Search Count: 4,549
All
Selected
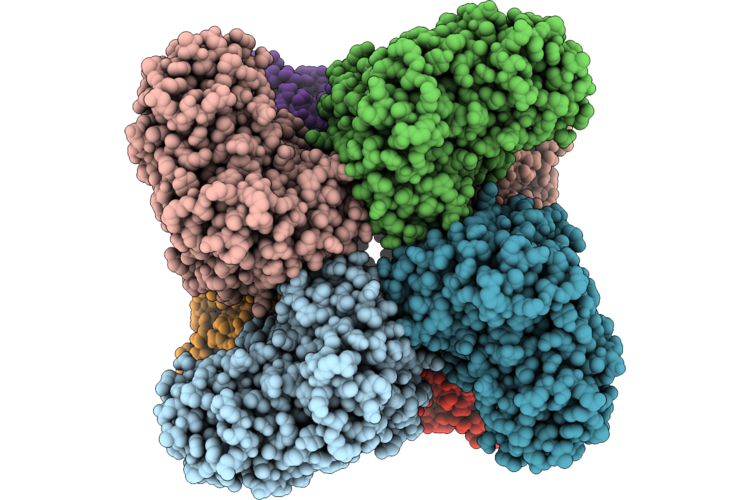 |
Cryo-Electron Microscopic Structure Of A Novel Amidohydrolase With Three Mutations
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN, 97U |
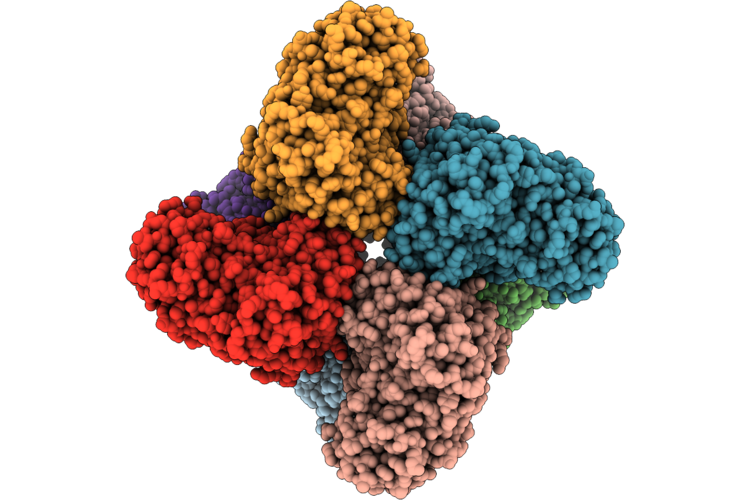 |
Cryo-Electron Microscopic Structure Of A Highly Efficient Ochratoxin Detoxification Enzyme Lladh
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN |
 |
Crystal Structure Of Afnth1:Tg-Dna Duplex Complex In A Pre-Intermediate State
Organism: Archaeoglobus fulgidus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2026-04-15 Classification: LYASE/DNA Ligands: SF4 |
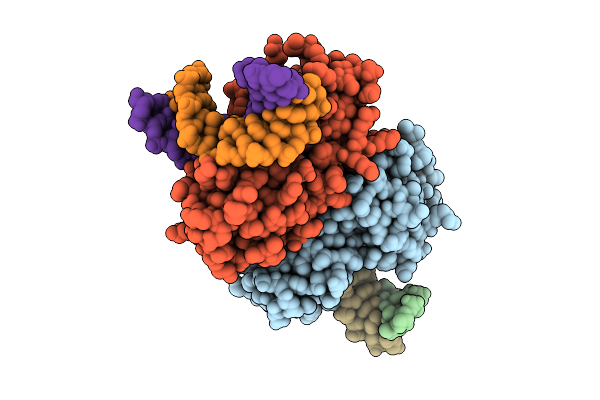 |
Crystal Structure Of Afnth1-K122A Mutant Bound To Tg-Dna Duplex Complex In An Intermediate State
Organism: Archaeoglobus fulgidus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2026-04-15 Classification: LYASE/DNA Ligands: SF4 |
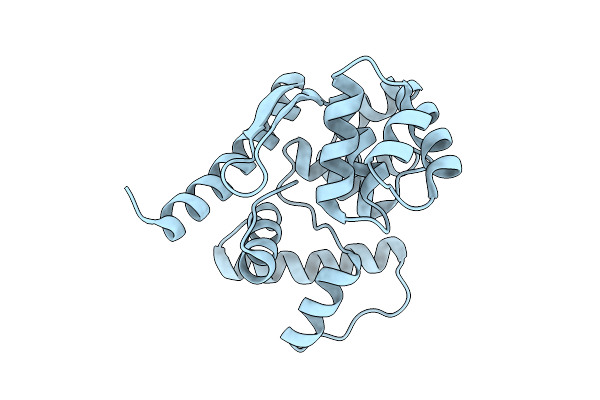 |
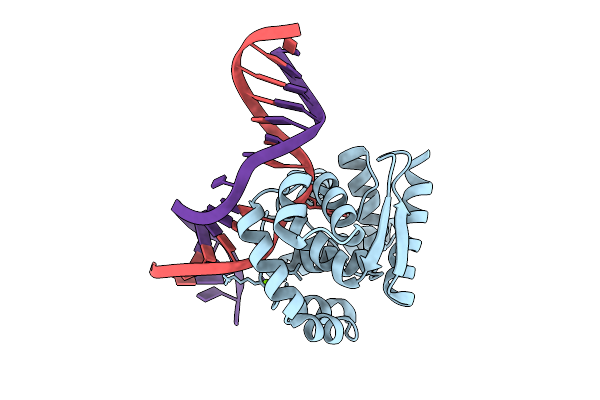
Organism: Archaeoglobus fulgidus
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2026-04-15 Classification: HYDROLASE, LYASE |
 |
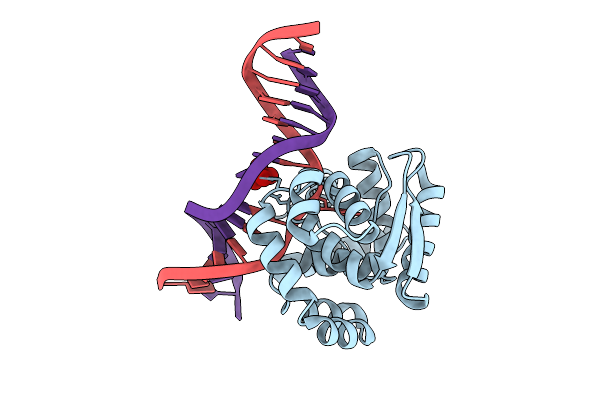
Crystal Structure Of Afogg1-D140N Mutant Bound To 8-Og Dna Duplex In An Intermediate State
Organism: Archaeoglobus fulgidus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-04-15 Classification: HYDROLASE, LYASE/DNA Ligands: MG |
 |
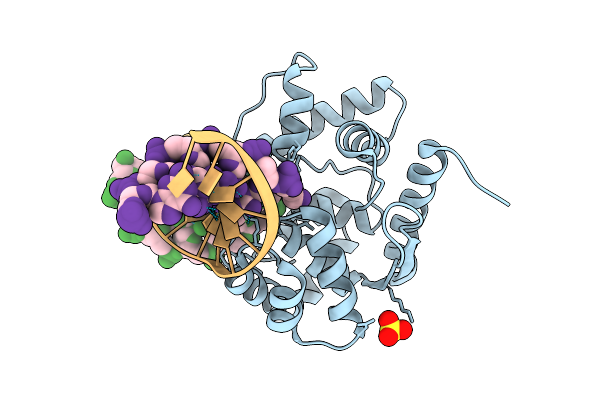
Crystal Structure Of Afogg1-K122S Mutant Bound To 8-Og Dna Duplex In An Intermediate State
Organism: Archaeoglobus fulgidus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-15 Classification: HYDROLASE, LYASE/DNA Ligands: MOO |
 |
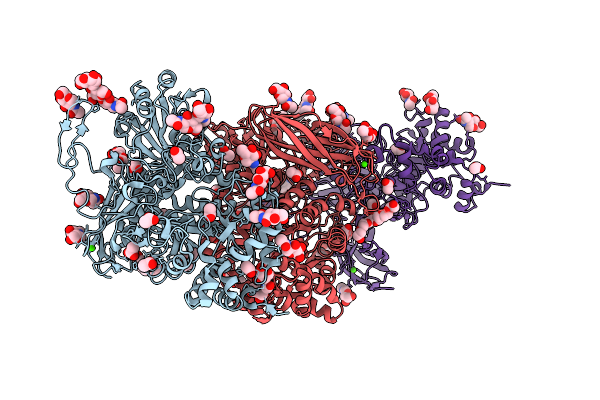
Organism: Archaeoglobus fulgidus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-15 Classification: HYDROLASE, LYASE/DNA Ligands: SO4 |
 |
Crystal Structure Of Isoprimeverose-Producing Enzyme From Phaeoacremonium Minimum
Organism: Rattus rattus, Phaeoacremonium minimum
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAG, ACT, PG4, CA, PGE, P6G |
 |
Organism: Gloeocapsa sp. pcc 73106
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-04-08 Classification: PHOTOSYNTHESIS |
