Search Count: 2,300
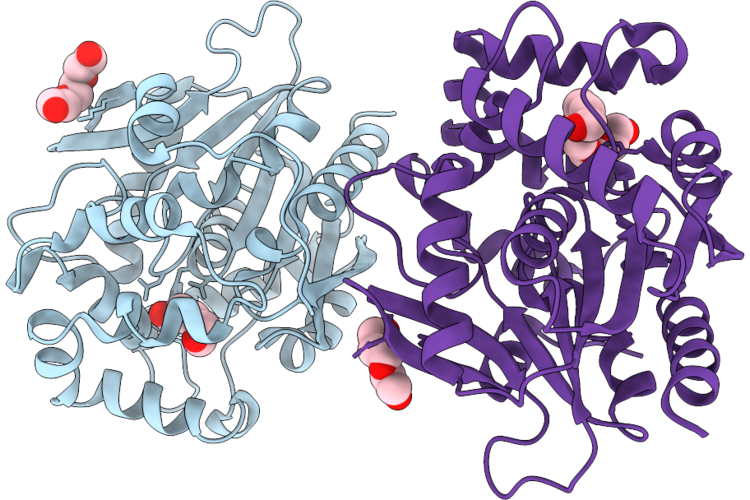 |
Structure Of An Ancestral Bifunctional Dehalogenase-Luciferase Enzyme Anc238Loc, Space Group C121
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-05-27 Classification: HYDROLASE Ligands: PGE, 1PE |
 |
Structure Of An Ancestral Bifunctional Dehalogenase-Luciferase Enzyme Anc238Loc, Space Group P21212
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-05-27 Classification: HYDROLASE Ligands: PGE |
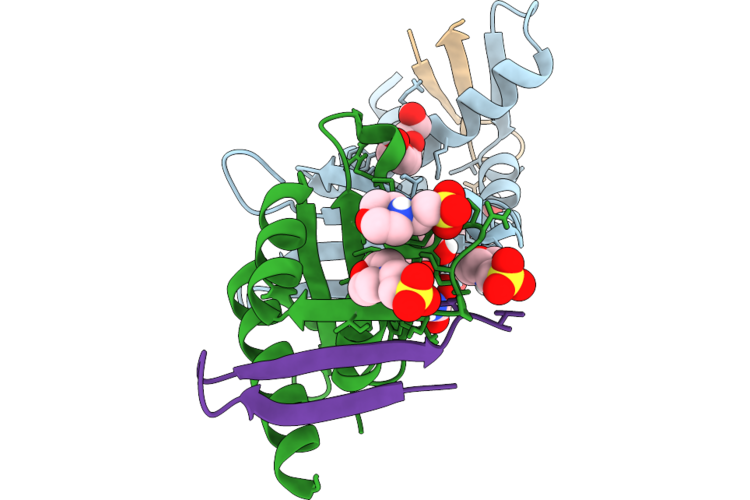 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-05-27 Classification: PROTEIN BINDING Ligands: 1PE, PGE, OXM, MES |
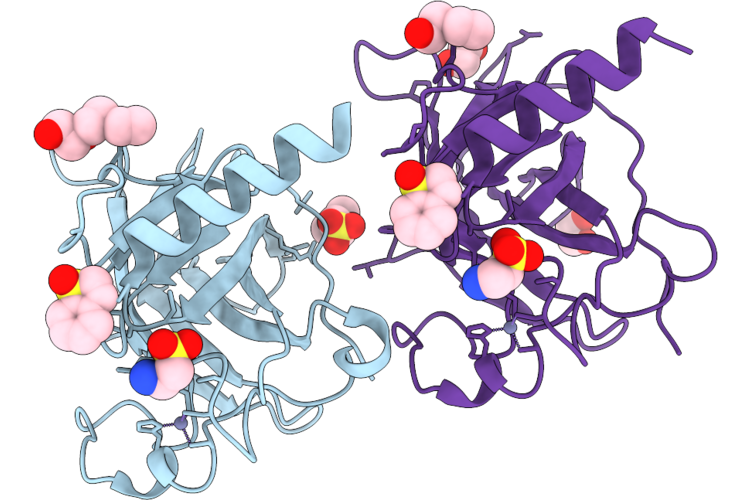 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-05-27 Classification: CELL CYCLE Ligands: A1JQW, EPE, ZN, PGE, PEG, EDO |
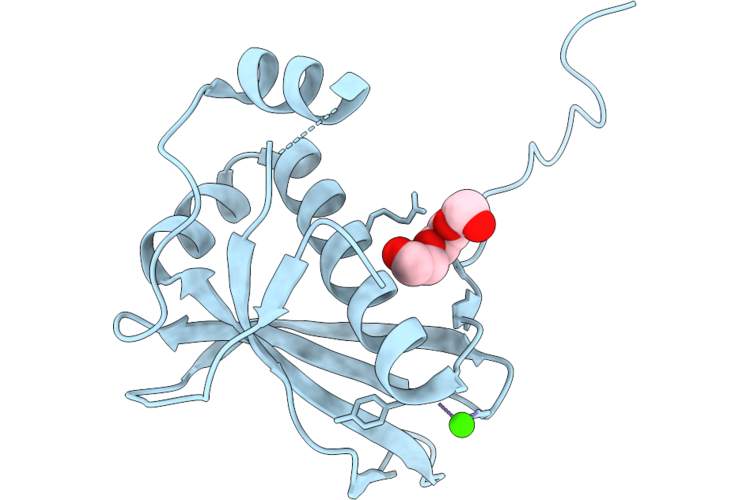 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2026-05-13 Classification: PROTEIN BINDING Ligands: PGE, CA |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2026-05-13 Classification: PROTEIN BINDING Ligands: PGE, CA |
