Search Count: 43,004
All
Selected
 |
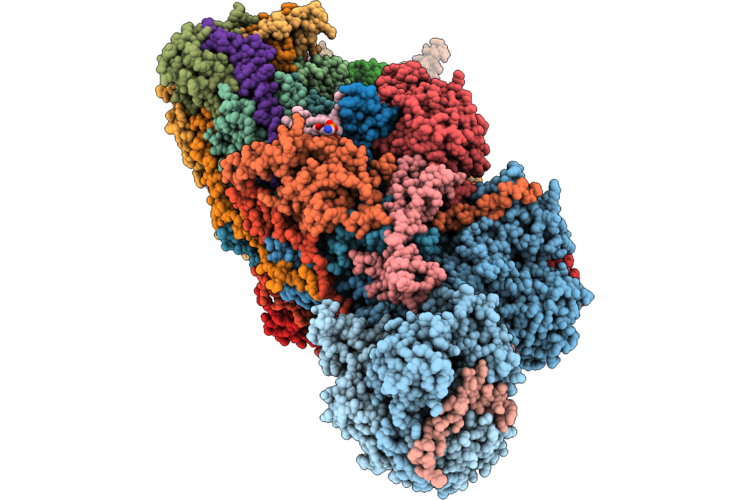
Organism: Streptococcus agalactiae
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: EDO, ZN |
 |
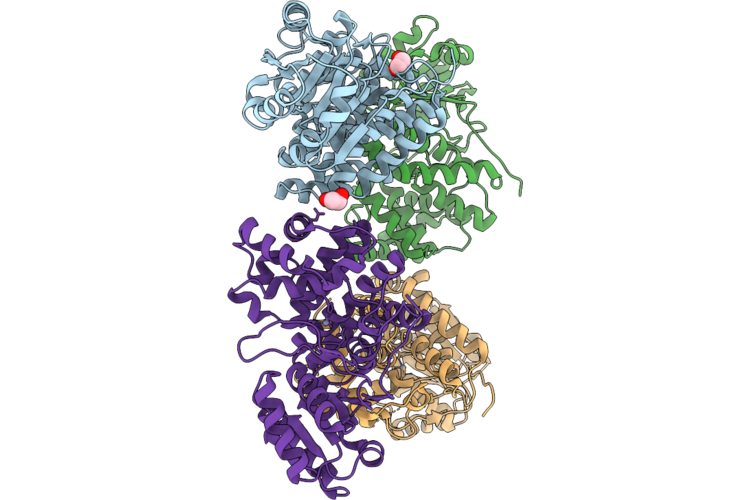
Xfel Structure Of Oxidised Ribonucleotide Reductase R2A Y122F Mutant From E. Coli
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE |
 |
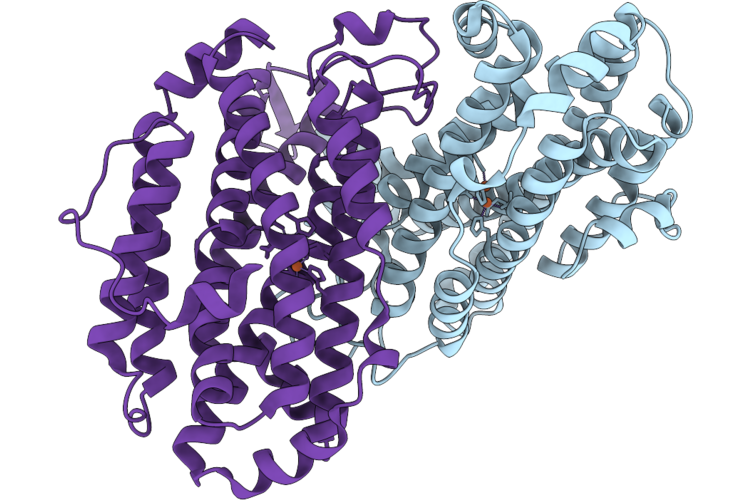
Xfel Structure Of Ribonucleotide Reductase R2A Y122F Mutant From E. Coli,Reduced Form
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE2 |
 |
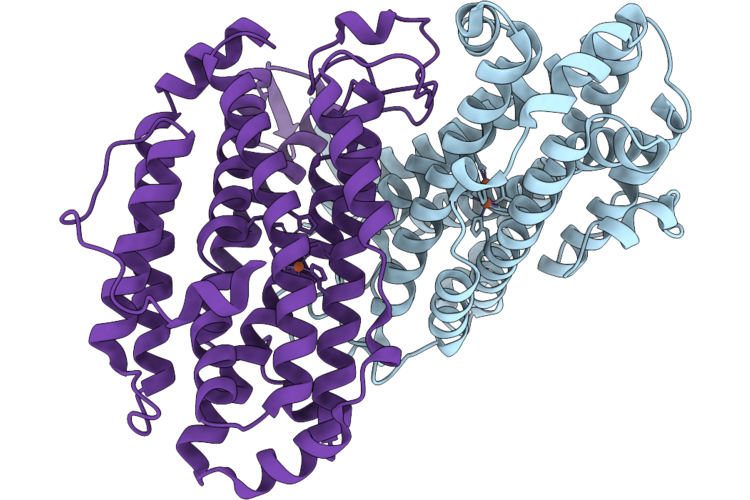
Coproheme Decarboxylase H117A Mutant From Listeria Monocytogenes In Complex With Iron Coproporphyrin Iii
Organism: Listeria monocytogenes egd-e
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FEC |
 |
Xfel Structure Of Oxidised Ribonucleotide Reductase R2A Y122F Mutant From E. Coli, Hexagonal P6122 Form
