Search Count: 73,386
All
Selected
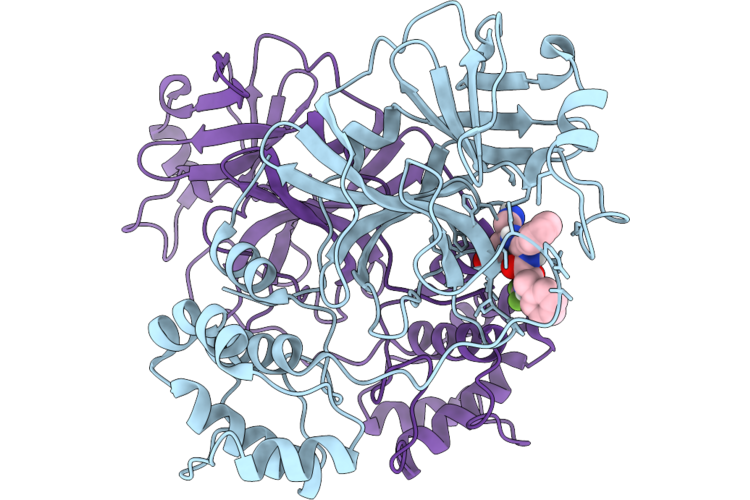 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-04-22 Classification: VIRAL PROTEIN Ligands: A1DAK |
 |
Organism: Manduca sexta
Method: SOLUTION NMR Release Date: 2026-04-22 Classification: ANTIMICROBIAL PROTEIN |
 |
Organism: Manduca sexta
Method: SOLUTION NMR Release Date: 2026-04-22 Classification: ANTIMICROBIAL PROTEIN |
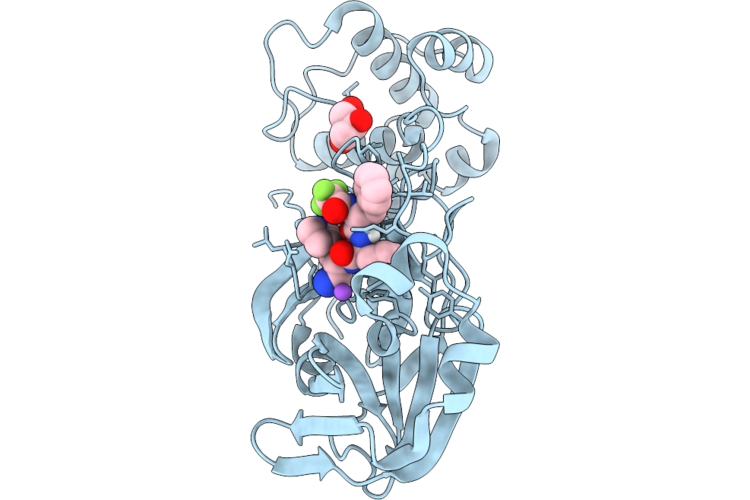 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-04-22 Classification: VIRAL PROTEIN Ligands: GOL, A1DA8, NA |
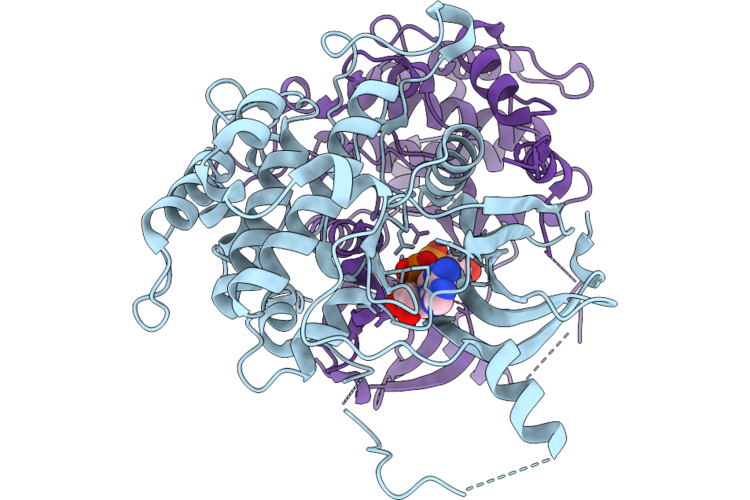 |
Crystal Structure Of Serine/Threonine-Protein Kinase (Aek1) From Trypanosoma Cruzi In Complex With Amp
Organism: Trypanosoma cruzi
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: AMP |
