Search Count: 26,847
All
Selected
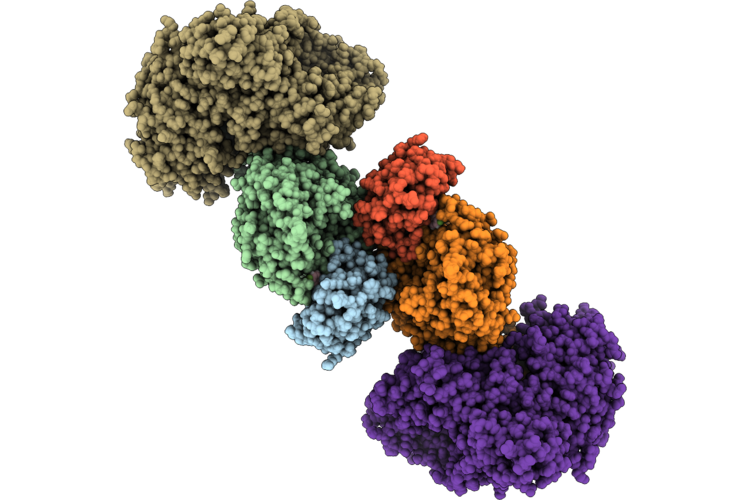 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: CYTOSOLIC PROTEIN Ligands: A1C4S, ZN |
 |
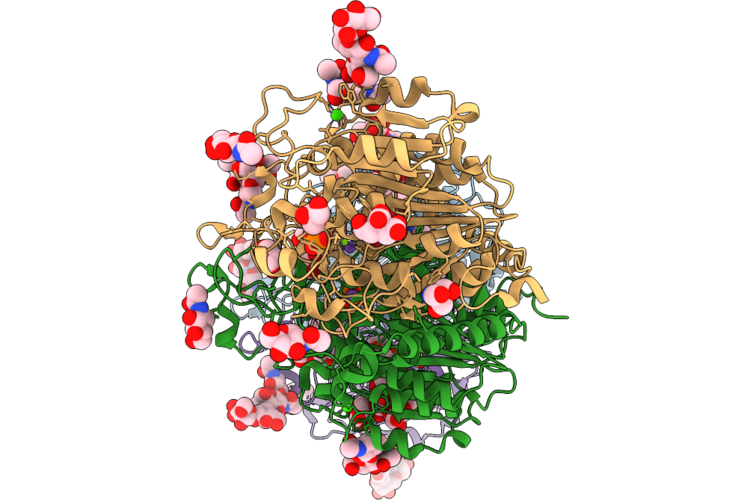
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: XDC, ZN, MG, CA, GOL, NA, NAG, MAN, PO4, TRS, FLC |
 |
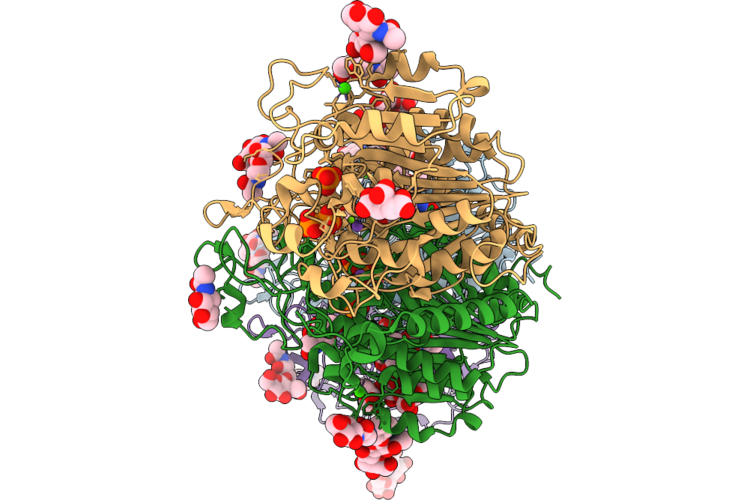
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PO4, ZN, MG, CA, NA, NAG, GOL, TRS, FLC |
 |
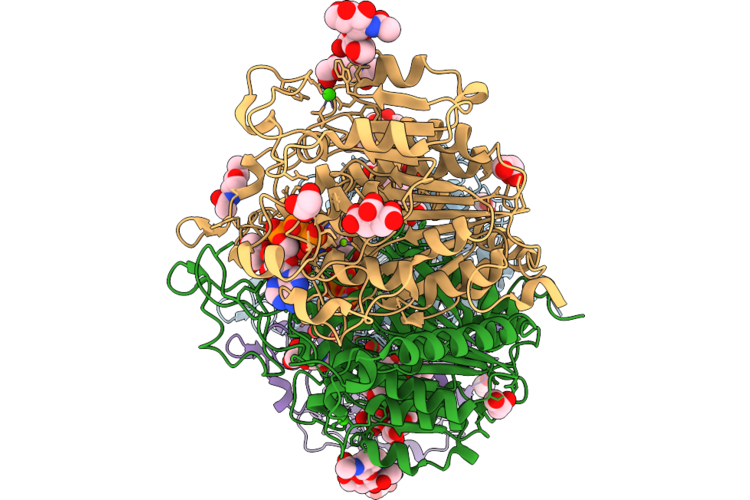
Tissue Non-Specific Alkaline Phosphatase -S110A Bound To Ppi With Ethylene Glycol
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: POP, ZN, MG, CA, NA, NAG, MAN, PO4, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ATP, ZN, MG, CA, GOL, NAG, FLC, PO4 |
 |
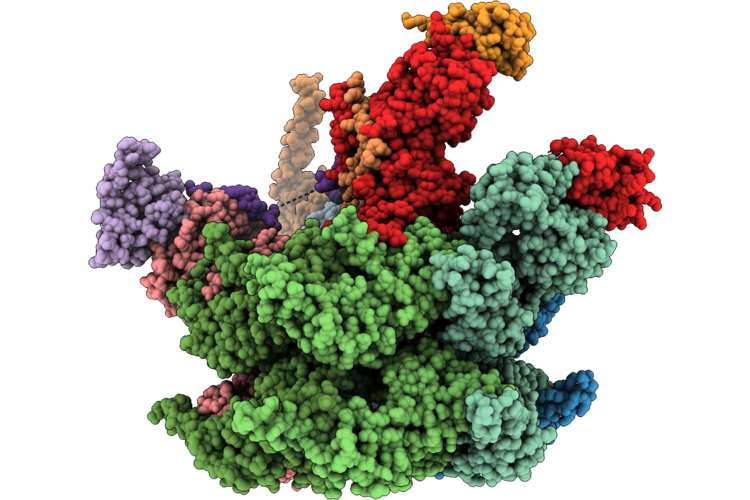
