Search Count: 23,292
All
Selected
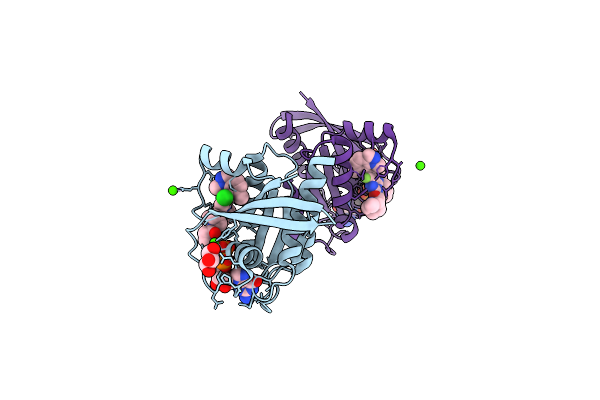 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-04-15 Classification: ONCOPROTEIN Ligands: CA, A1C6Y, GOL, GDP |
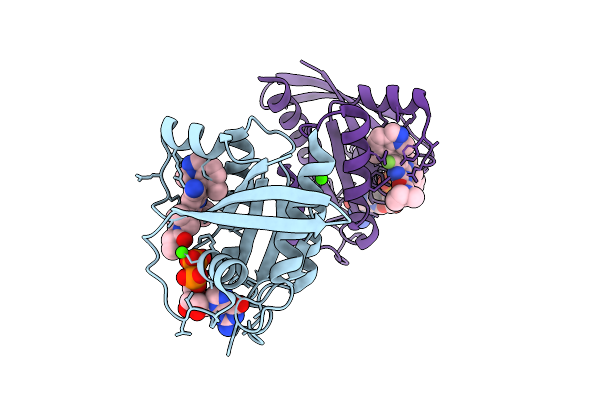 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.52 Å Release Date: 2026-04-15 Classification: ONCOPROTEIN Ligands: GDP, A1C60, CA |
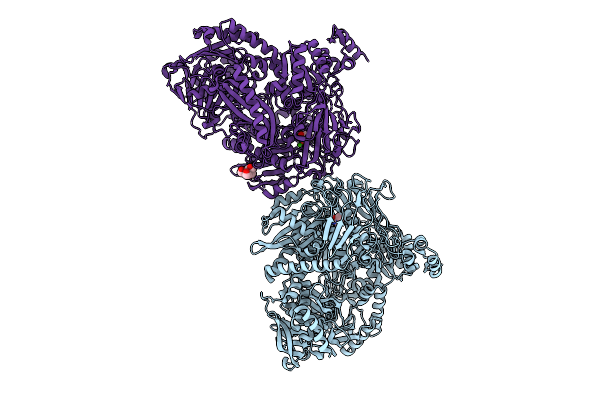 |
Crystal Structure Of N-Methylhydantoinase In Complex With 1-Methylimidazolidine-2,4-Dione
Organism: Glutamicibacter protophormiae
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: CA, A1BC1, NH4, BTB |
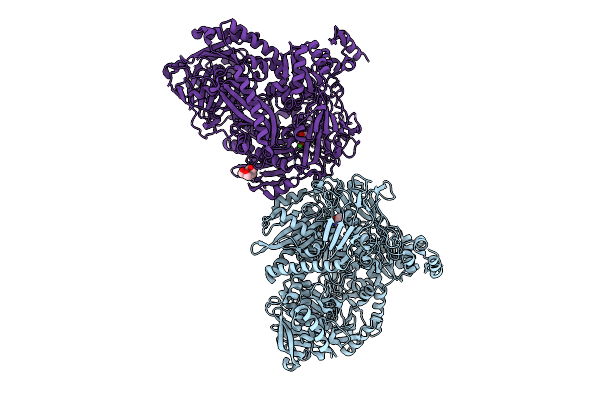 |
Crystal Structure Of N-Methylhydantoinase In Complex With 1-Methylimidazolidine-2,4-Dione, Iodide Soak
Organism: Glutamicibacter protophormiae
Method: X-RAY DIFFRACTION Resolution:2.62 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: CA, A1BC1, NH4, IOD, BTB |
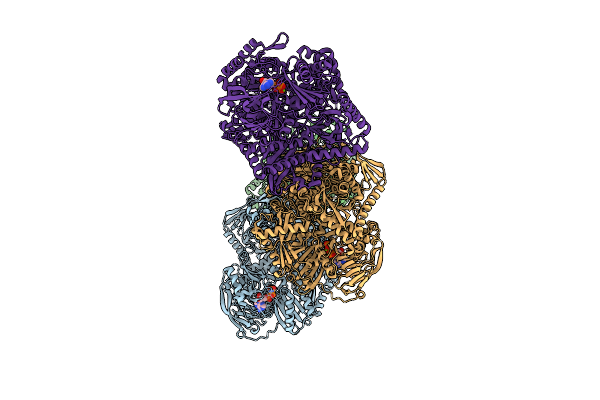 |
Organism: Glutamicibacter protophormiae
Method: X-RAY DIFFRACTION Resolution:3.13 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: ANP, MG, NH4 |
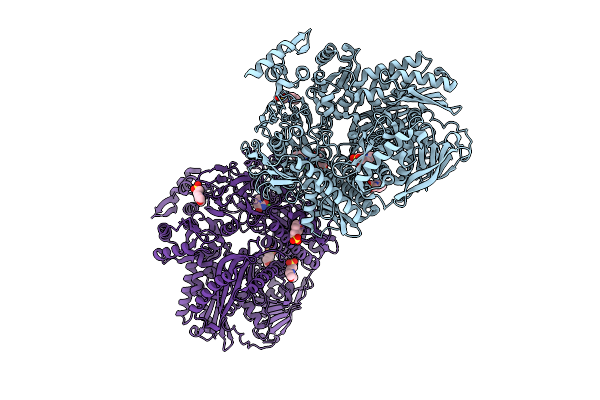 |
Crystal Structure Of N-Methylhydantoinase In Complex With 1-Methylimidazolidine-2,4-Dione, C2221 Form
