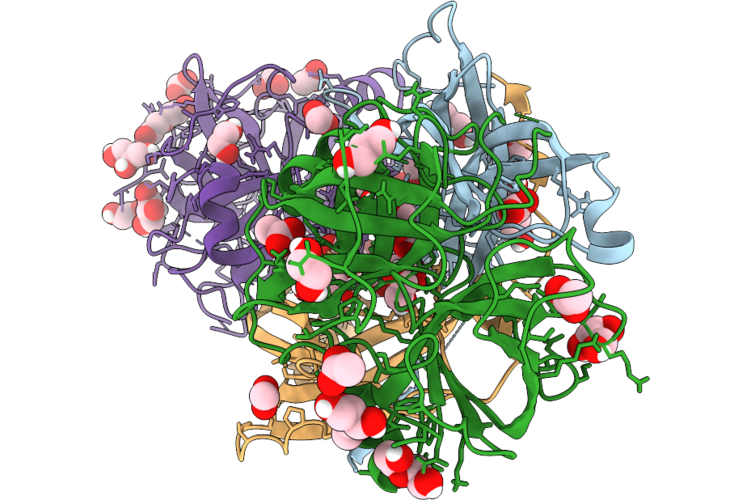 |
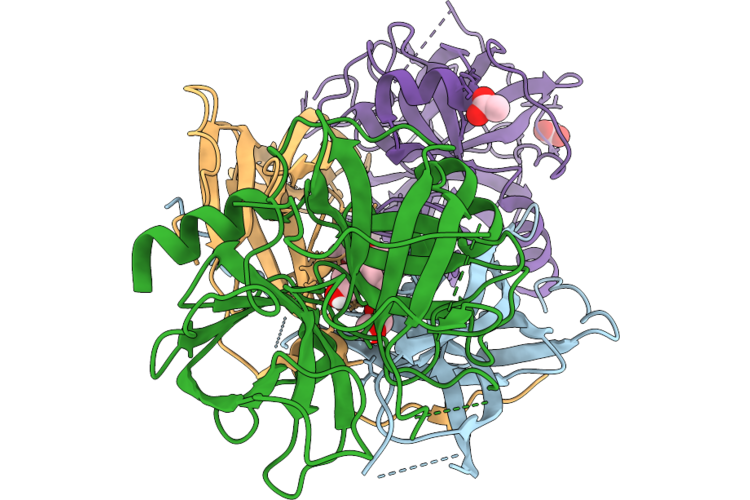
Organism: Leishmania braziliensis, Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: PEG, ACT, GOL, EDO, CL |
 |
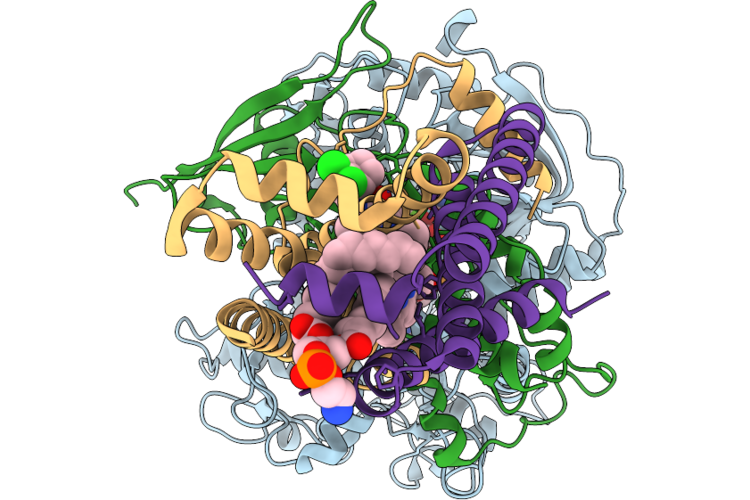
Organism: Bos taurus, Leishmania major
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ACT, GOL, PG4 |
 |
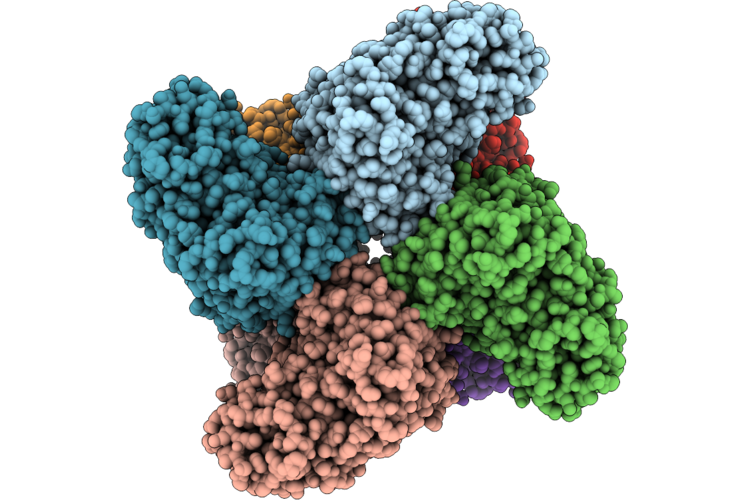
The Cryo-Em Structure Of Human Succinate Dehydrogenase In Complex With Benzovindiflupyr
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FAD, FES, SF4, F3S, A1EGM, HEM, PEV |
 |
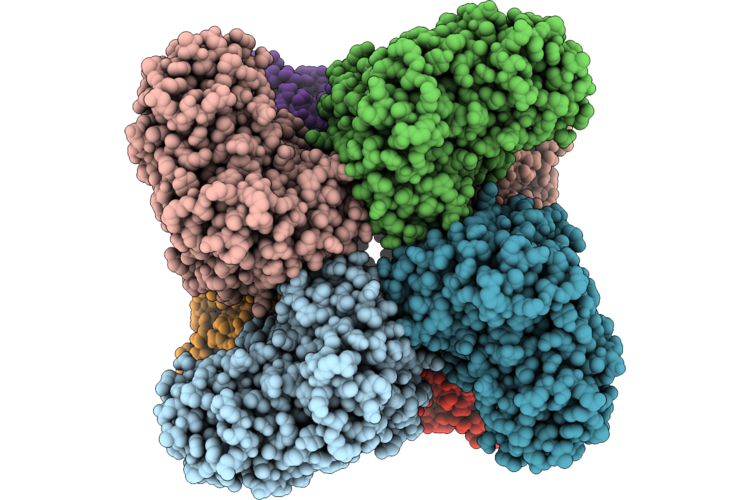
Cryo-Em Structure And Rational Engineering Of A Novel Efficient Ochratoxin A-Detoxifying Amidohydrolase
Organism: Pseudoxanthomonas wuyuanensis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN, 97U |
 |
Cryo-Electron Microscopic Structure Of A Novel Amidohydrolase With Three Mutations
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN, 97U |

