Search Count: 109
All
Selected
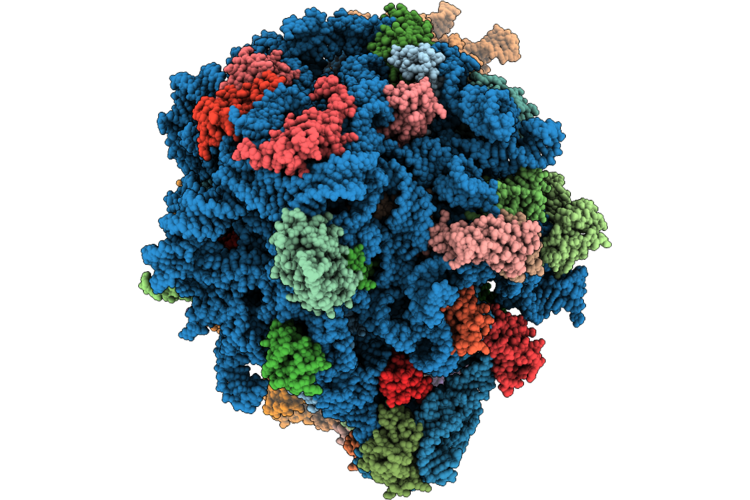 |
Organism: Francisella tularensis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: RIBOSOME Ligands: MG, ZN |
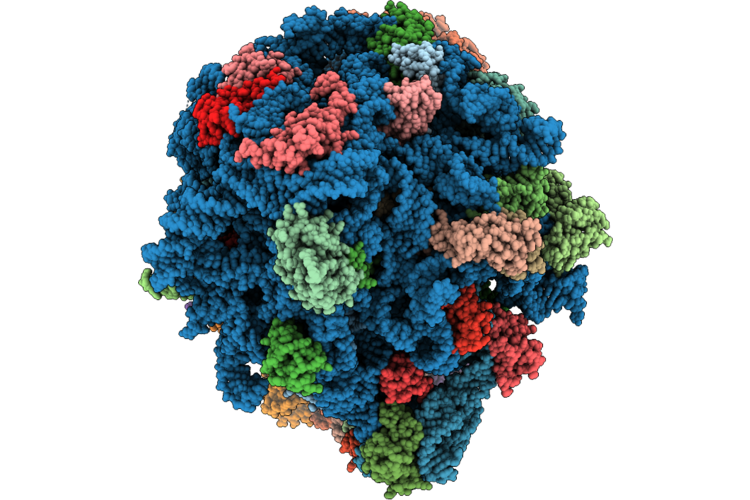 |
Cryo-Em Structure Of The 70S Ribosome From Francisella Tularensis Bound To A Hibernation-Promoting Factor
Organism: Francisella tularensis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: RIBOSOME Ligands: MG, ZN |
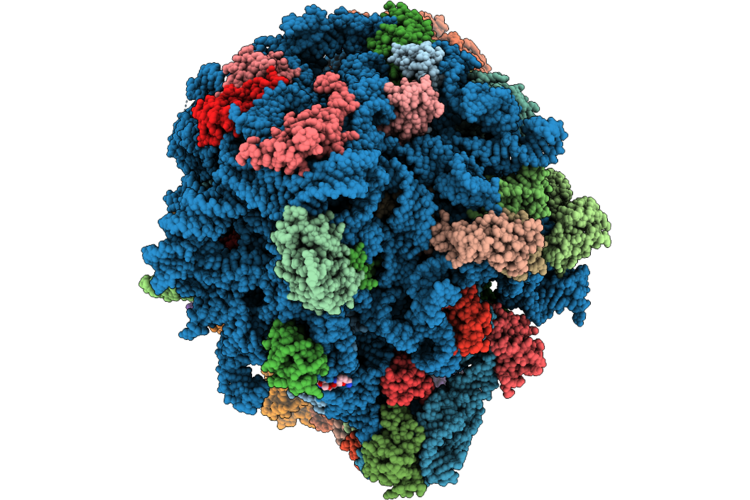 |
Cryo-Em Structure Of The 70S Ribosome From Francisella Tularensis Bound To Antibiotics Chloramphenicol And Gentamicin
Organism: Francisella tularensis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: RIBOSOME Ligands: LLL, MG, CLM, ZN |
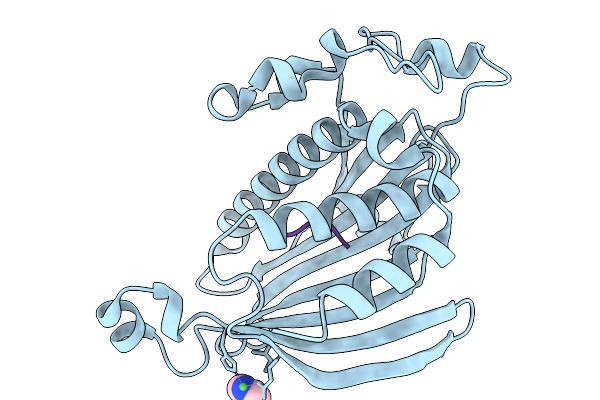 |
Organism: Homo sapiens, Moorena producens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-02-04 Classification: LIPID TRANSPORT Ligands: IMD, NI |
 |
Organism: Borreliella burgdorferi
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: MN, FE, AMP |
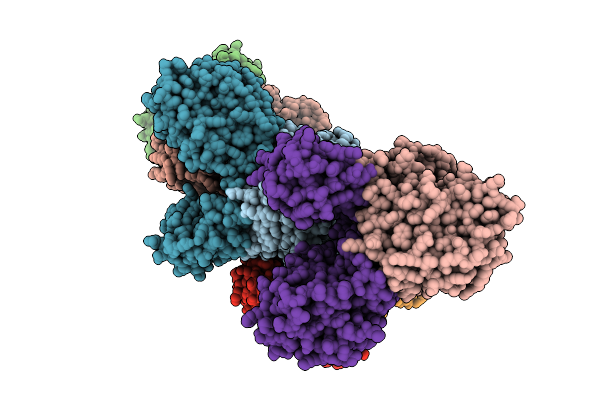 |
Organism: Borreliella burgdorferi
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: MN, FE, AMP |
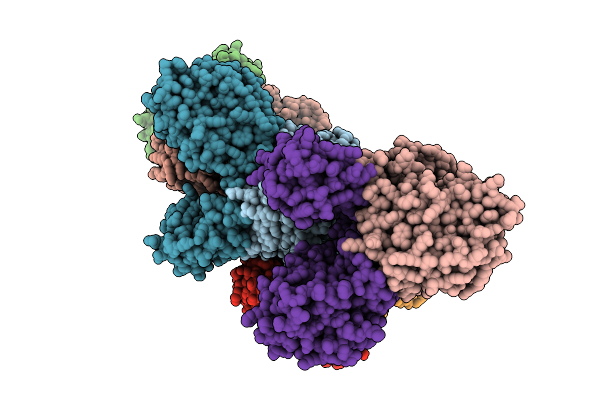 |
Organism: Borreliella burgdorferi
