Search Count: 87,025
All
Selected
 |
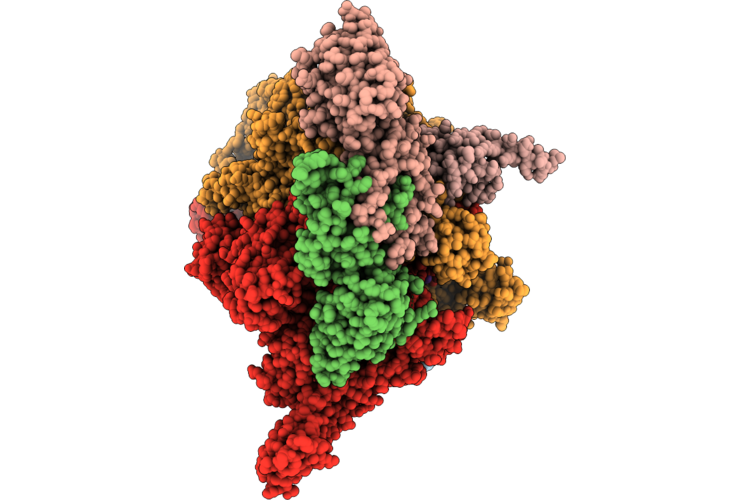
Closed Eco-Epec: Cryo-Em Structure Of Eco Rnap His-Elemental Paused Elongation Complex With A Closed Active Site (Closed Tl, Si3 And Rh-Fl)
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-29 Classification: Transcription/DNA/RNA Ligands: 1N7, MG, ZN |
 |
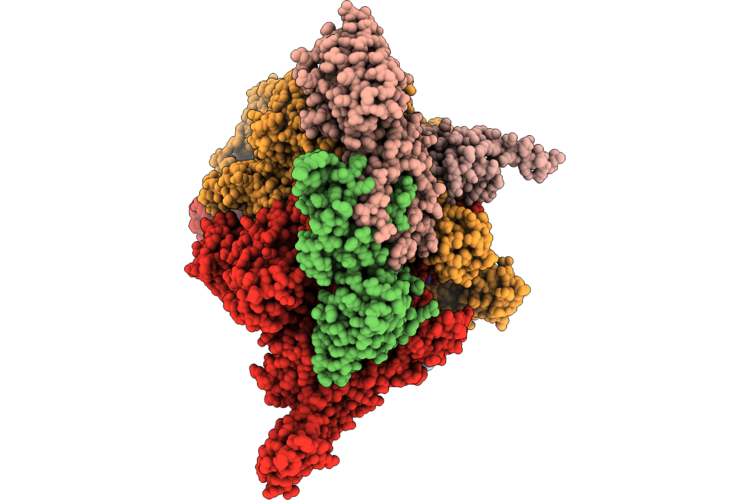
Open1 Eco-Epec: Cryo-Em Structure Of Eco Rnap His-Elemental Paused Elongation Complex With An Open Active Site (Open Tl, Si3 And Rh-Fl)
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-29 Classification: TRANSCRIPTION/DNA/RNA Ligands: 1N7, MG, ZN |
 |
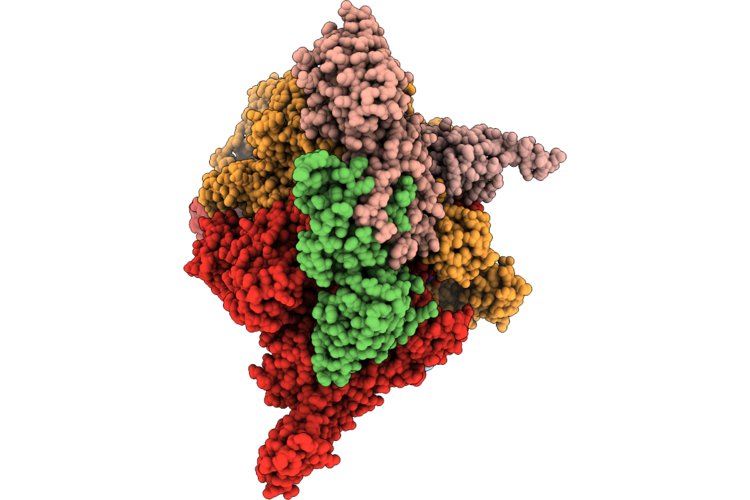