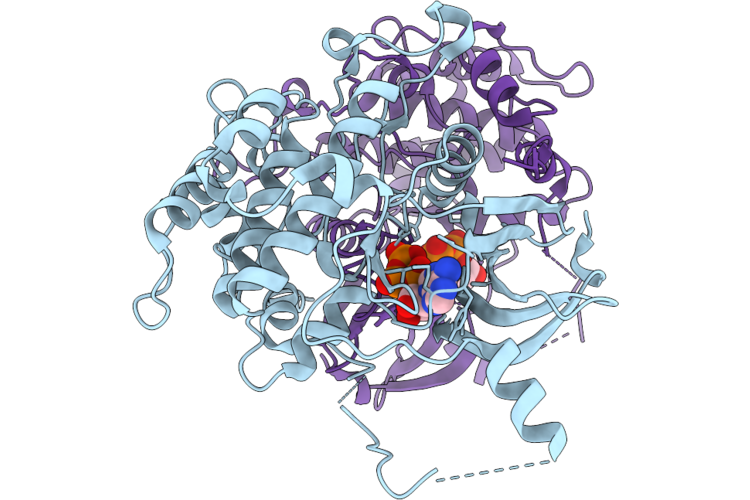 |
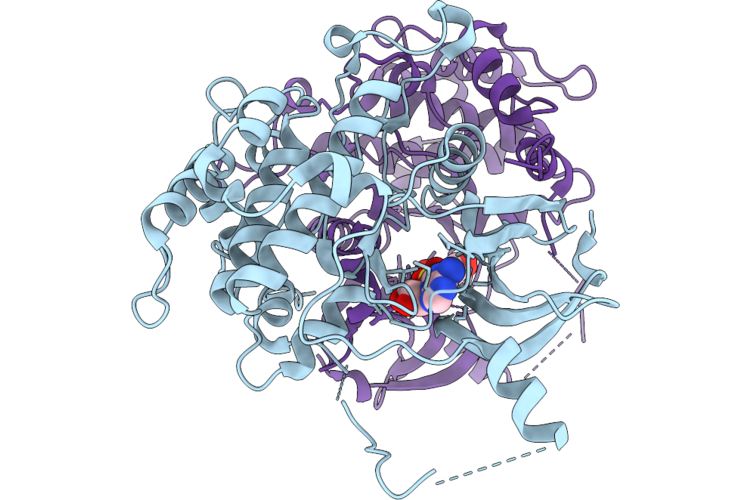
Crystal Structure Of Serine/Threonine-Protein Kinase (Aek1) From Trypanosoma Cruzi In Complex With Atp
Organism: Trypanosoma cruzi
Method: X-RAY DIFFRACTION Resolution:2.68 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: ATP |
 |
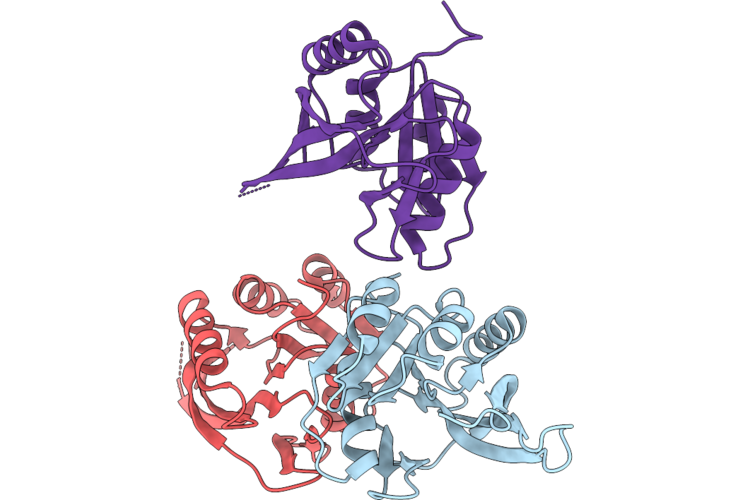
Crystal Structure Of Serine/Threonine-Protein Kinase (Aek1) From Trypanosoma Cruzi In Complex With Lms
Organism: Trypanosoma cruzi
Method: X-RAY DIFFRACTION Resolution:2.82 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: LMS |
 |
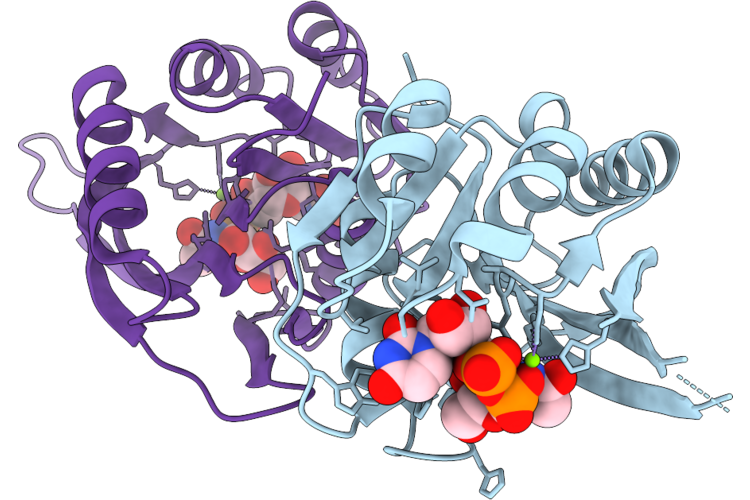
Organism: Bacillus cereus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-22 Classification: TRANSFERASE |
 |
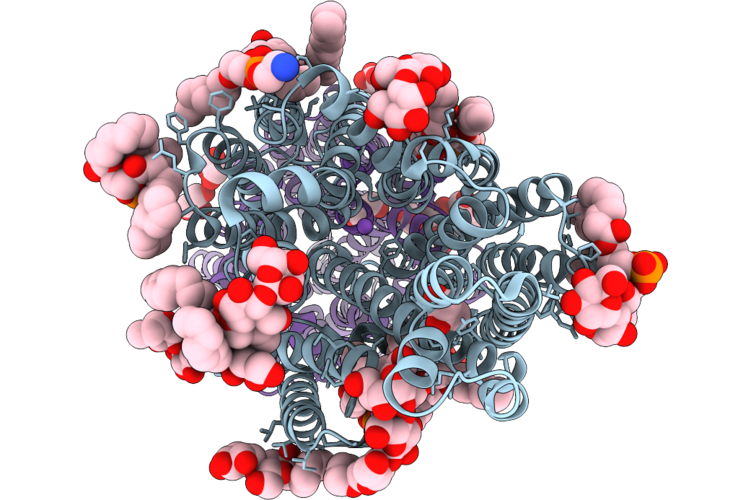