 |
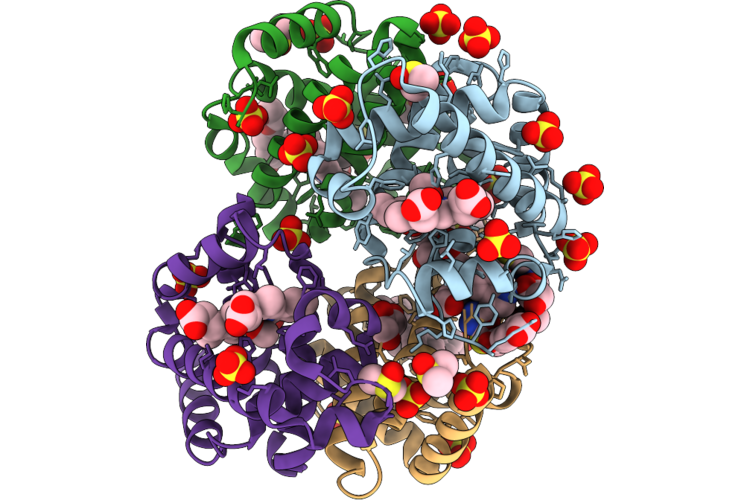
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: PGE, PG4 |
 |
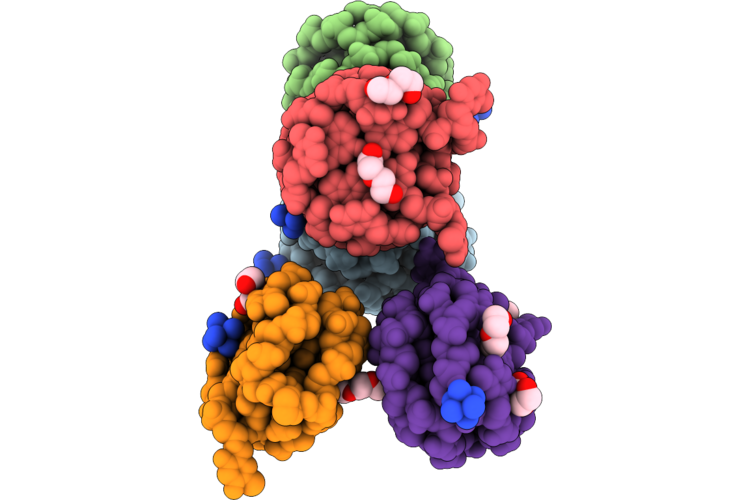
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: HEM, GOL, A1ITR, DMS, CMO, ACT, SO4, PGE |
 |
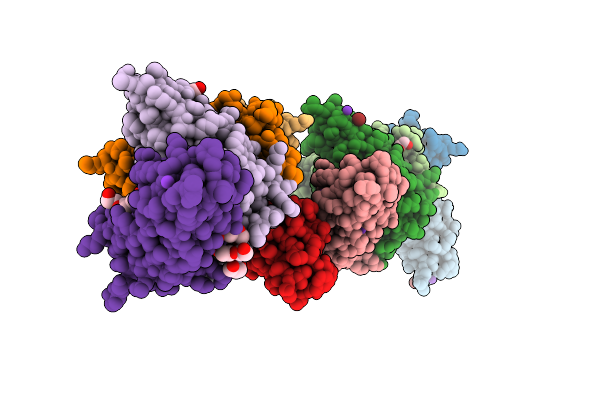
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2026-05-06 Classification: DNA Ligands: K, NCO, PEG, PG4, PGE, MES |
 |
Crystal Structure Of The Third Immunoglobulin-Like Domain Of Human Muscle-Specific Kinase
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-15 Classification: SIGNALING PROTEIN Ligands: BR, K, PEG, PG4, PGE |
 |
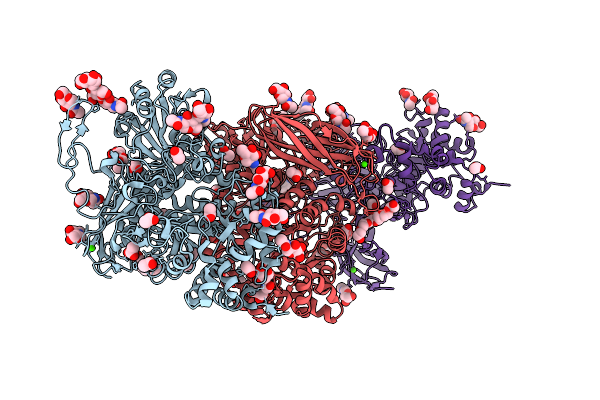
Long-Wavelength Sad Crystal Structure Of The Third Immunoglobulin-Like Domain Of Human Muscle-Specific Kinase (Musk)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-15 Classification: SIGNALING PROTEIN Ligands: K, BR, PEG, PGE |