Organism: Glutamicibacter protophormiae
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: CA, A1BC1, NH4, MES, SO4 |
 |
Organism: Glutamicibacter protophormiae
Method: X-RAY DIFFRACTION Resolution:2.58 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: ANP, ZN, NH4, ACT |
 |
Crystal Structure Of N-Methylhydantoinase In Complex With 1-Methylimidazolidine-2,4-Dione, C-Terminal Residues Visible
Organism: Glutamicibacter protophormiae
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: CA, A1BC1, BTB, NH4, SO4 |
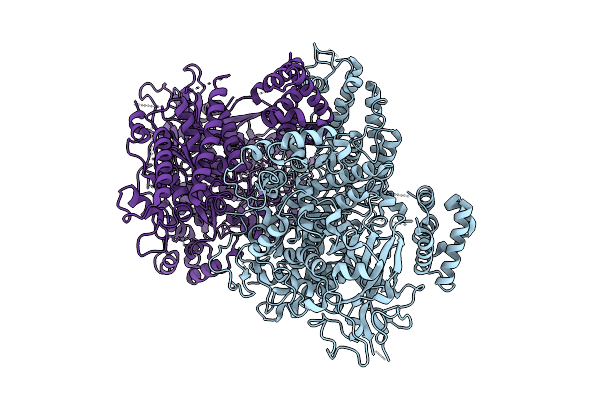 |
Single Stranded Dna-Binding Protein (Icp8) From Herpes Simplex Virus-1, Apo Form. Mutations: C254S, C455S, K769A, E770A
Organism: Human alphaherpesvirus 1
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN Ligands: ZN |
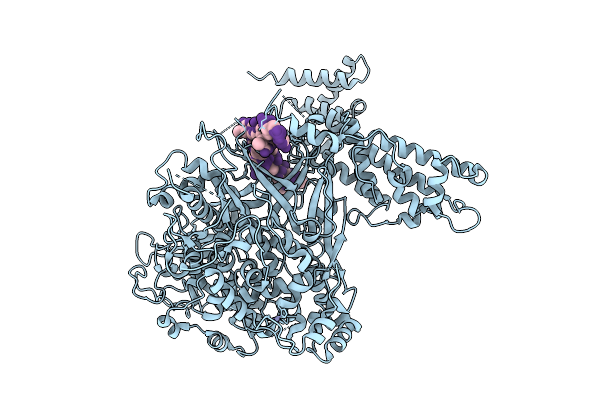 |
Poly Dt Bound Form Of Single Stranded Dna-Binding Protein(Icp8) From Herpes Simplex Virus-1. Mutations: C254S, C455S
Organism: Human alphaherpesvirus 1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN/DNA Ligands: ZN |
 |
Poly Da Bound Form Of Single Stranded Dna-Binding Protein(Icp8) From Herpes Simplex Virus-1. Mutations: C254S, C455S
Organism: Human alphaherpesvirus 1, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN/DNA |