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: MN, FE, AMP |
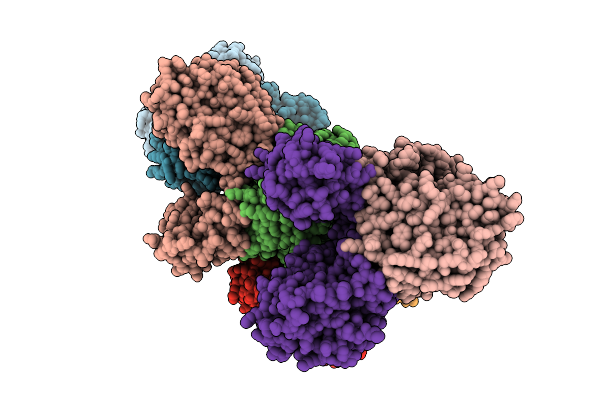 |
Organism: Borreliella burgdorferi
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: MN, FE, A1JDM |
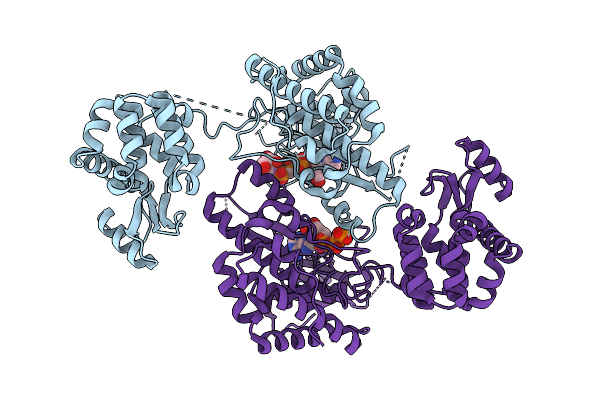 |
Crystal Structure Of Phosphatidyl Inositol 4-Kinase Ii Beta In Complex With Hh5129
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2025-12-03 Classification: TRANSFERASE Ligands: A1IVA |
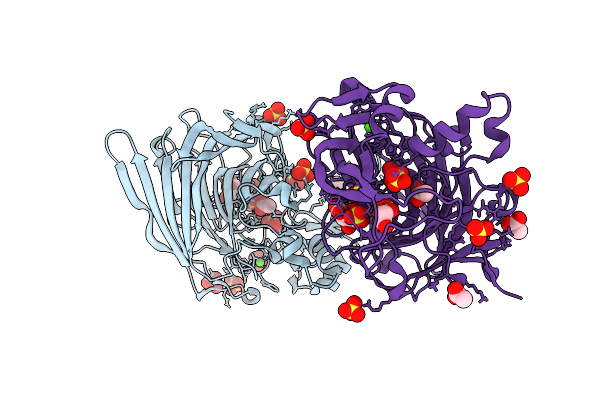 |
Structure Of Heparinase I From Bacteroides Eggerthii In Complex With Calcium Cofactor And Delta-Ua(1-4)Glc 3, 6, N-Sulphated Disaccharide
Organism: Bacteroides eggerthii
Method: X-RAY DIFFRACTION Resolution:1.72 Å Release Date: 2025-10-08 Classification: LYASE Ligands: ACT, SO4, CA |
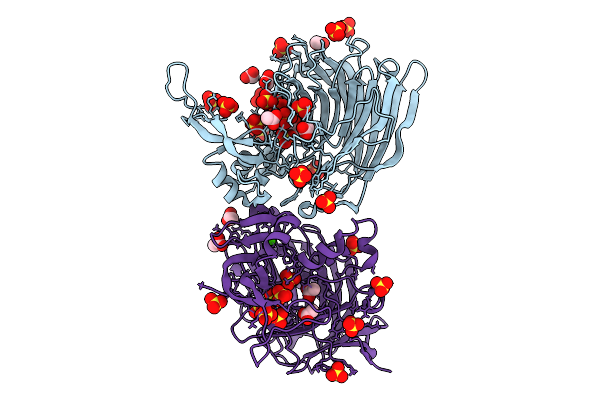 |
Structure Of Heparinase I From Bacteroides Eggerthii In Complex With Calcium Cofactor
Organism: Bacteroides eggerthii
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2025-09-24 Classification: LYASE Ligands: CA, ACT, SO4 |
 |
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: IMP |
 |
Crystal Structure Of M. Smegmatis Gmp Reductase With Xmp* Intermidiate In Complex With Nadp+ And Imp.
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: IMP, NAP |