Open2 Eco-Epec: Cryo-Em Structure Of Eco Rnap His-Elemental Paused Elongation Complex With An Open Active Site (Open Tl, Si3 And Rh-Fl)
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-29 Classification: Transcription/DNA/RNA Ligands: 1N7, MG, ZN |
 |
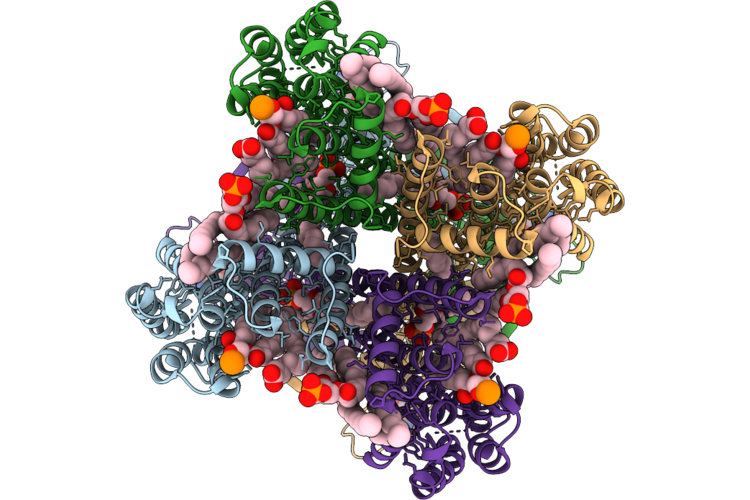
Open3 Eco-Epec: Cryo-Em Structure Of Eco Rnap His-Elemental Paused Elongation Complex With An Open Active Site (Open Tl, Si3 And Rh-Fl)
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-04-29 Classification: TRANSCRIPTION Ligands: 1N7, MG, ZN |
 |
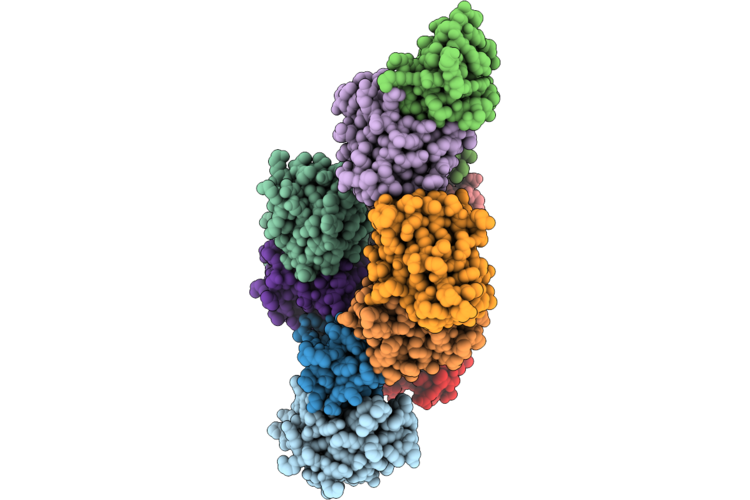
Organism: Winmispira thermophila
Method: ELECTRON MICROSCOPY Resolution:2.51 Å Release Date: 2026-04-29 Classification: TRANSPORT PROTEIN Ligands: CMP, PCW |
 |
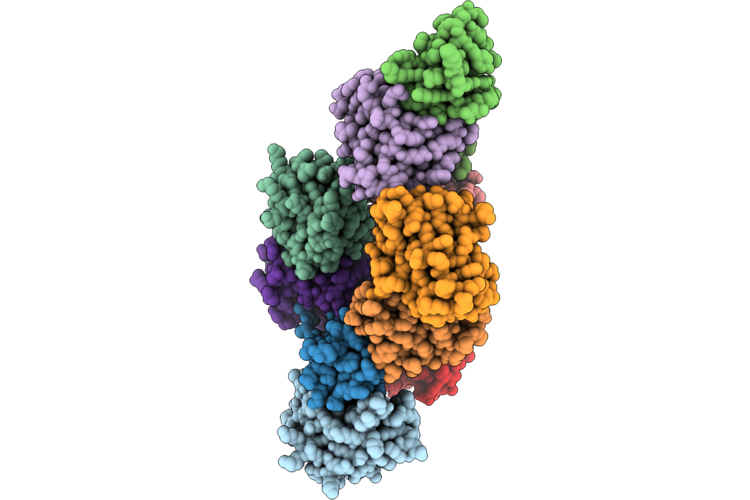