Cryo-Em Structure Of Substrate Engaged P97-Ufd1-Npl4-Faf1 Complex (Npl4 Focused)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ZN |
 |
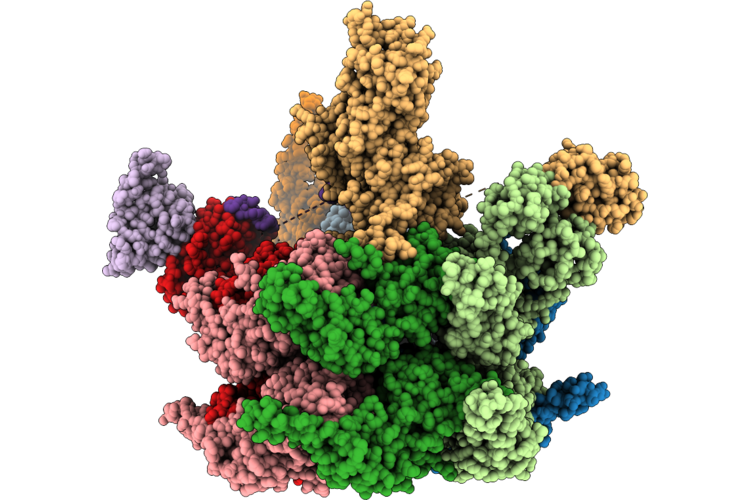
Cryo-Em Structure Of Substrate Engaged P97-Ufd1-Npl4-Faf1 Complex (Motor Focused)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ZN, ATP, ADP |
 |
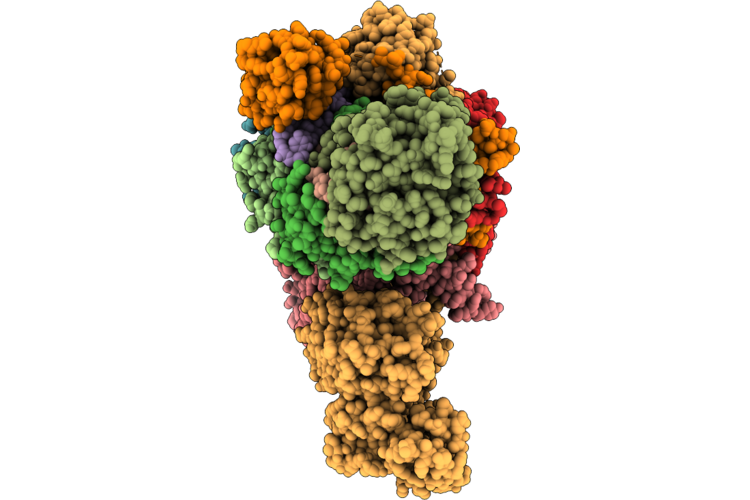
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.85 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ZN, ATP, ADP |
 |
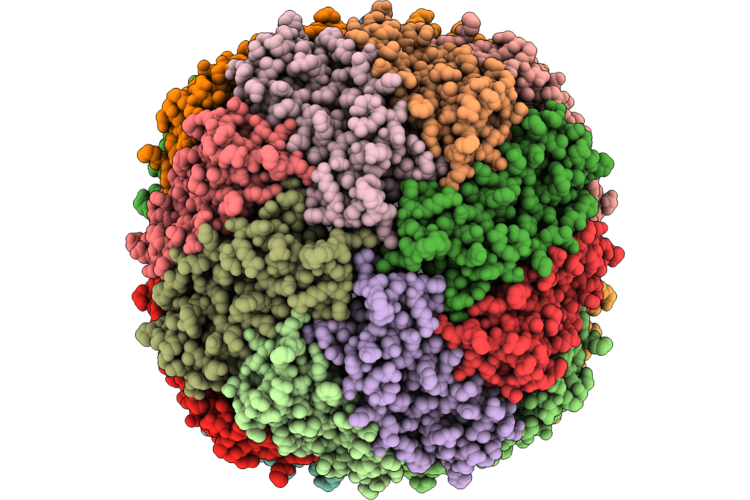
Cryo-Em Structure Of Type Vii Crispr-Cas Complex At The Target Engagement State
Organism: Metagenome
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: ANTIVIRAL PROTEIN/RNA Ligands: ZN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: RECOMBINATION Ligands: FE, ZN |
 |
Organism: Mycobacterium phage ds6a
