Search Count: 64,474
All
Selected
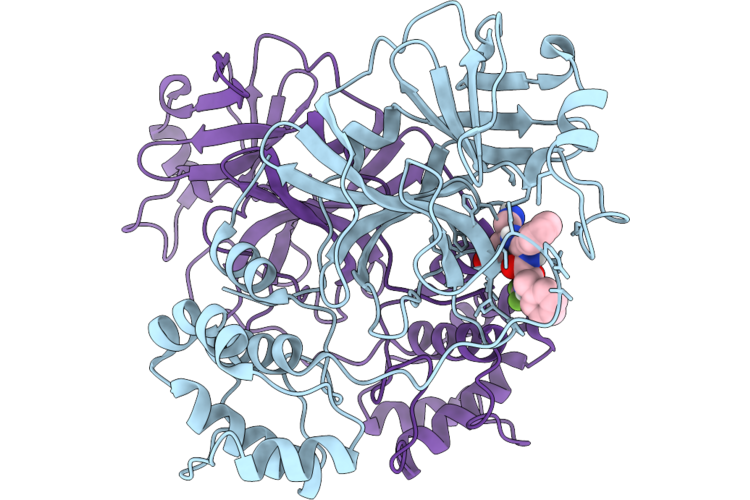 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-04-22 Classification: VIRAL PROTEIN Ligands: A1DAK |
 |
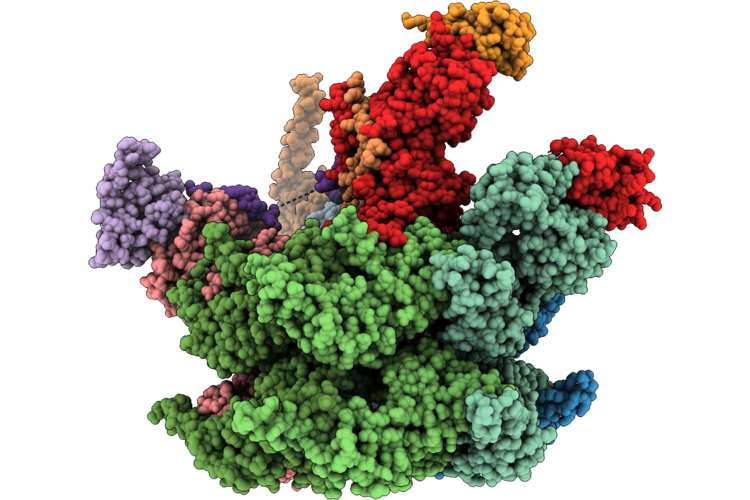
Cryo-Em Structure Of Substrate Engaged P97-Ufd1-Npl4-Faf1 Complex (Npl4 Focused)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ZN |
 |
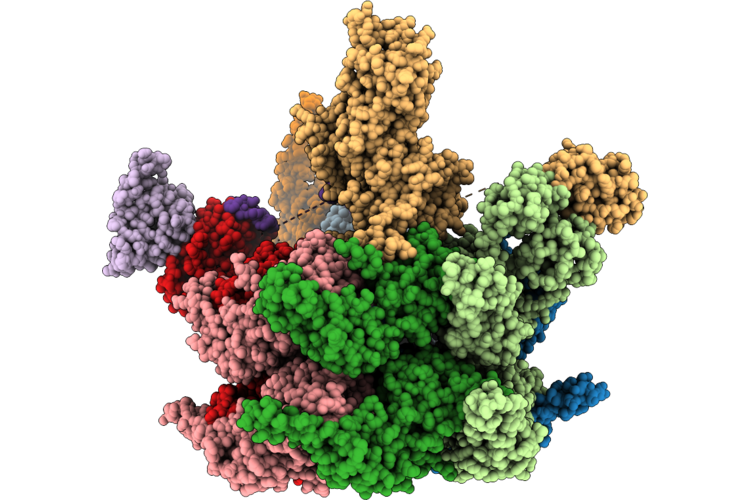
Cryo-Em Structure Of Substrate Engaged P97-Ufd1-Npl4-Faf1 Complex (Motor Focused)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ZN, ATP, ADP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.85 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ZN, ATP, ADP |
