Search Count: 32,509
All
Selected
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.42 Å Release Date: 2026-05-20 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: A1E5S |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.42 Å Release Date: 2026-05-20 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: A1E5T |
 |
Organism: Escherichia coli, Homo sapiens, Synthetic construct, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-05-20 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens, Mus musculus, Escherichia coli, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-05-20 Classification: MEMBRANE PROTEIN |
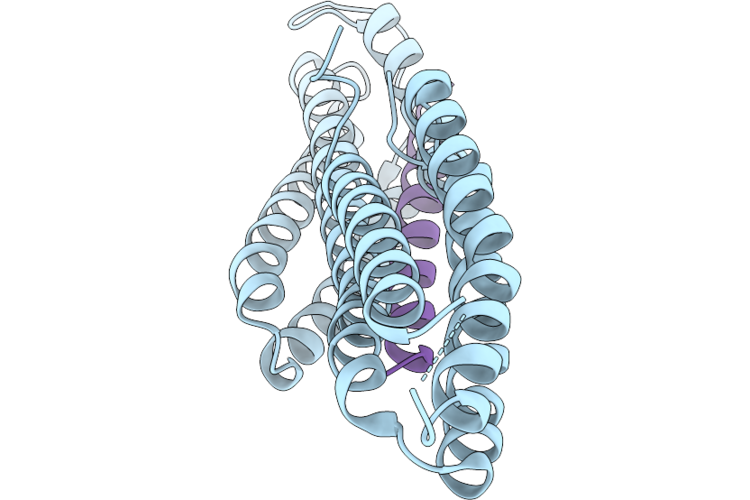 |
Mouse Vinculin Head Domain 1 (Vd1) (Residues 2-258) With A50I Mutation In Complex With Mouse Talin 1 Helix 50 (H50, Residues 2072-2103)
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-05-20 Classification: CELL ADHESION |