Crystal Structure Of Geocas9 Hnh Domain Bound To S78A Mutant Anti-Crispr Acriic1
Organism: Geobacillus stearothermophilus
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-29 Classification: DNA BINDING PROTEIN |
 |
Organism: Geobacillus stearothermophilus, Neisseria meningitidis
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-29 Classification: DNA BINDING PROTEIN |
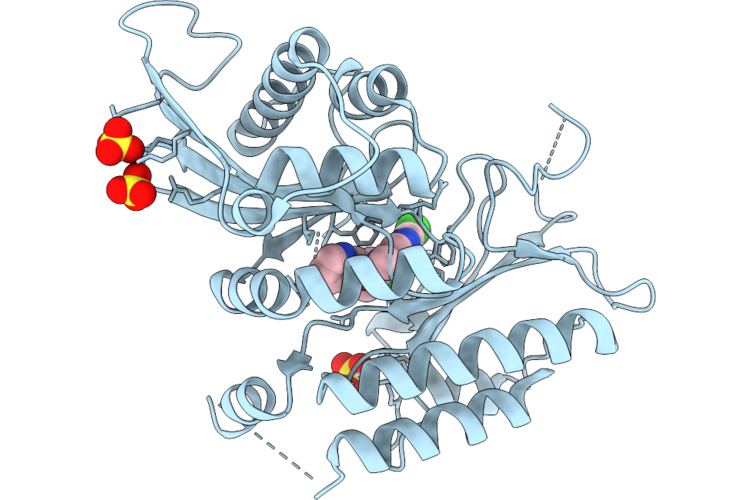 |
Structural Basis For High-Affinity Inhibitor Binding To Lipid Kinases Pip4K2A And Pip4K2B
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.79 Å Release Date: 2026-04-29 Classification: TRANSFERASE Ligands: SO4, A1C8V |
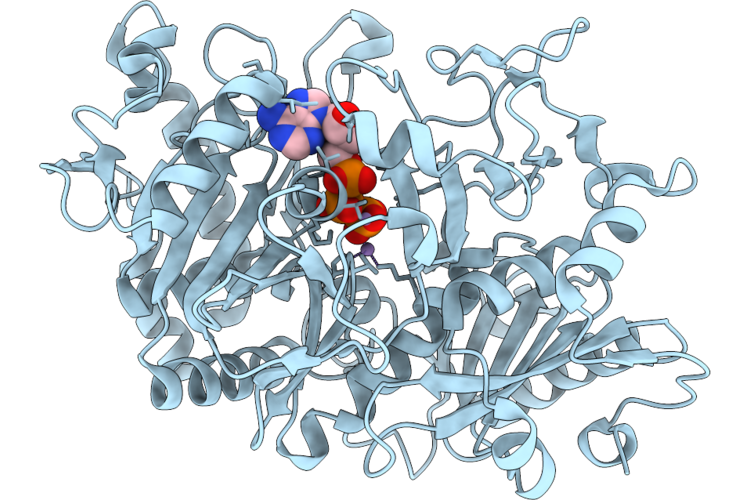 |