Crystal Structure Of Bacillus Cereus Gmar In Complex With Udp-Glcnac And Mg2+
Organism: Bacillus cereus
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: UD1, MG |
 |
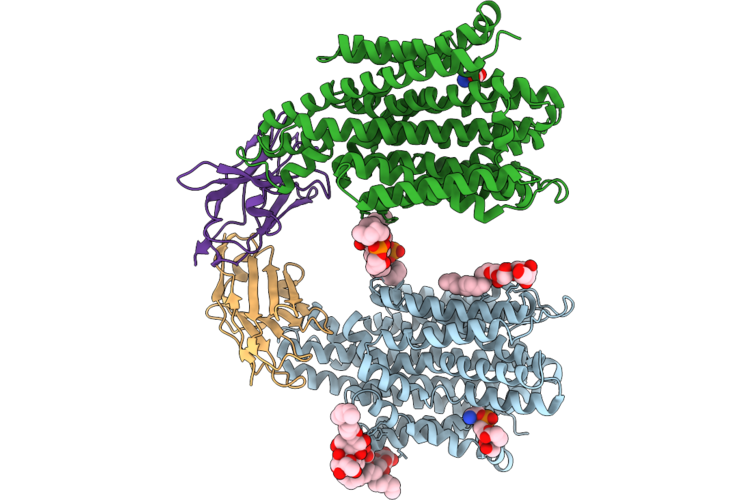
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Outward Open State
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.83 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: PGT, LMU, PTY, 3PH, GOL, NA |
 |
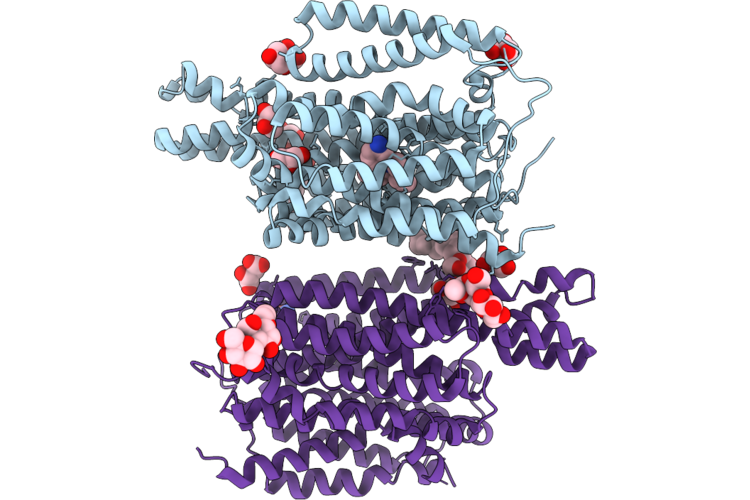
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Inward Open State In Complex With Nanobody 89
Organism: Staphylococcus aureus, Lama glama
Method: X-RAY DIFFRACTION Resolution:3.32 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: PTY, PGT, LMU, OXM |
 |
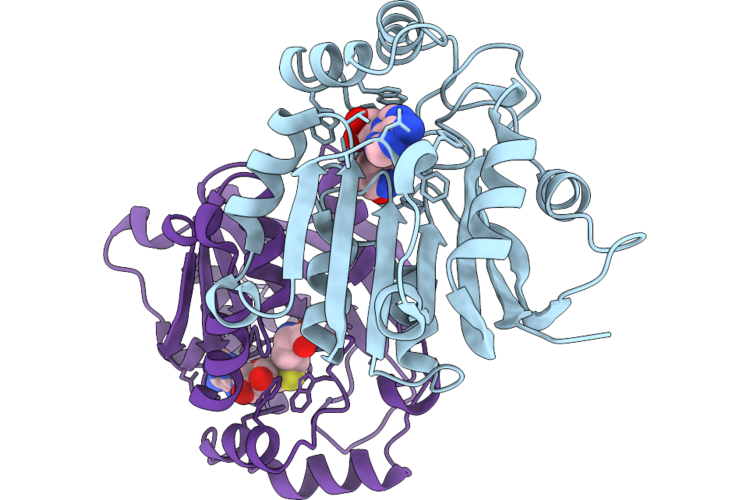