 |
Crystal Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp And Atp.
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: ATP, 5GP |
 |
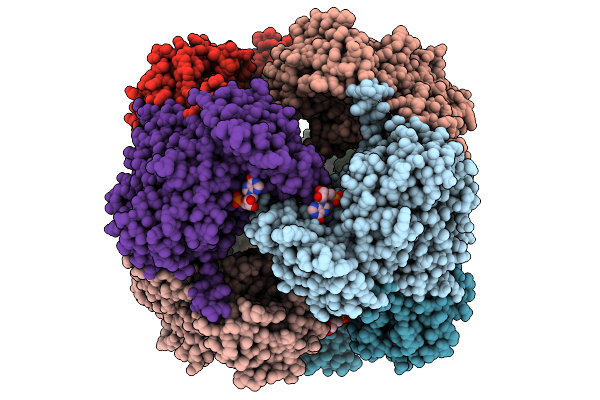
Crystal Structure Of M. Smegmatis Gmp Reductase In Complex With Imp And Atp.
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: IMP, ATP |
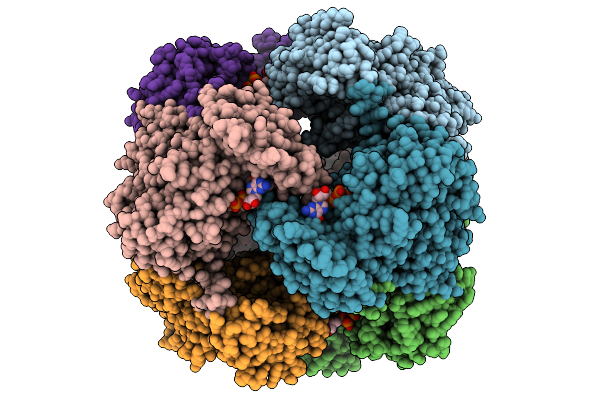 |
Crystal Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp And Gtp.
Organism: Mycolicibacterium smegmatis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: 5GP, GTP |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp At Ph 6.6, Extended Conformation.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: 5GP |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp At Ph 6.6, Compressed Conformation.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: 5GP |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp And Gtp At Ph 6.6, Extended Conformation I.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: 5GP, GTP |
 |
Cryoem Structure Of M. Smegmatis Gmp Reductase In Complex With Gmp And Gtp At Ph 6.6, Extended Conformation Ii.
Organism: Mycolicibacterium smegmatis
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: OXIDOREDUCTASE Ligands: 5GP, GTP |