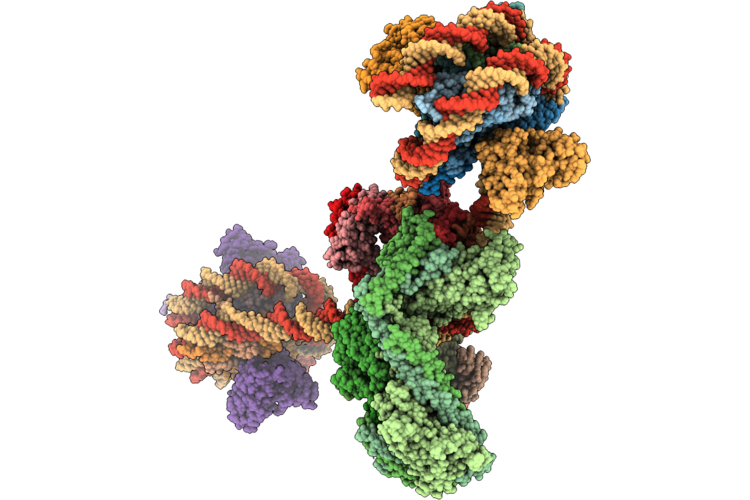 |
Structure Of The Human Inner Kinetochore Ccan Bound To A Di-Cenp-A Nucleosome
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:15.50 Å Release Date: 2026-05-20 Classification: CELL CYCLE |
 |
Cryo-Em Structure Of Active Mutant Human Green Cone Opsin (E129Q) In Complex With Chimeric G Protein (Minigist)
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: SIGNALING PROTEIN Ligands: RET |
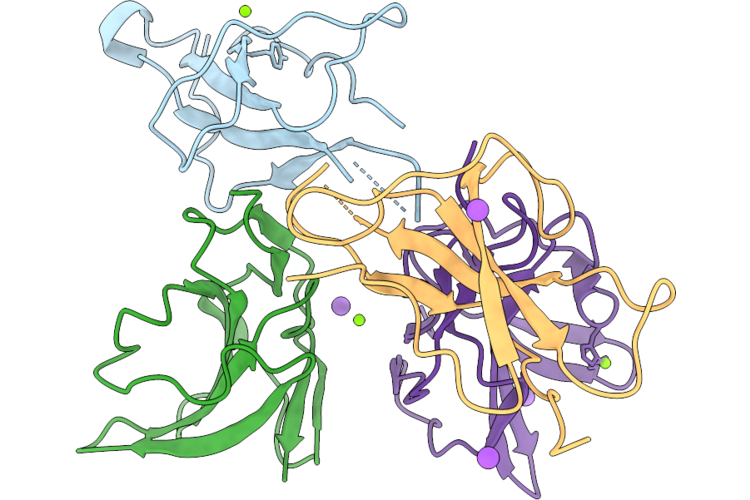 |
Organism: Heligmosomoides polygyrus, Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.52 Å Release Date: 2026-05-20 Classification: SIGNALING PROTEIN Ligands: MG, NA, CL |
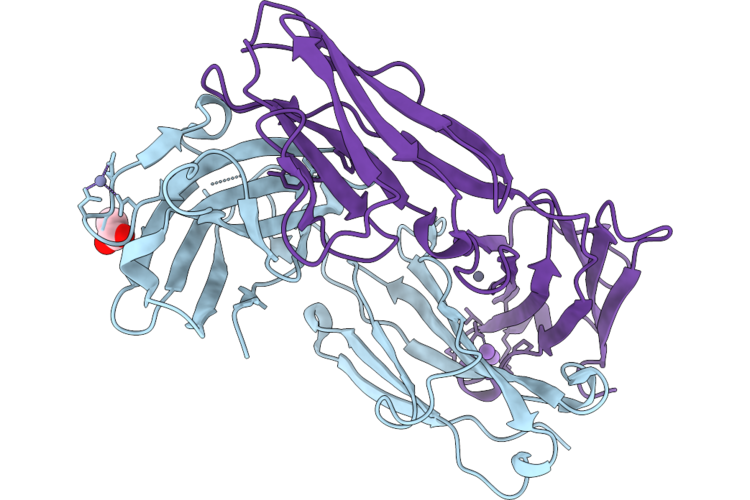 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.48 Å Release Date: 2026-05-13 Classification: PROTEIN BINDING Ligands: ACT, ZN, NA |
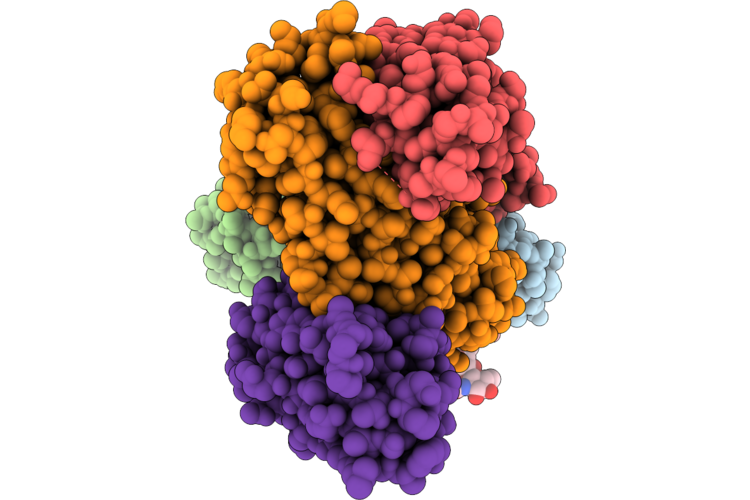 |