 |
Single Stranded Dna-Binding Protein (Icp8) From Herpes Simplex Virus-1, Apo Form.Mutations: K166A, E167A, C254S, C455S
Organism: Human alphaherpesvirus 1
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN Ligands: ZN, SO4, GOL |
 |
Organism: Yersinia phage yera41
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: DNA BINDING PROTEIN |
 |
Organism: Plasmodium falciparum, Aequorea victoria
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: TRANSPORT PROTEIN Ligands: A1IVP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: ZN, A1AID |
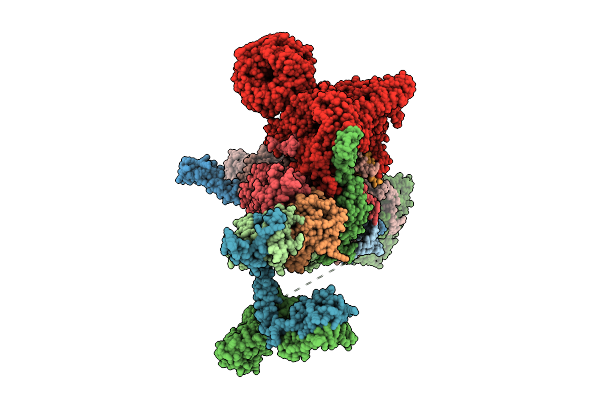 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Proximal Dhx15 State)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
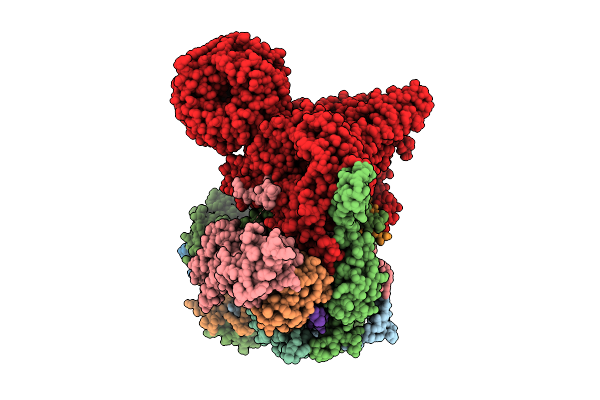 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Core)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
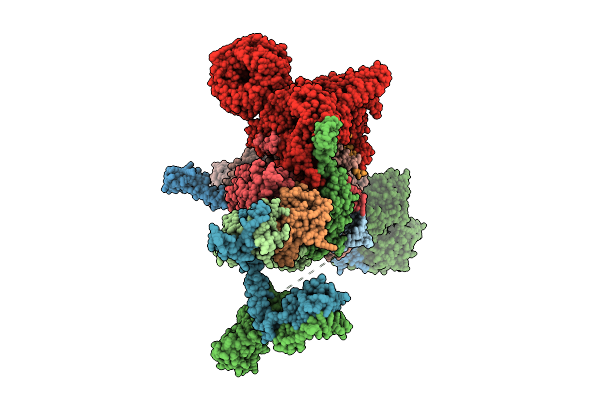 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Distal Dhx15 State)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
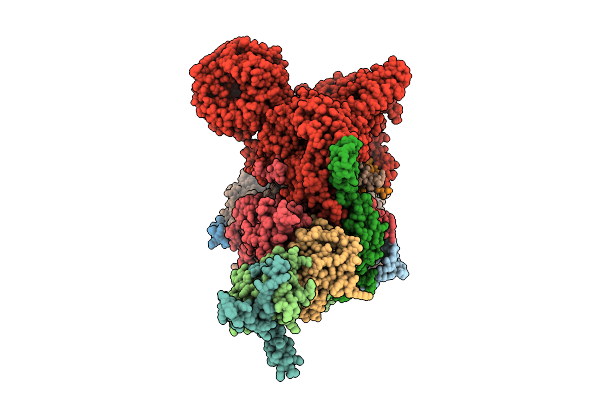 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Sf3A State 1)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |