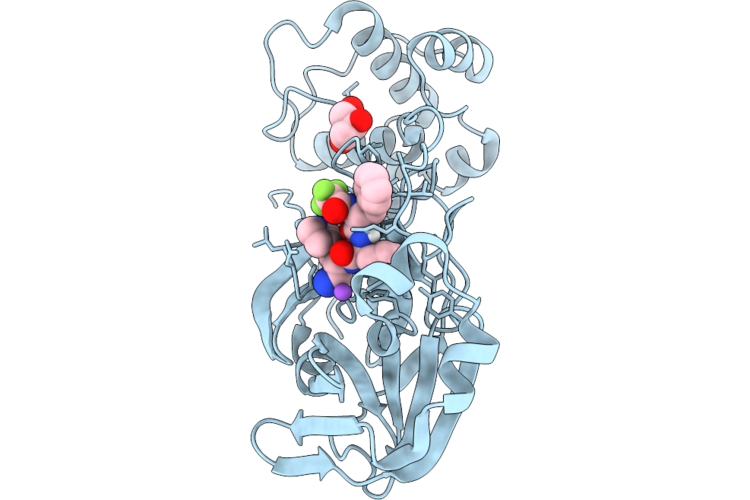 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-04-22 Classification: VIRAL PROTEIN Ligands: GOL, A1DA8, NA |
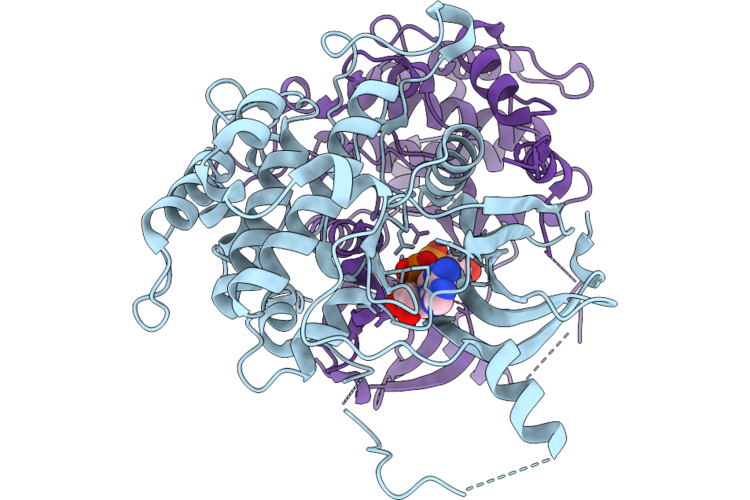 |
Crystal Structure Of Serine/Threonine-Protein Kinase (Aek1) From Trypanosoma Cruzi In Complex With Amp
Organism: Trypanosoma cruzi
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: AMP |
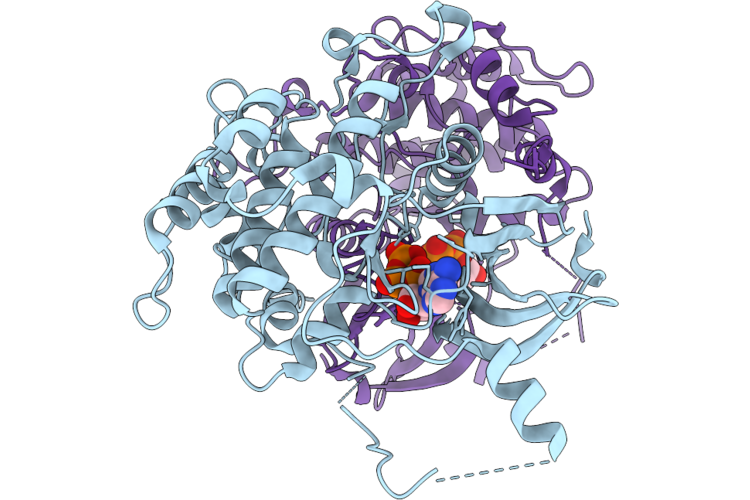 |
Crystal Structure Of Serine/Threonine-Protein Kinase (Aek1) From Trypanosoma Cruzi In Complex With Atp
Organism: Trypanosoma cruzi
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: ATP |
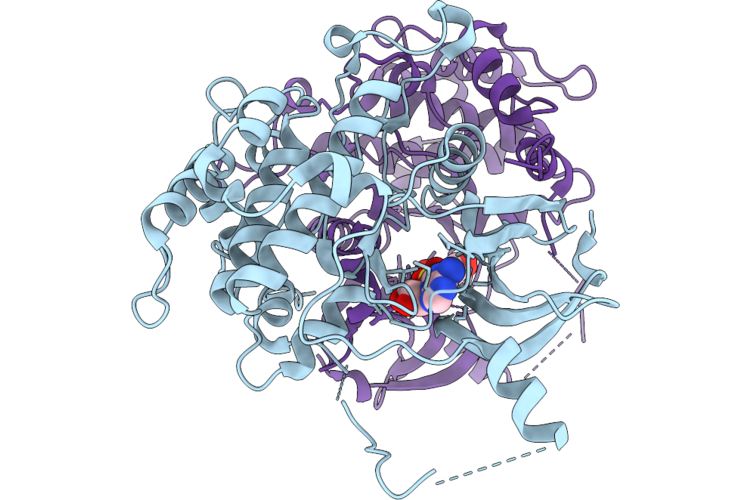 |