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE |
 |
Xfel Structure Of Ribonucleotide Reductase R2A Y122F Mutant From E. Coli,Reduced Form, Hexagonal P6122
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE2 |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.05 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: 3PE, HEM, HEO, CU, UQ8 |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.56 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: HEM, HEO, CU, 3PE, LMG, UQ8 |
 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.12 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: HEM, HEO, CU, 3PE, LMG, UQ8 |
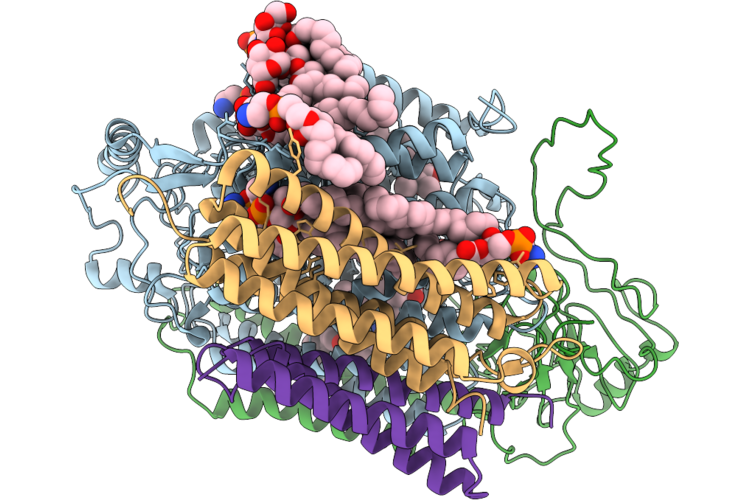 |
Organism: Acinetobacter baumannii
Method: ELECTRON MICROSCOPY Resolution:3.28 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: HEM, HEO, CU, 3PE, LMG, UQ8 |
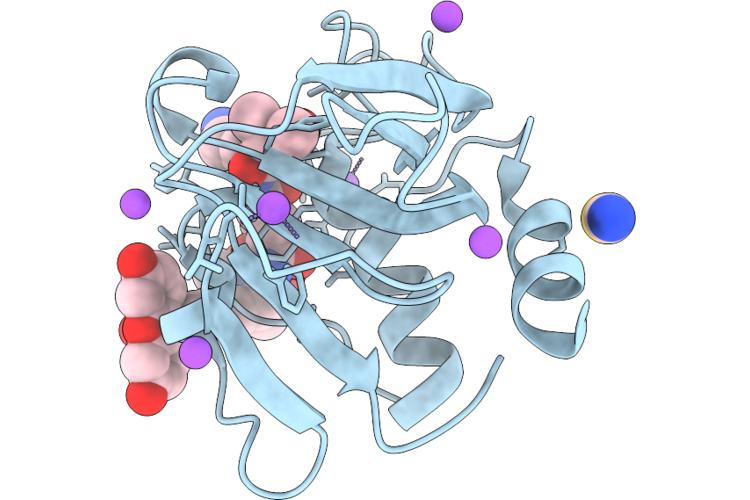 |