Structure Of The Porcine Deltacoronavirus (Pdcov) Receptor-Binding Domain Bound To The Rbd Minibinder 11, The Pd3 Fab, And The Kappa Light Chain Nanobody
Organism: Porcine deltacoronavirus, Synthetic construct, Mus musculus, Lama glama
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-05-13 Classification: VIRAL PROTEIN |
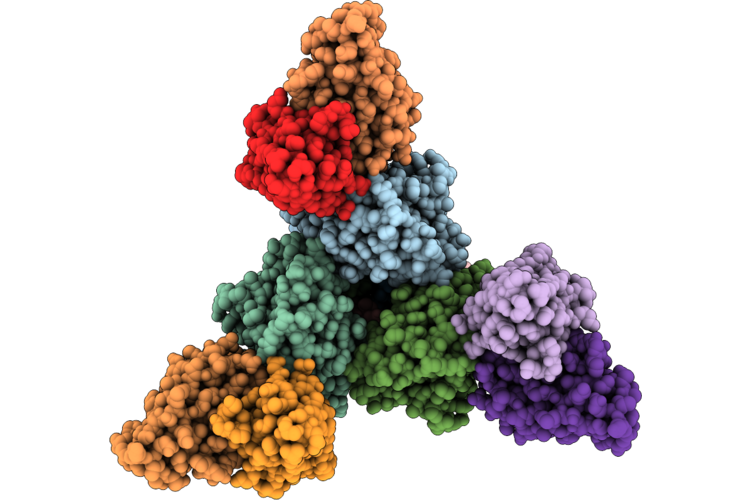 |
Organism: Influenza a virus, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.87 Å Release Date: 2026-05-13 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
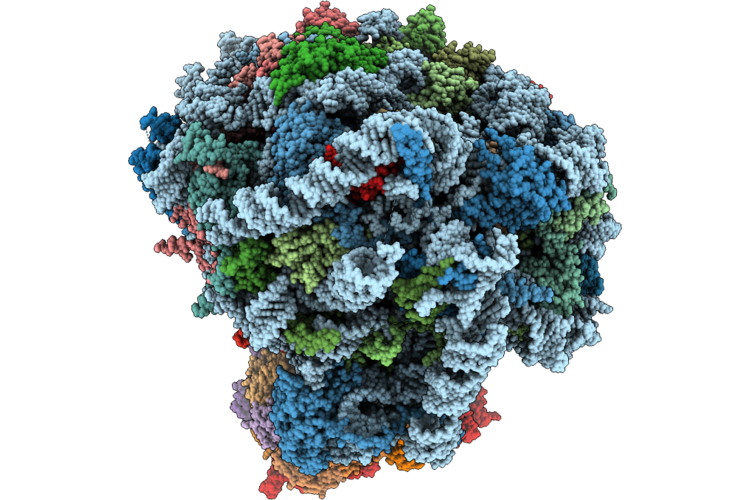 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
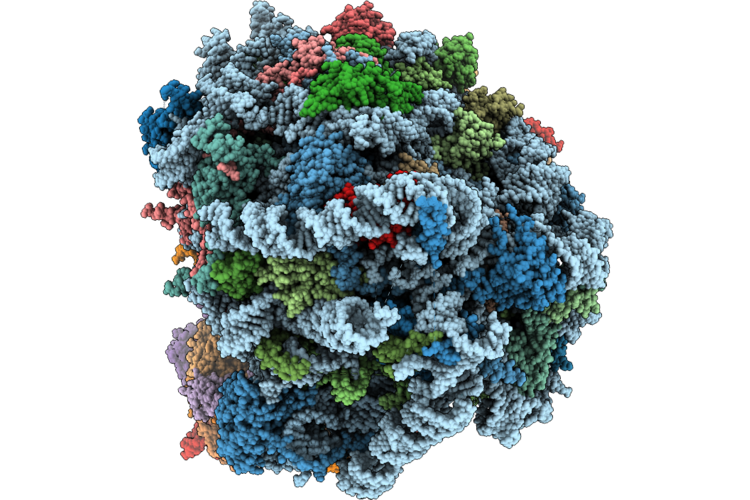 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
 |
Organism: Mus musculus, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.48 Å Release Date: 2026-05-13 Classification: TRANSFERASE/DNA Ligands: ZN |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2026-05-13 Classification: SIGNALING PROTEIN Ligands: GOL |
