Search Count: 27,506
All
Selected
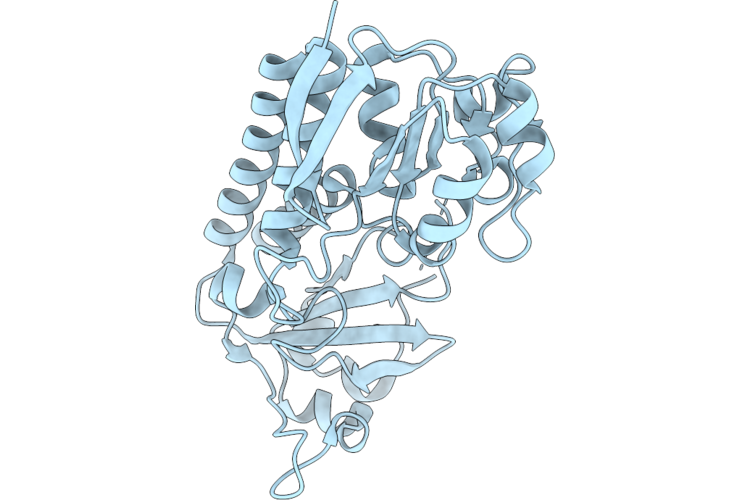 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus In The Apo-Form
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN |
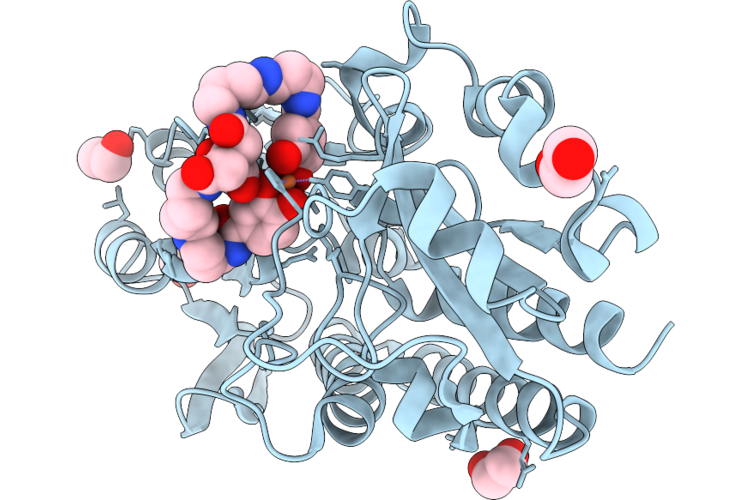 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus Complexed With Ferric Petrobactin
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN Ligands: F8W, FE, EDO |
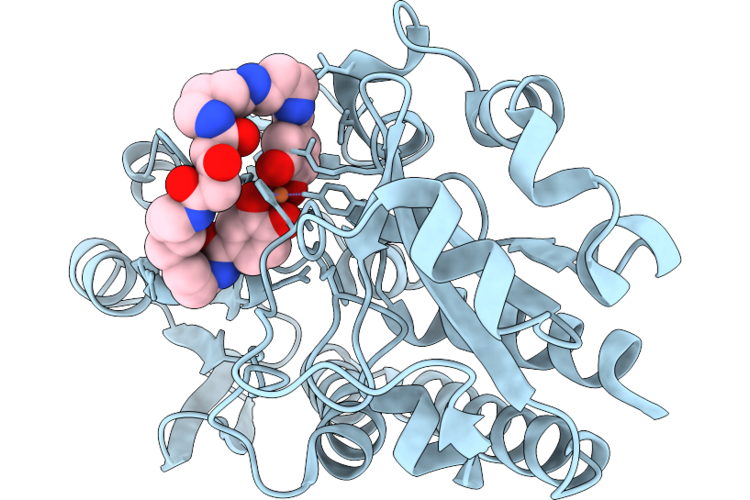 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus Complexed With Ferric Petrobactin Photoproduct, Fepbv
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN Ligands: A1MDH, FE |