 |
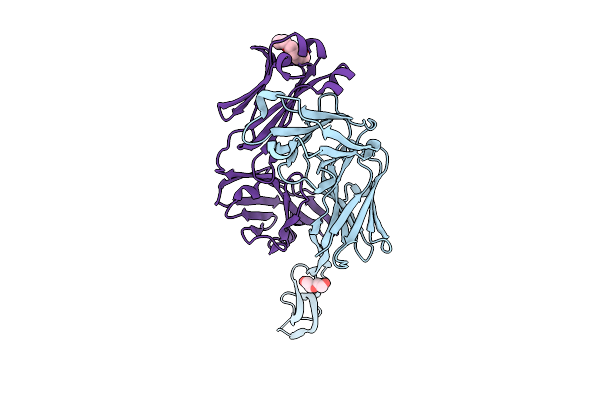
Crystal Structure Of Isoprimeverose-Producing Enzyme From Phaeoacremonium Minimum
Organism: Rattus rattus, Phaeoacremonium minimum
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAG, ACT, PG4, CA, PGE, P6G |
 |
Organism: Bos taurus
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-04-08 Classification: IMMUNE SYSTEM Ligands: PEG, PGE |
 |
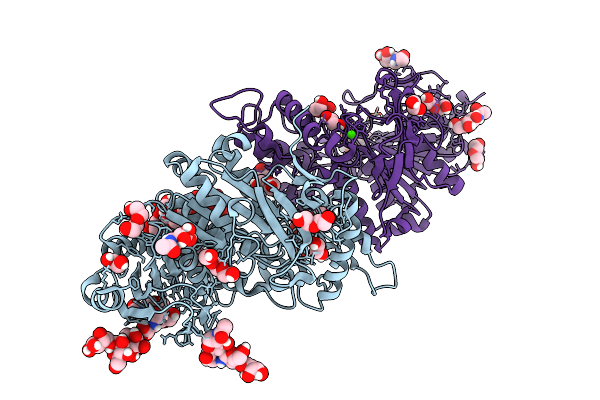
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2026-04-01 Classification: OXYGEN BINDING Ligands: HEM, PG4, EDO, SO4, PEG, TRS, OXY, PGE |
 |
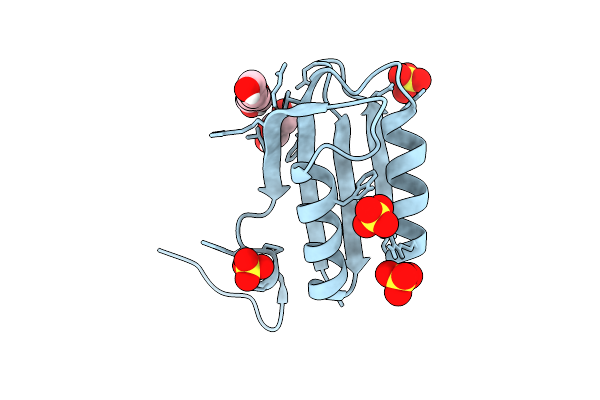
Crystal Structure Of Feruloyl Esterase From Fusarium Oxysporum G122S Variant
Organism: Fusarium oxysporum
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-01 Classification: HYDROLASE Ligands: NAG, PEG, EDO, PGE, CA |
 |