Structure Of Escherichia Coli Phosphoenolpyruvate Carboxykinase With Bound Atp, Mg2+, And Mn2+
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-04-29 Classification: LYASE Ligands: ATP, MN, MG |
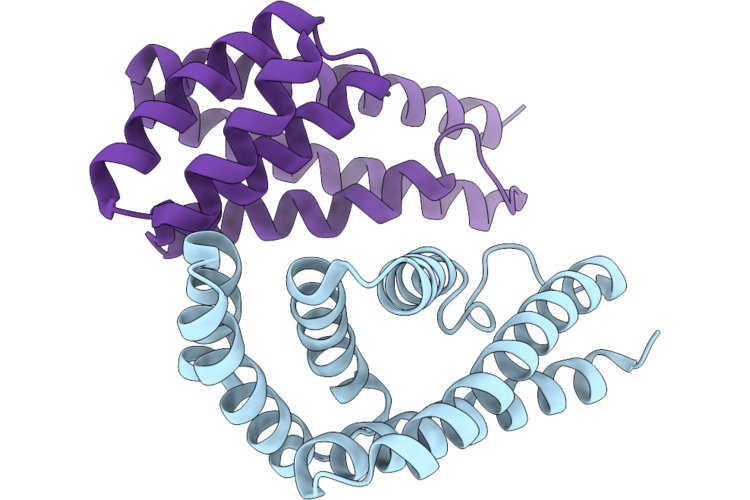 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2026-04-29 Classification: DE NOVO PROTEIN |
 |
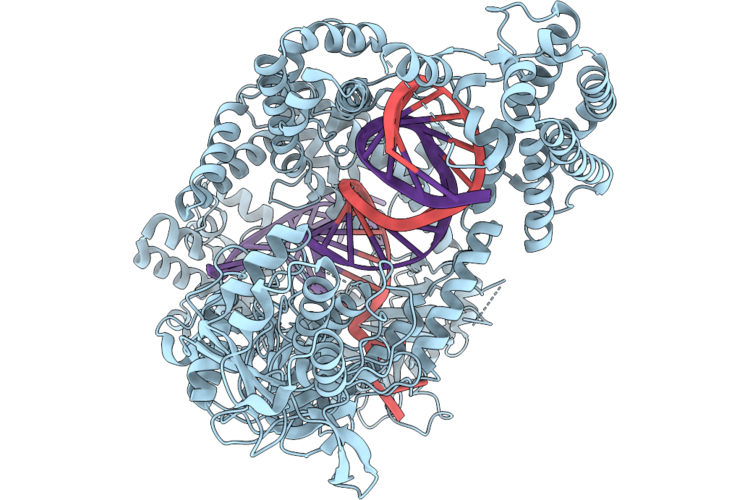
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-04-29 Classification: DE NOVO PROTEIN Ligands: A1DD0 |
 |
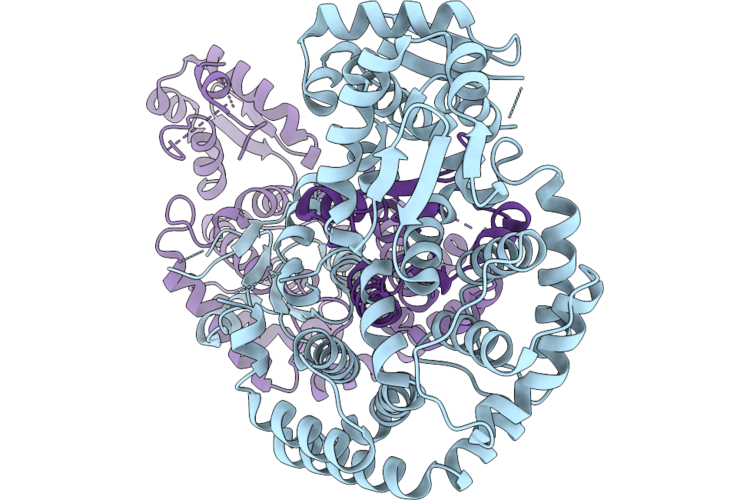
Organism: Acidaminococcus sp. bv3l6, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.17 Å Release Date: 2026-04-29 Classification: DNA BINDING PROTEIN |
 |
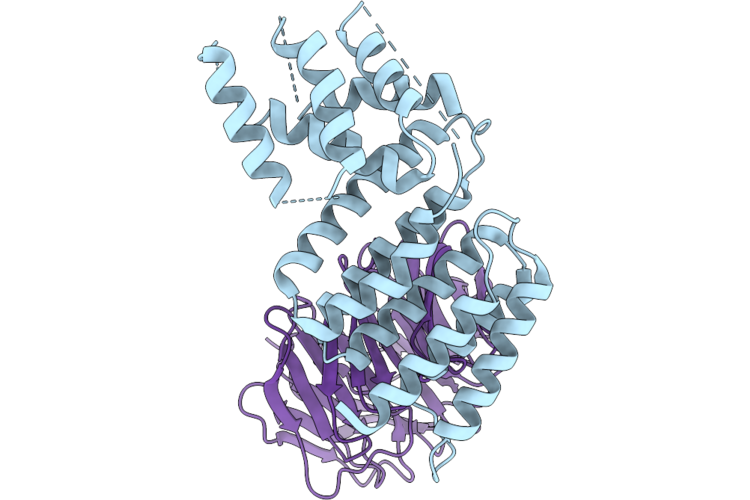