Chap Domain Of Staphylococcus Aureus-Specific Lysin L1 Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:1.09 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA, SCN |
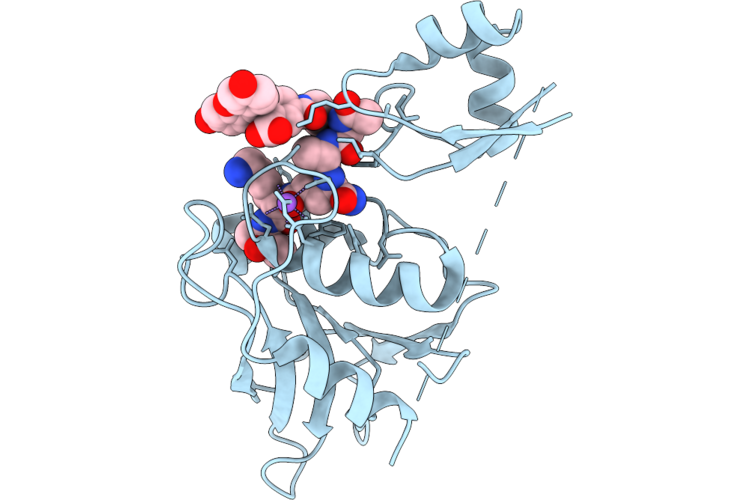 |
Staphylococcus Aureus-Specific Lysin L1-3 (Lysm-Chap) Covalently Complexed To Pep1A-Cmk Substrate Mimic
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: A1DEZ, NA |
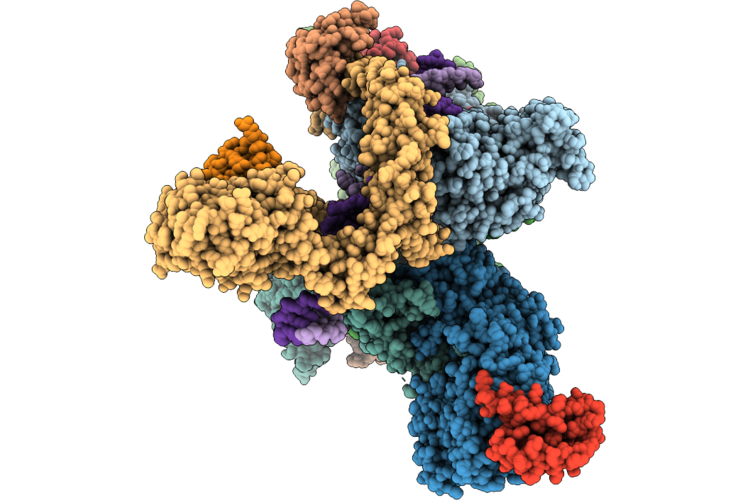 |
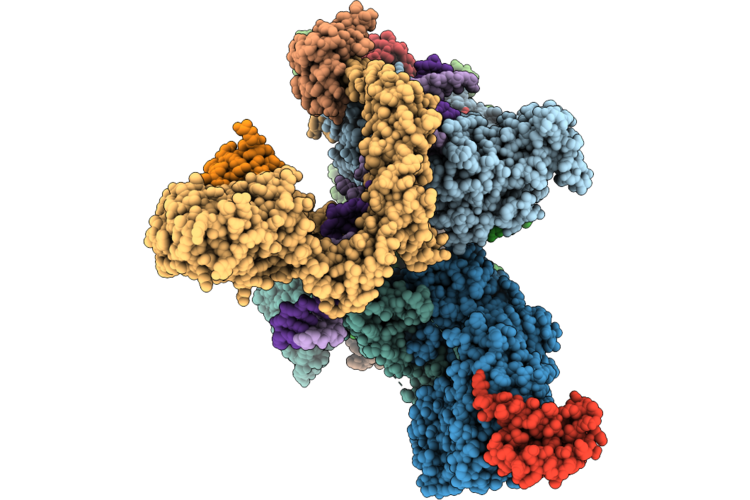
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Composite Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.32 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
 |
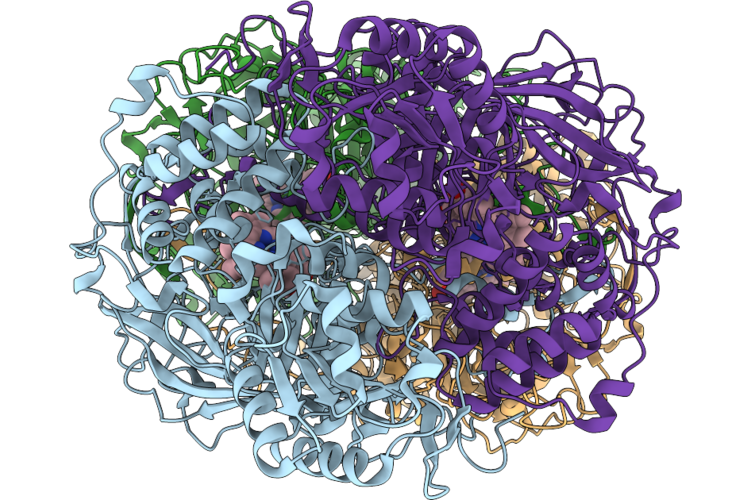
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Consensus Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.91 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
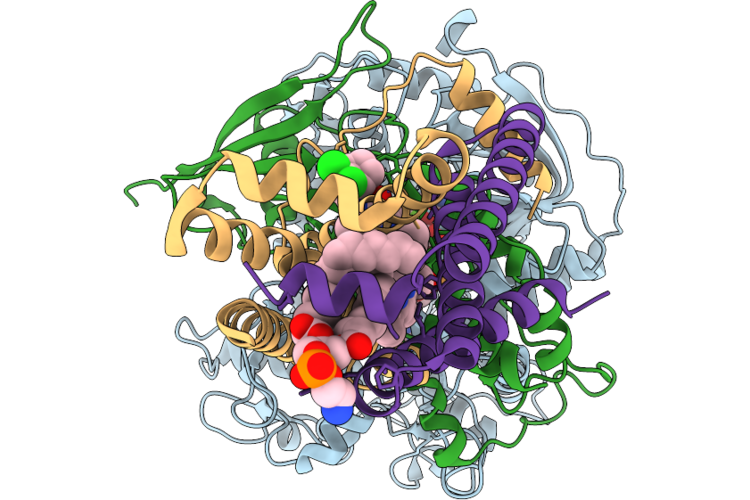
Aetf, A Single-Component Flavin-Dependent Tryptophan Halogenase, In Complex With Tryptoline