 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus Complexed With Ferric Siderophore Mimic, Fe(3,4-Dhb)2
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN Ligands: DHB, FE, EDO, IPA |
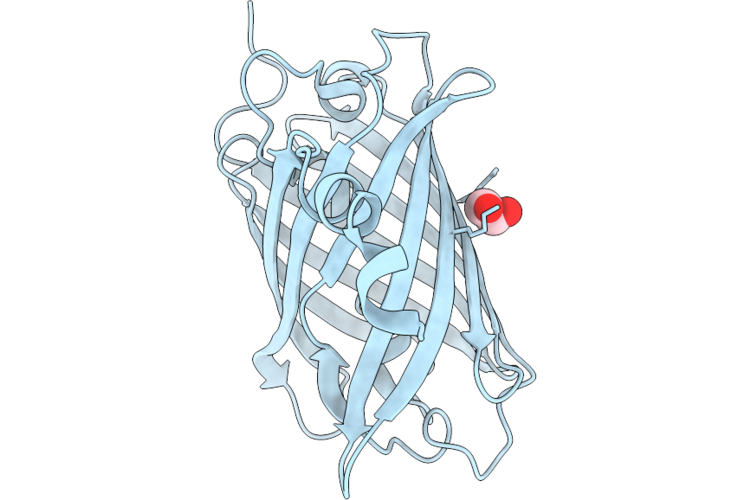 |
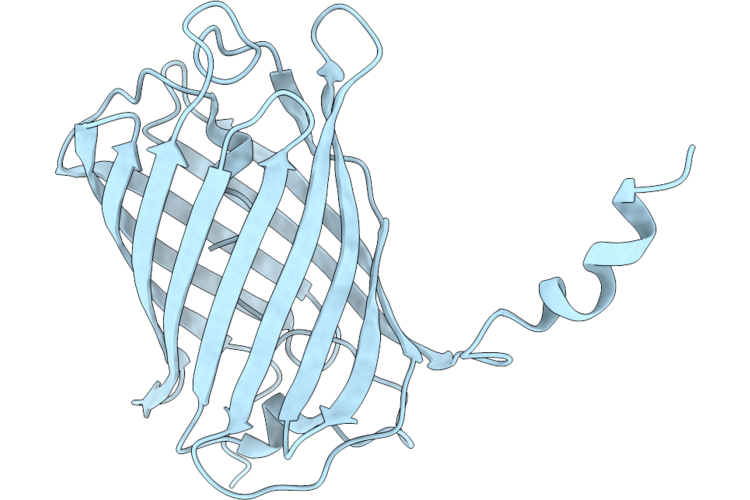
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN Ligands: EDO |
 |
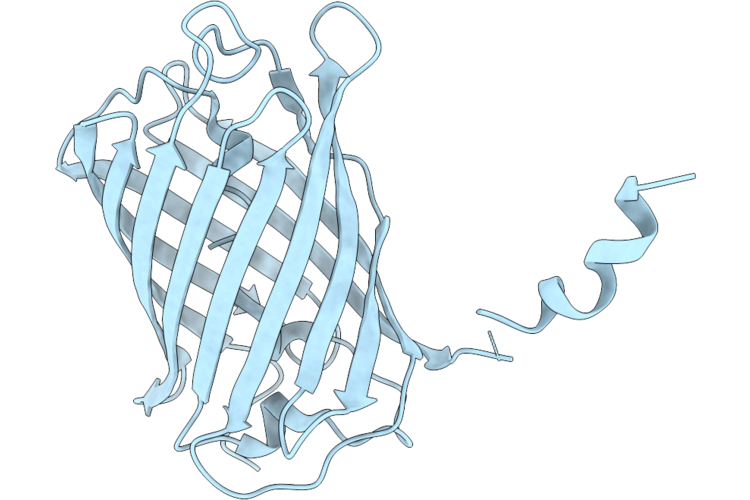
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN |
 |
Crystal Structure Of Lssmorange Under Room Temperature At Ph 8.0, 250Ps After Pump Laser, Refined Against 8.33 Times Extrapolated Structure Factor
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN |
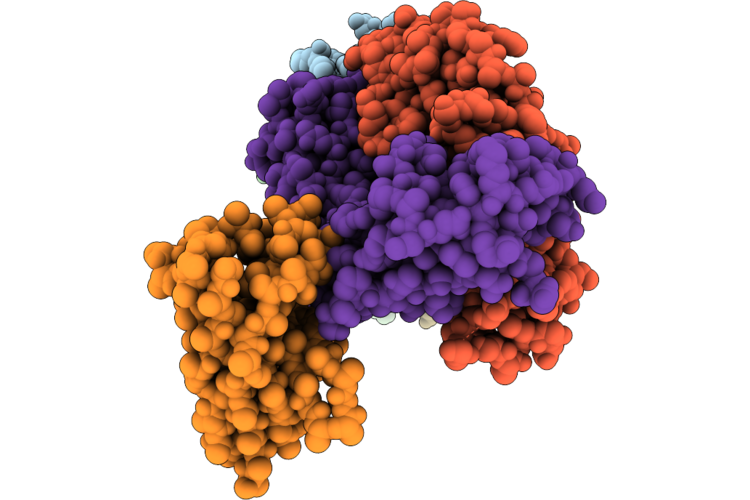 |