Organism: Enterococcus faecalis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-29 Classification: TRANSCRIPTION |
 |
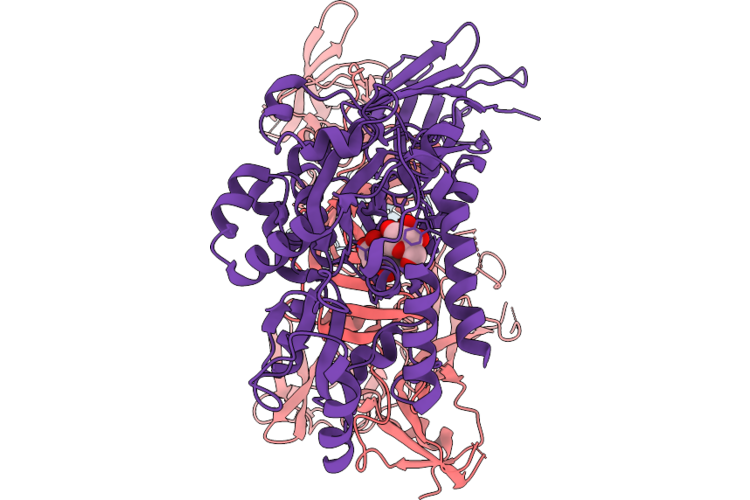
Stabilized Complex Of Chlamydia Trachomatic Efector Ct622 In Complex With Human Wd40 Domain Of Atg16L1
Organism: Escherichia coli o157:h7, Chlamydia trachomatis, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: PROTEIN TRANSPORT |
 |
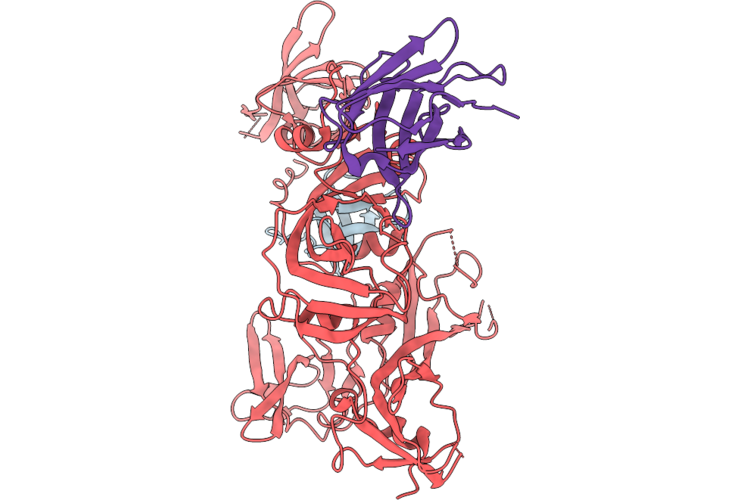
Organism: Homo sapiens, Synthetic construct, Escherichia coli
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: SIGNALING PROTEIN |
 |
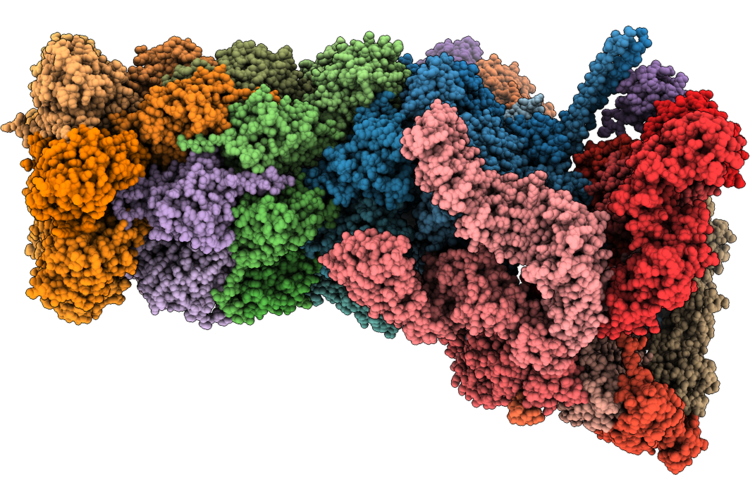
Cryoem Structure Of The Themis:Grb2 Complex With Bound Promacrobody 256, Local Refinement