Crystal Structure Of Serine/Threonine-Protein Kinase (Aek1) From Trypanosoma Cruzi In Complex With Lms
Organism: Trypanosoma cruzi
Method: X-RAY DIFFRACTION Resolution:2.82 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: LMS |
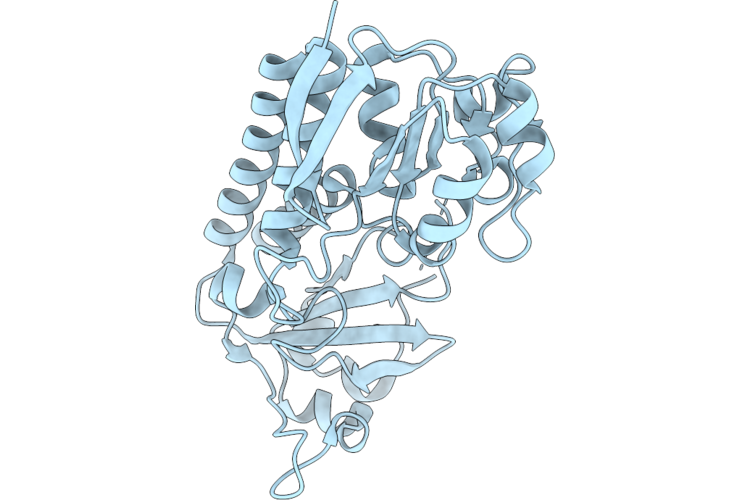 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus In The Apo-Form
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN |
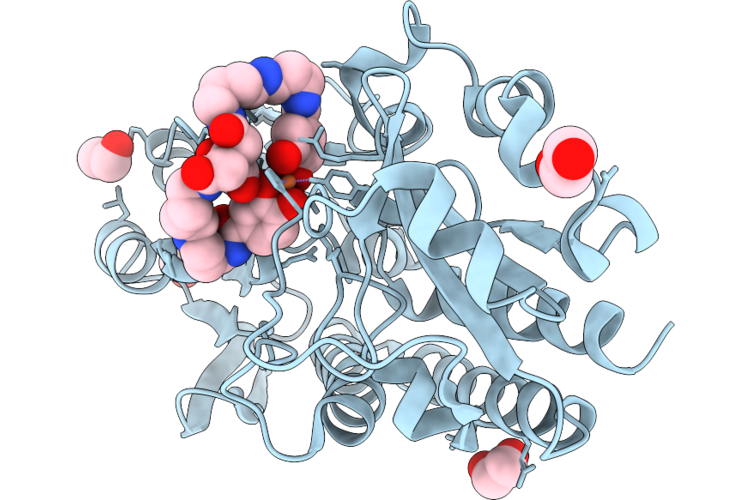 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus Complexed With Ferric Petrobactin
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN Ligands: F8W, FE, EDO |
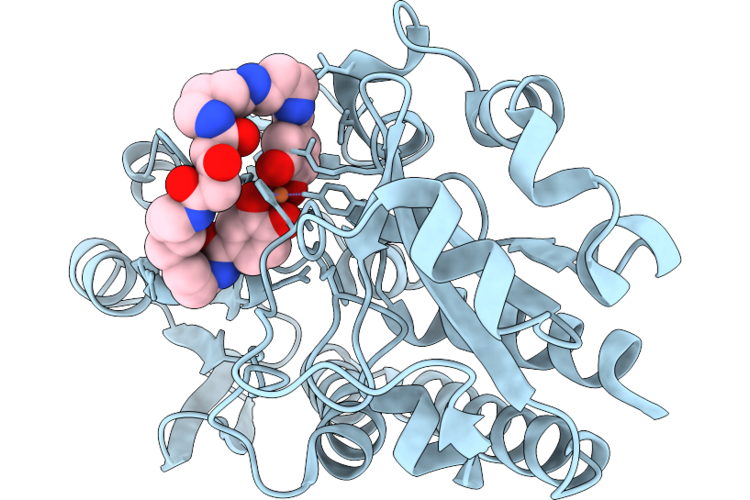 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus Complexed With Ferric Petrobactin Photoproduct, Fepbv