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: IMMUNE SYSTEM/Hydrolase |
 |
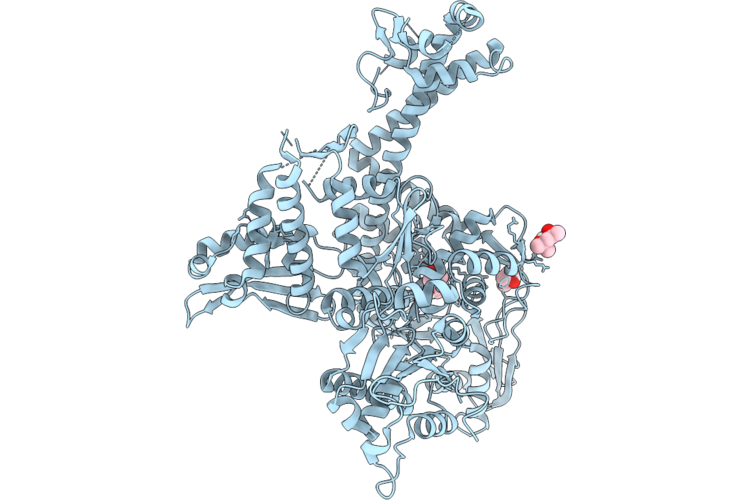
Crystal Structure Of A P. Aeruginosa Gyrase Chimera In Complex With 20Mer Dna And Ciprofloxacin
Organism: Pseudomonas aeruginosa, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-04-22 Classification: DNA BINDING PROTEIN Ligands: CIT, CPF, SO4, MN, CL |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-22 Classification: DNA BINDING PROTEIN |
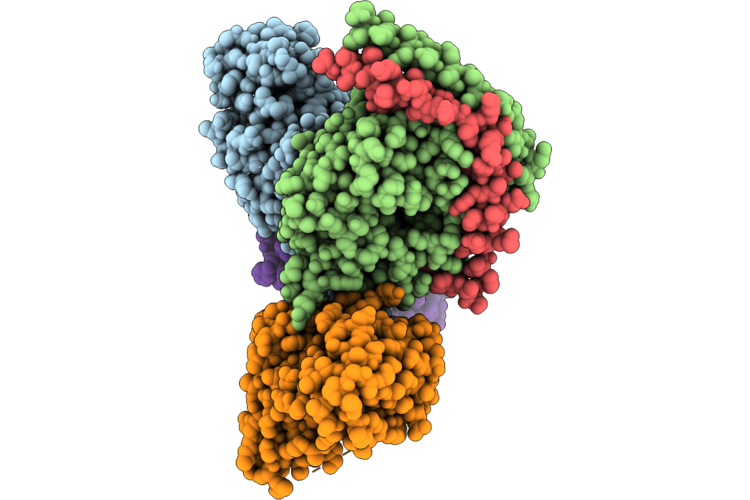 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: GOQ |
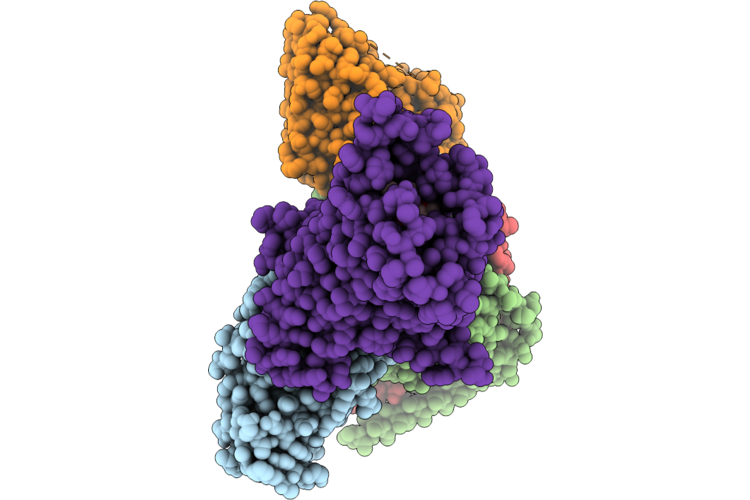 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: GOQ |
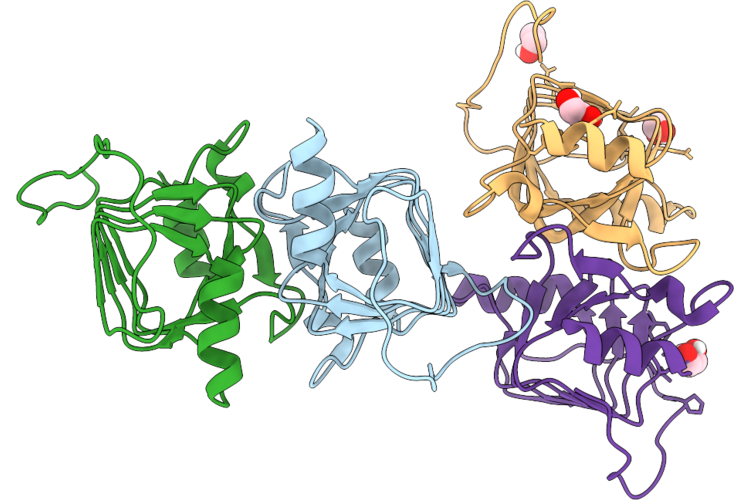 |