Crystal Structure Of The Staphylococcal Efflux Pump Qaca In The Outward Open State Bound To Ethidium
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.79 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ET, GOL, TLA, CL, LMU, OXM |
 |
Crystal Structure Of Halide Methyltransferase From Moniliophthora Roreri In Complex With Sah
Organism: Moniliophthora roreri
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: SAH |
 |
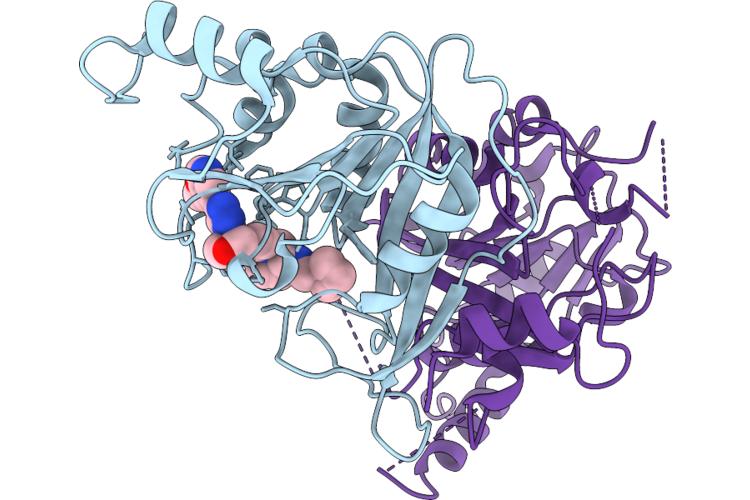
Crystal Structure Of Halide Methyltransferase From Dichomitus Squalens In Complex With Sah
Organism: Dichomitus squalens
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: SAH, ACT |
 |
Crystal Structure Of The Human Mettl3-Mettl14 In Complex With Small Molecule Inhibitor Compound 8