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-05-06 Classification: FLAVOPROTEIN Ligands: FAD, WYH, PEG, CL |
 |
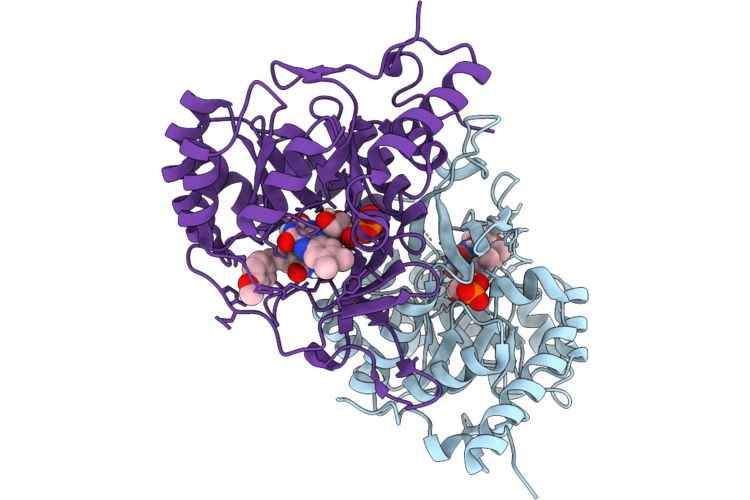
The Cryo-Em Structure Of Human Succinate Dehydrogenase In Complex With Benzovindiflupyr
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FAD, FES, SF4, F3S, A1EGM, HEM, PEV |
 |
Crystal Structure Of Dihydroorotate Dehydrogenase From Leishmania Brasiliensis In Complex With 5-[(E)-3-(P-Methoxyphenyl)-2-Propenylidene]-2,4,6(1H,3H,5H)-Pyrimidinetrione
Organism: Leishmania braziliensis
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FMN, 5TI |
 |
Crystal Structure Of Dihydroorotate Dehydrogenase From Leishmania Brasiliensis In Complex With (E)-5-(3-(4-Nitrophenyl)Allylidene)Pyrimidine-2,4,6(1H,3H,5H)-Trione
Organism: Leishmania braziliensis
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FMN, DMS, A1CAU, SO4, GOL |