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-04-22 Classification: VIRUS Ligands: ZN, PO4 |
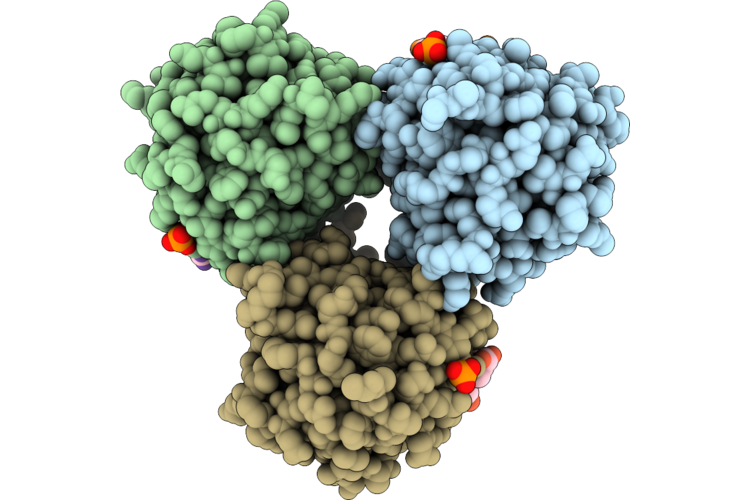 |
Crystal Structure Of Ami2B Domain Of Ds6A-Lysa In Complex With L-Ala-D-Iso-Gln-L-Lys- D-Ala-D-Ala
Organism: Mycobacterium phage ds6a, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PO4, ZN |
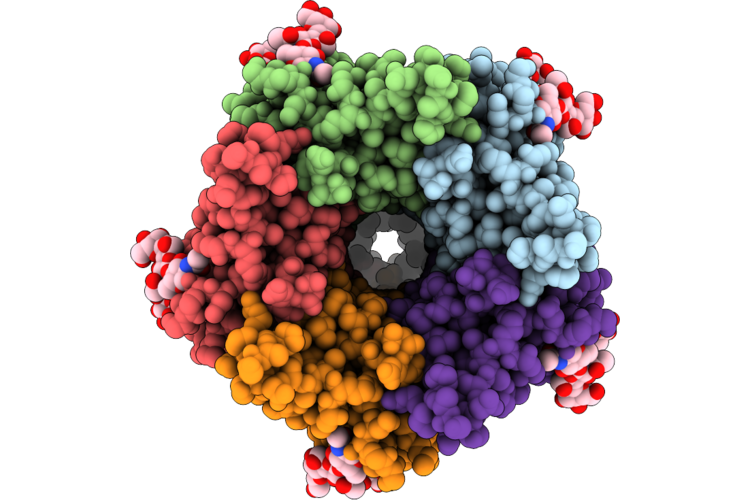 |
Organism: Homo sapiens, Enterovirus a71
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ZN |
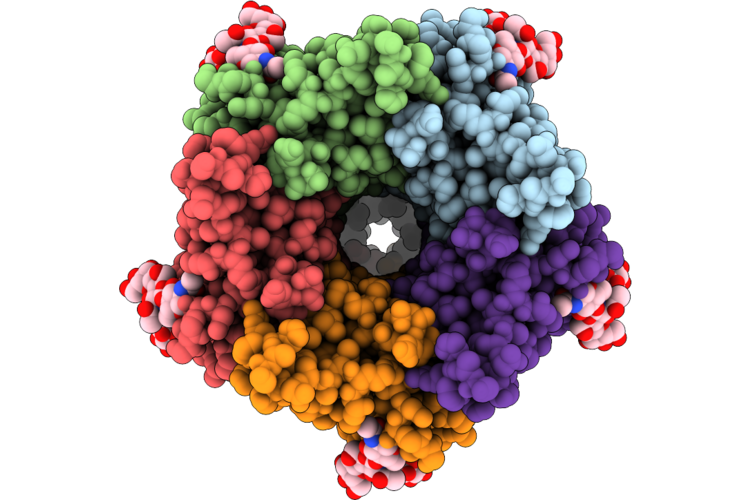 |
Organism: Homo sapiens, Human enterovirus 71
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ZN |
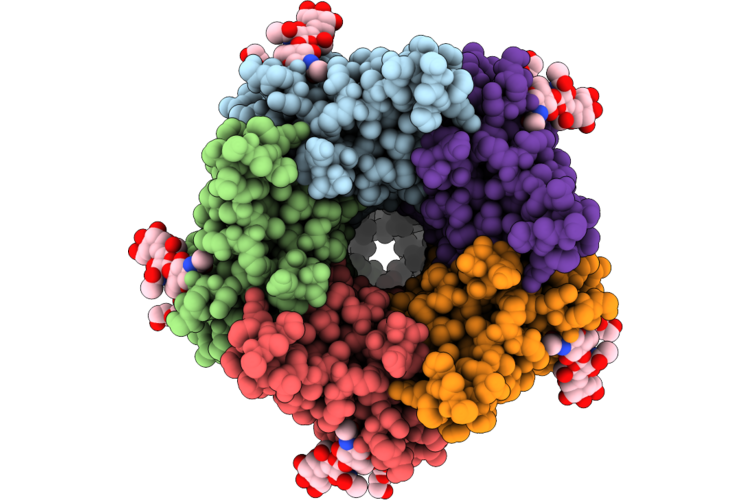 |
Organism: Homo sapiens, Human enterovirus 71
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ZN |
 |
Cryo-Em Structure Of Human Zac In Complex With N-(4-(Tert-Butyl)Thiazol-2-Yl)-3-Fluorobenzamide (Ttfb)
Organism: Homo sapiens, Enterovirus a71
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: A1EKJ, ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.13 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Major State, Active Conformation
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.15 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Oxidizing Conditions, Dimer, Minor State
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.45 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |
 |
Cryo-Em Structure Of H. Neapolitanus Csosca In Reducing Conditions, Hexamer
Organism: Escherichia coli k-12, Halothiobacillus neapolitanus c2
Method: ELECTRON MICROSCOPY Resolution:2.06 Å Release Date: 2026-04-22 Classification: LYASE Ligands: ZN |