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN Ligands: A1MDH, FE |
 |
Crystal Structure Of The Petrobactin-Binding Protein Fatb From Bacillus Cereus Complexed With Ferric Siderophore Mimic, Fe(3,4-Dhb)2
Organism: Bacillus cereus atcc 14579
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-04-22 Classification: METAL TRANSPORT PROTEIN Ligands: DHB, FE, EDO, IPA |
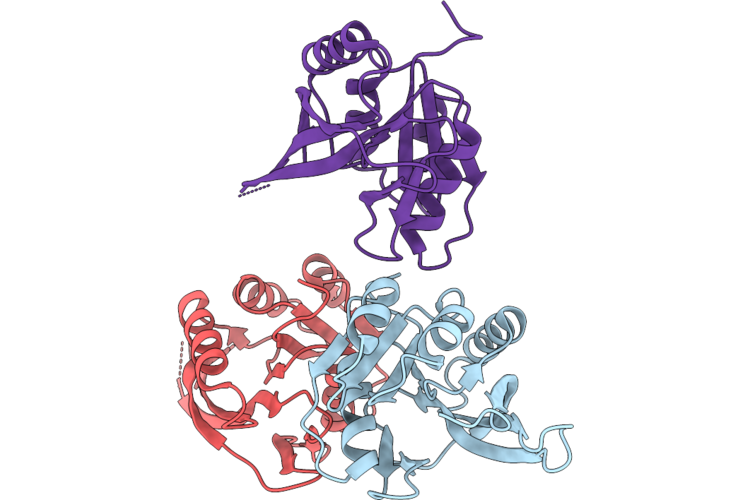 |
Organism: Bacillus cereus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-22 Classification: TRANSFERASE |
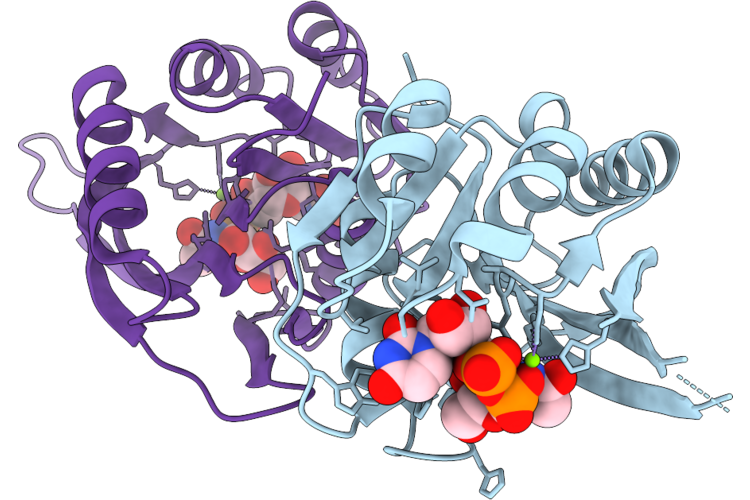 |
Crystal Structure Of Bacillus Cereus Gmar In Complex With Udp-Glcnac And Mg2+
Organism: Bacillus cereus