Organism: Homo sapiens, Synthetic construct, Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-04-29 Classification: SIGNALING PROTEIN |
 |
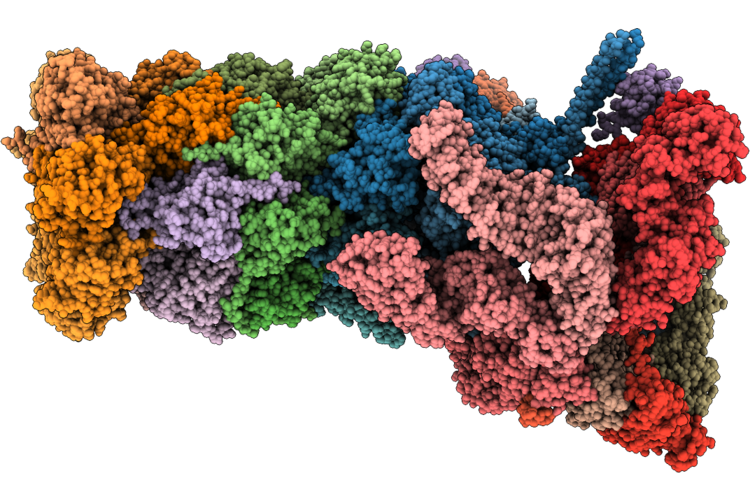
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ea1.0
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ea1.1
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
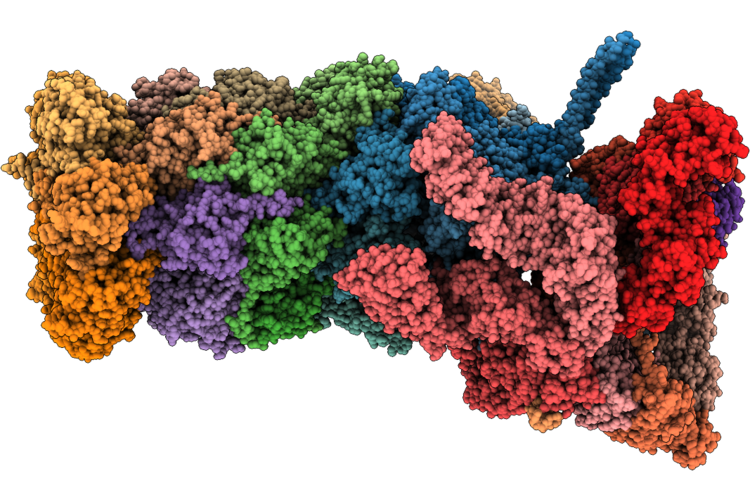 |
Structure Of Substrate-Engaged 26S Proteasome Rp-Cp Subcomplex In State Ea1.2
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |
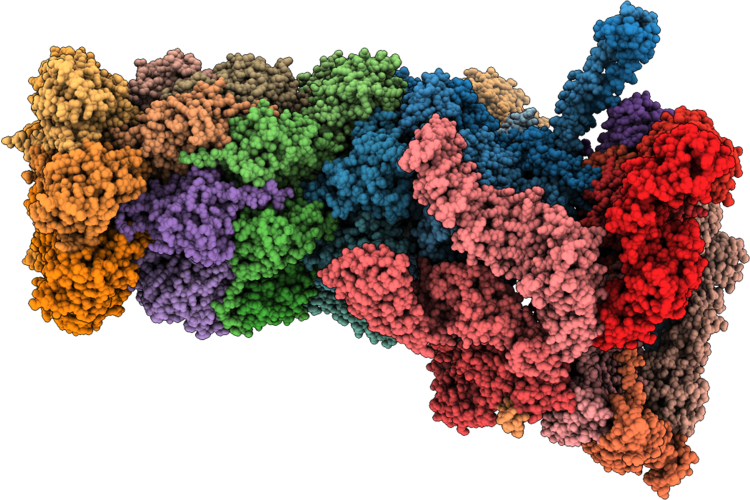 |
Structure Of Substrate-Engaged Human 26S Proteasome Rp-Cp Subcomplex In State Ea2.1
Organism: Homo sapiens, Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: HYDROLASE Ligands: ATP, MG, ADP, ZN |