Organism: Brevundimonas sp. sh203
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-22 Classification: UNKNOWN FUNCTION Ligands: EDO |
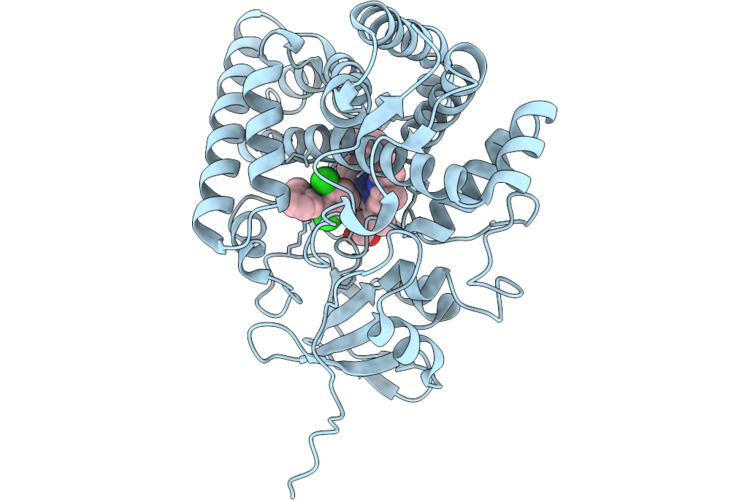 |
Crystal Structure Of Cyp105A1 R84A Complexed With Diclofenac (Dif) At Room Temperature
Organism: Streptomyces griseolus
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, DIF |
 |
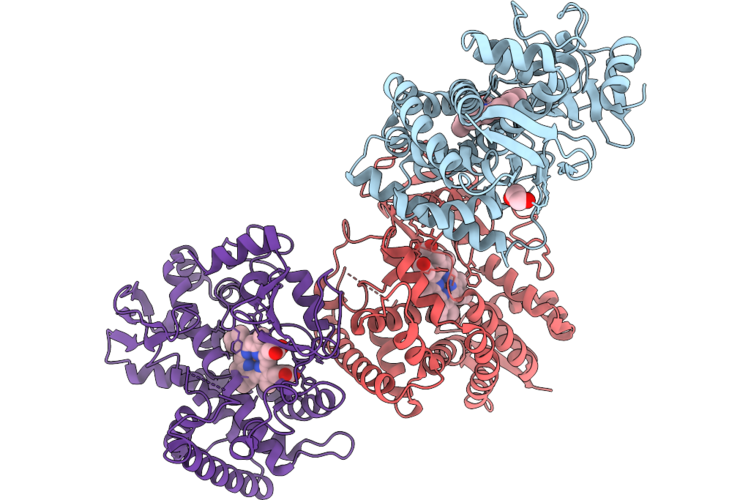
Organism: Micromonospora rosaria
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
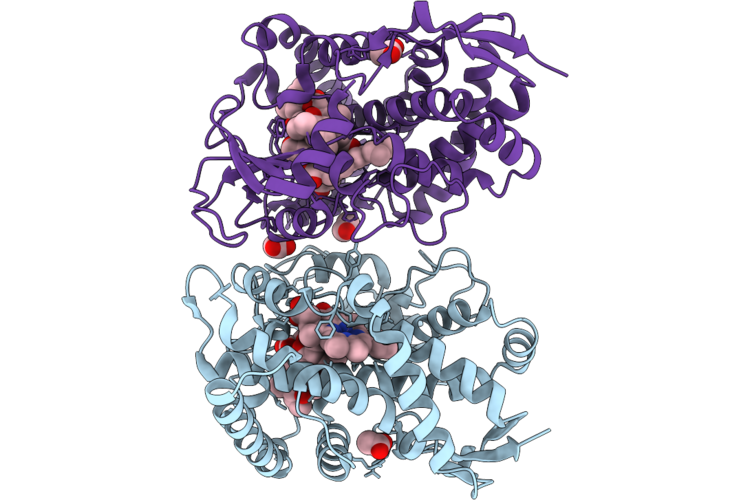
Organism: Micromonospora rosaria
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, EDO |
 |
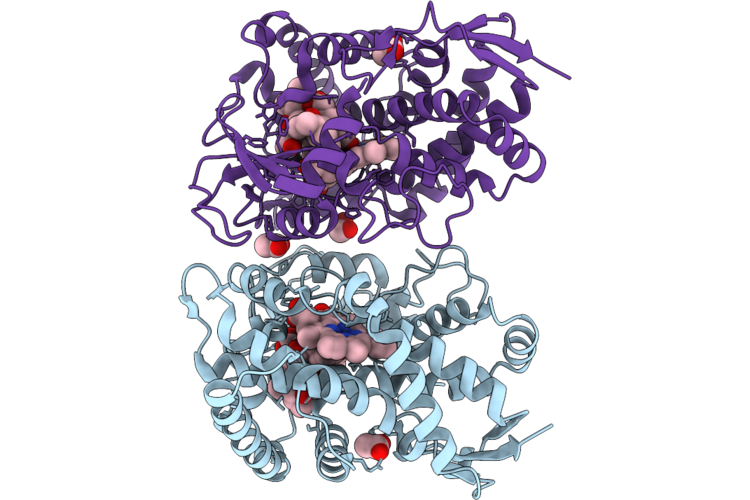
Organism: Micromonospora rosaria
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: A1MAI, HEM, ACT, MLT, EDO |
 |
Crystal Structure Of The Cytochrome P450 Rosc In Complexed With 20-Dihydrorosamicin
Organism: Micromonospora rosaria
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: L0O, HEM, ACT, MLT, EDO |
 |
Crystal Structure Of The Cytochrome P450 Rosc In Complexed With 20-Deoxo-20-Dihydrorosamicin
Organism: Micromonospora rosaria
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: A1MAJ, HEM, ACT, MLT, EDO |
 |
Ago Maturation Complex (Amc): Ago2-Mirna Duplex In Complex With Hsp90 Beta And Co-Chaperone P23
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: CHAPERONE Ligands: ATP, MG |