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: UD1, MG |
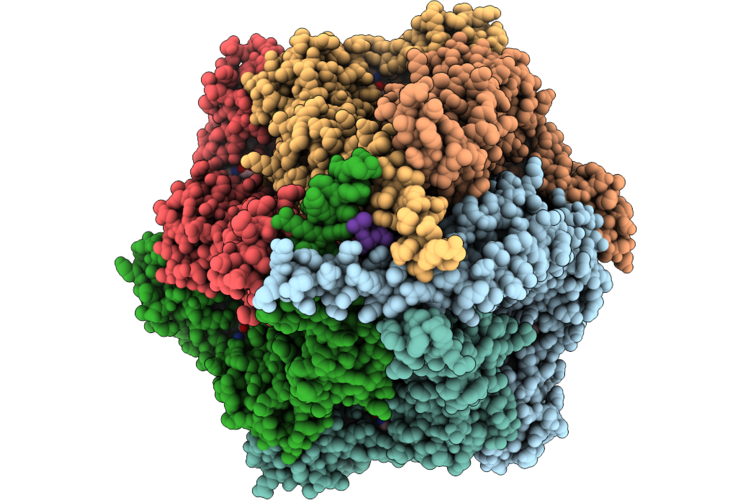 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.22 Å Release Date: 2026-04-22 Classification: STRUCTURAL PROTEIN Ligands: AGS, ADP, MG |
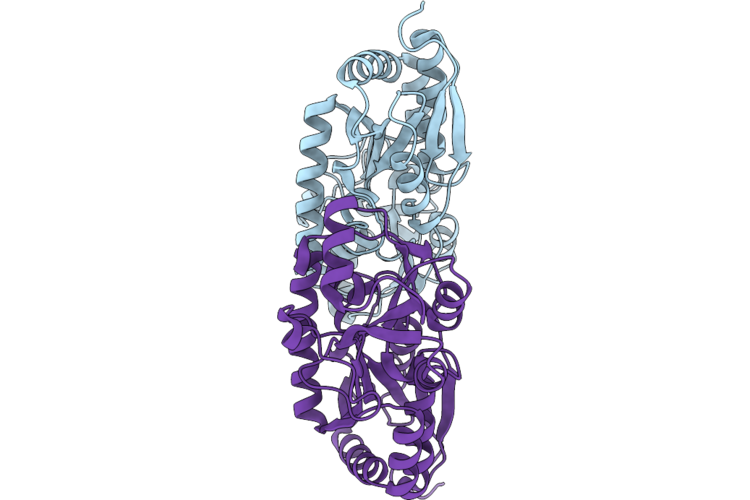 |
Crystal Structure Of A Tctc Solute Binding Protein From Vibrio Sp. C42, No Ligand
Organism: Vibrio atlanticus
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING |
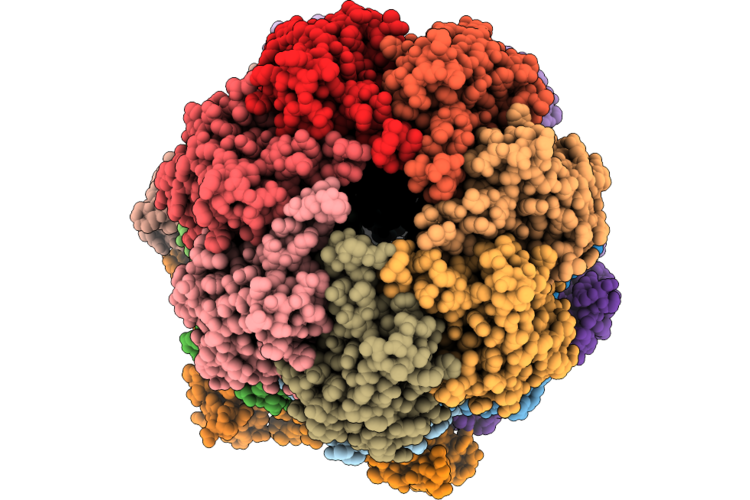 |
Organism: Enterobacter
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-22 Classification: CHAPERONE Ligands: ATP |
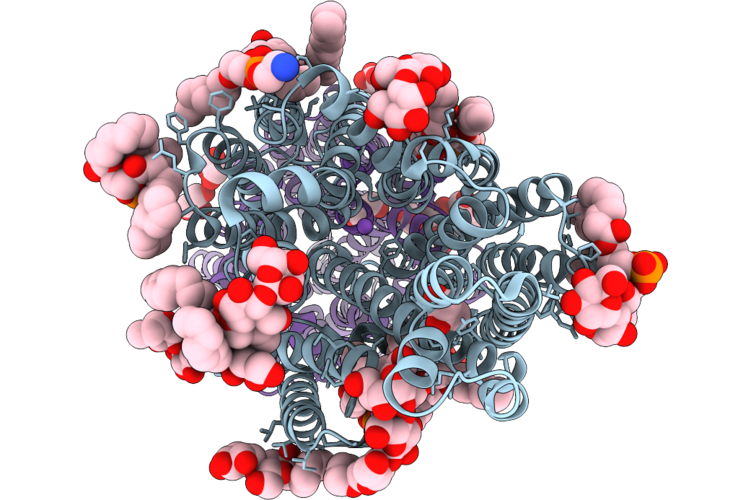 |
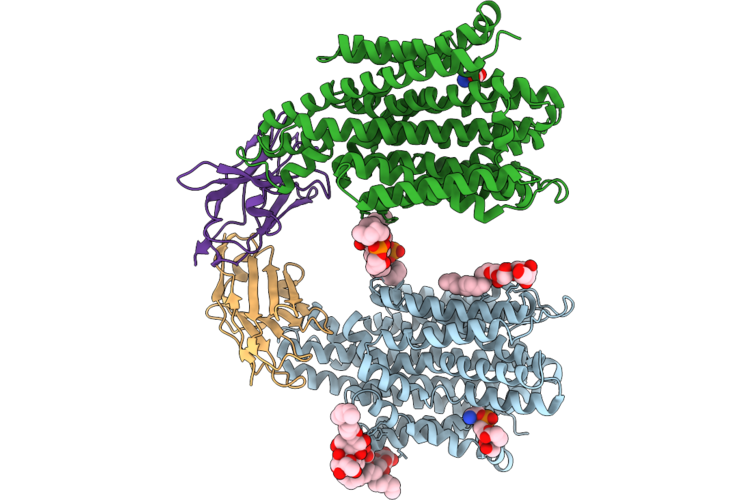