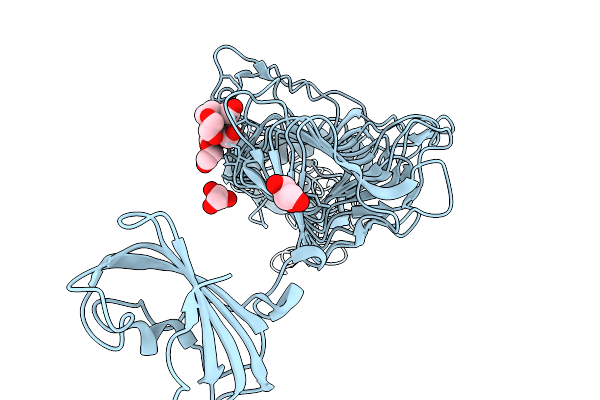
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-04-01 Classification: HYDROLASE Ligands: GOL, PGE, SO4 |
 |
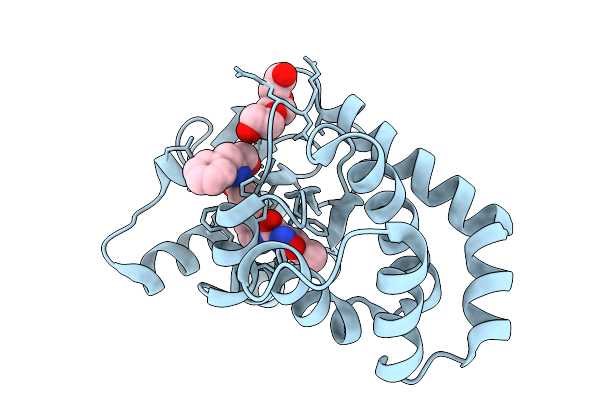
Crystal Structure Of A Tailspike Depolymerase (Belisarius_Gp86) From Acinetobacter Phage Belisarius
Organism: Acinetobacter phage
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-03-25 Classification: VIRAL PROTEIN Ligands: GOL, PGE |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJN, PGE |
 |
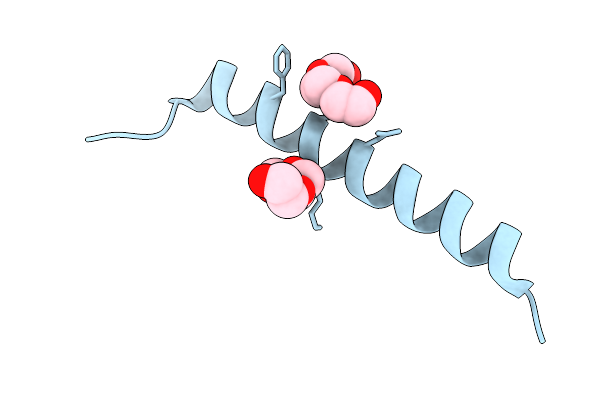
De Novo Photoenzyme Variant Able-E116C-Txt H49A-L42T (Photoable1 Precursor) In Complex With Product Formed In Crystallo After Irradiation
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-03-25 Classification: DE NOVO PROTEIN Ligands: A1JUX, A1JUW, PGE, EDO, GOL |
 |
Organism: Heloderma suspectum cinctum
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2026-03-25 Classification: HORMONE Ligands: PGE |