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-04-22 Classification: RNA BINDING PROTEIN Ligands: A1I4E |
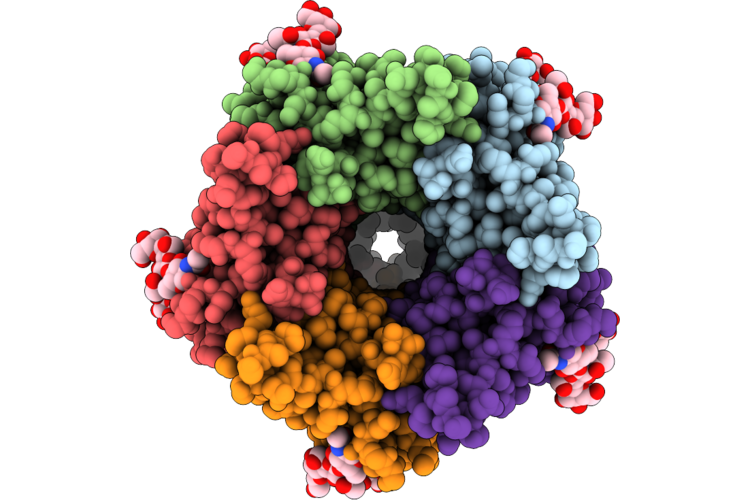 |
Organism: Homo sapiens, Enterovirus a71
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ZN |
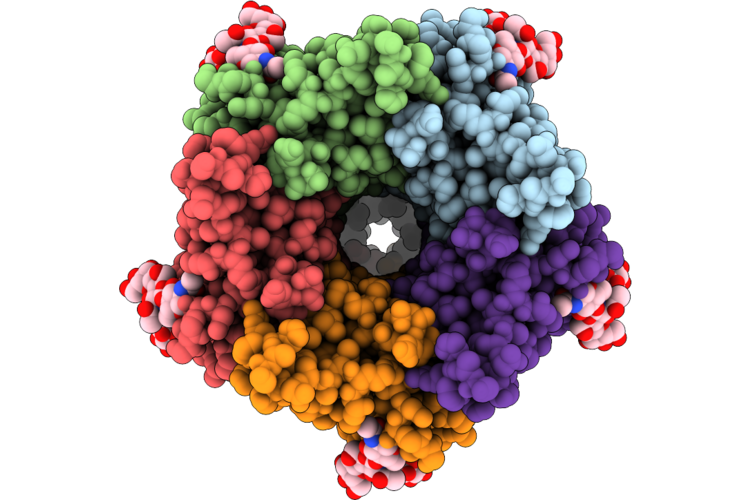 |
Organism: Homo sapiens, Human enterovirus 71
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ZN |
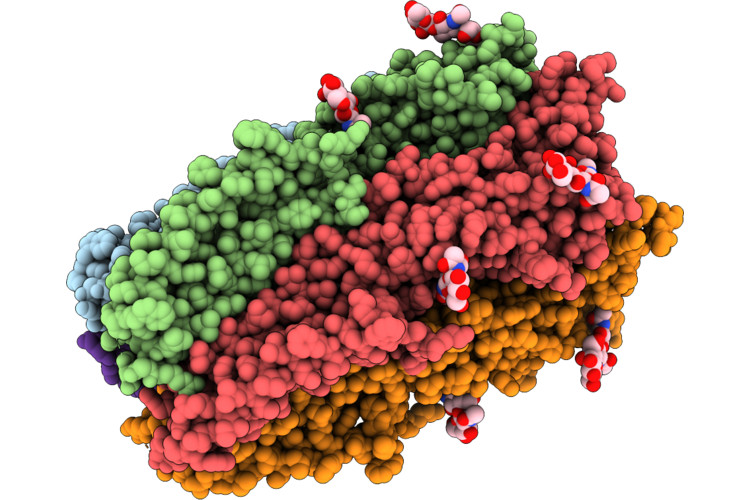 |
Organism: Homo sapiens, Enterovirus a71
Method: ELECTRON MICROSCOPY Resolution:3.35 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN |
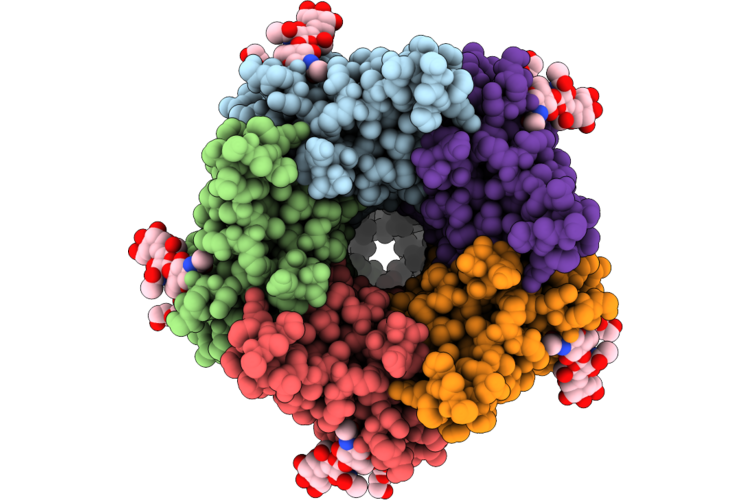 |
Organism: Homo sapiens, Human enterovirus 71
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ZN |
 |
Organism: Homo sapiens, Human enterovirus 71
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: TC9 |