 |
Organism: Neisseria gonorrhoeae fa 1090
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-01 Classification: BIOSYNTHETIC PROTEIN Ligands: SO4, PEG, GOL |
 |
Crystal Structure Of Selenomethionine-Substituted N-Oxygenase Rohs From Pseudomonas Syringae Pv. Tomato Str. Dc3000 (Pstorohs)
Organism: Pseudomonas syringae pv. tomato str. dc3000
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-01 Classification: BIOSYNTHETIC PROTEIN |
 |
Sub-Atomic Resolution (0.95 A) Xfel Structure Of As-Isolated Copper Nitrite Reductase From Achromobacter Cycloclastes Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox)
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:0.95 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
Sub-Atomic Resolution (0.95 A) Xfel Structure Of Nitrite-Bound Copper Nitrite Reductase From Achromobacter Cycloclastes Determined By Serial Femtosecond Rotation Crystallography (Sf-Rox) At 100 K
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:0.95 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: CU, SO4, NO2 |
 |
X-Ray Crystal Structure Of Pseudoazurin Met16Arg Variant, Oxidized Form At Ph 7.0
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2026-03-11 Classification: ELECTRON TRANSPORT Ligands: CU, PO4 |
 |
Low-Dose (14.9 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Room Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
Higher-Dose (283.1 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Room Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
Low-Dose (33.3 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Cryogenic Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
Higher-Dose (1332 Kgy) Structure Of Copper-Containing Nitrite Reductase From Achromobacter Cycloclastes At Cryogenic Temperature
Organism: Achromobacter cycloclastes
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-03-04 Classification: OXIDOREDUCTASE Ligands: CU, SO4 |
 |
Crystal Structure Of The Flavoprotein Monooxygenase Rslo9 From Streptomyces Bottropensis
Organism: Streptomyces bottropensis
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-11 Classification: OXIDOREDUCTASE Ligands: FAD, CL |