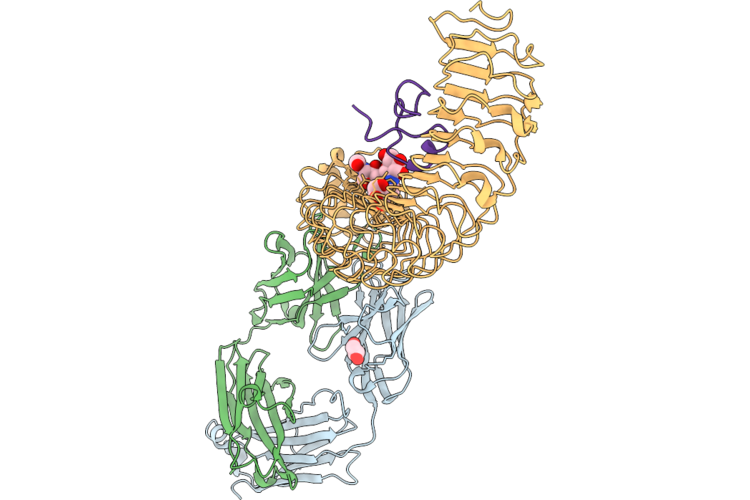 |
Mouse Lrrc15 Extracellular Domain In Complex With Disulfide Constrained Peptide Ml-Ysd-07 And Fab-E1
Organism: Homo sapiens, Mus musculus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.57 Å Release Date: 2026-05-13 Classification: SIGNALING PROTEIN/Immune System Ligands: GOL |
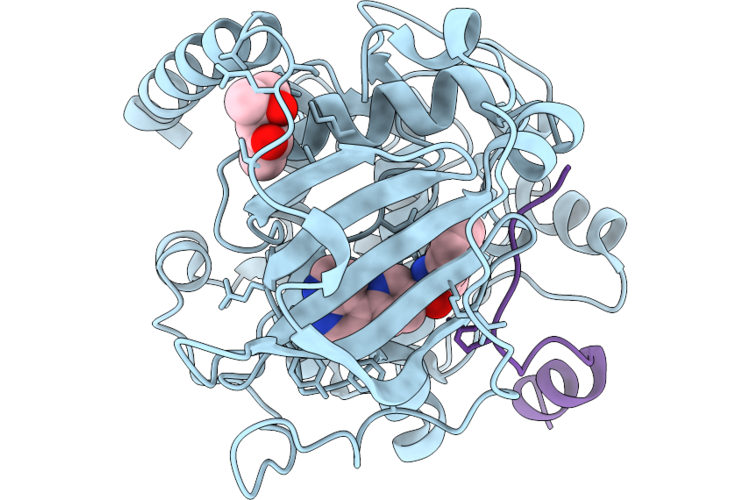 |
Co-Crystal Structure Of The Camp-Dependent Protein Kinase Catalytic Subunit Alpha With The Inhibitor Blu0588
Organism: Mus musculus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2026-05-13 Classification: TRANSFERASE Ligands: MPD, A1CHQ |
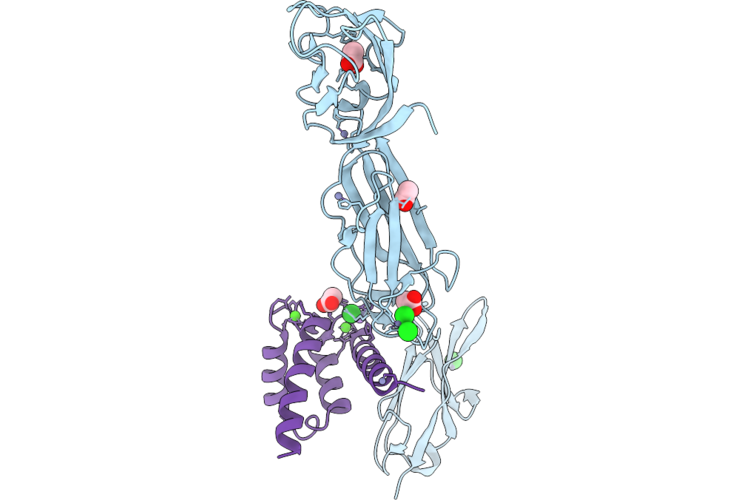 |
Crystal Structure Of The Human Rage Ectodomain In Complex With Murine S100A6 Mutant Y84C
Organism: Homo sapiens, Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-05-13 Classification: SIGNALING PROTEIN Ligands: ZN, CL, ACT, CA, NA |