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Outward Open State
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: PGT, LMU, PTY, 3PH, GOL, NA |
 |
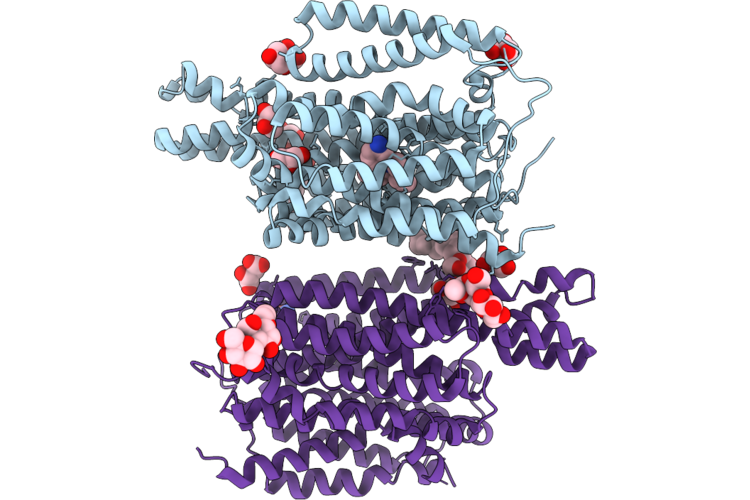
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Inward Open State In Complex With Nanobody 89
Organism: Staphylococcus aureus, Lama glama
Method: X-RAY DIFFRACTION Resolution:3.32 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: PTY, PGT, LMU, OXM |
 |
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Outward Open State Bound To Ethidium
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.79 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ET, GOL, TLA, CL, LMU, OXM |

