Search Count: 5,463
All
Selected
 |
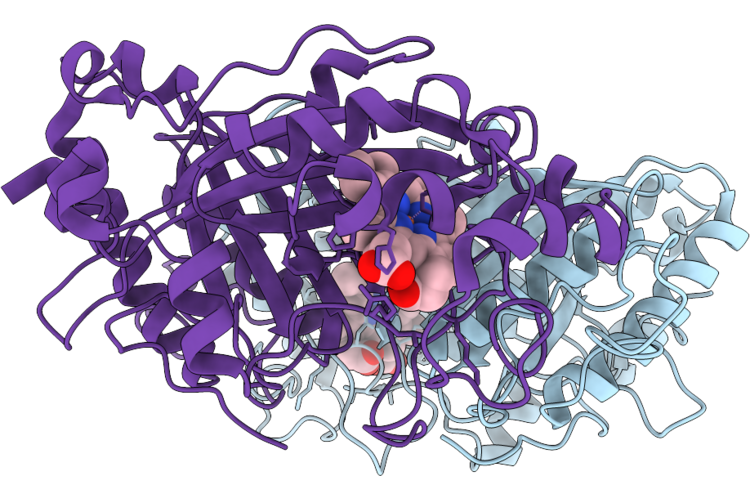
Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans.
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Reprocessing And Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
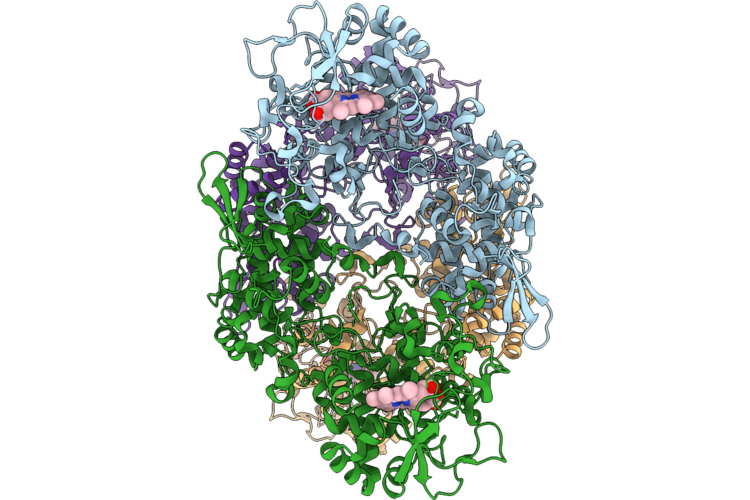 |
Cryoem Structure Of Eckatg S-Trp105 At 2.22 Angstrom Resolution Revealing An Asymmetric Sulfur Center In O=S-Trp
Organism: Escherichia coli k-12
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
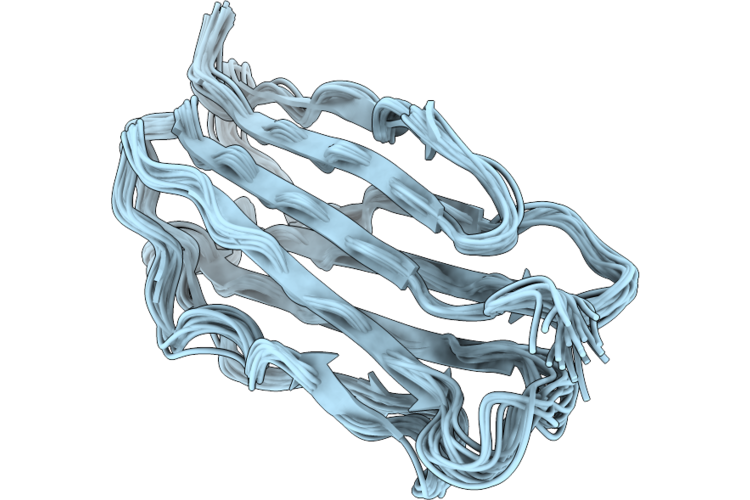 |
Organism: Mattirolomyces terfezioides
Method: SOLUTION NMR Release Date: 2026-04-22 Classification: UNKNOWN FUNCTION |
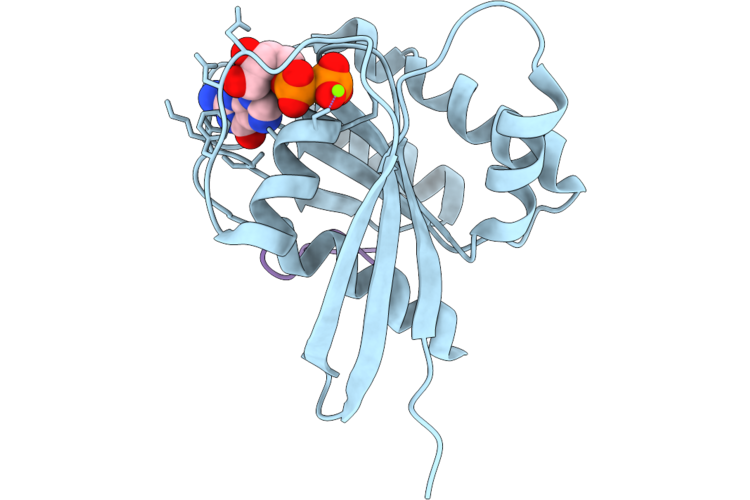 |
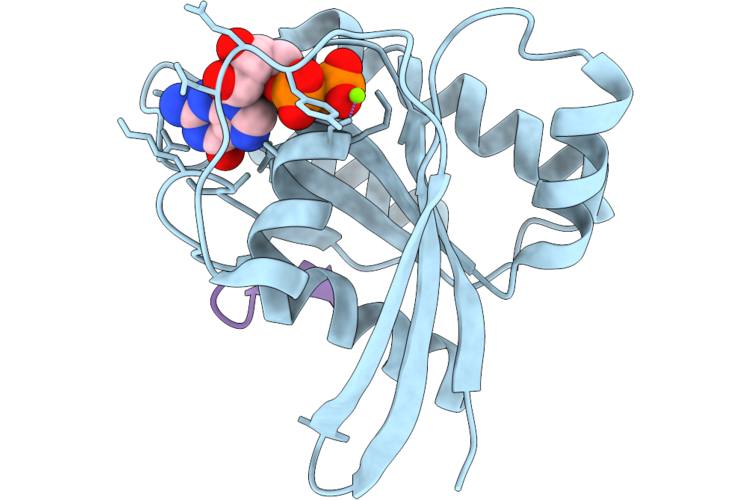
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
 |
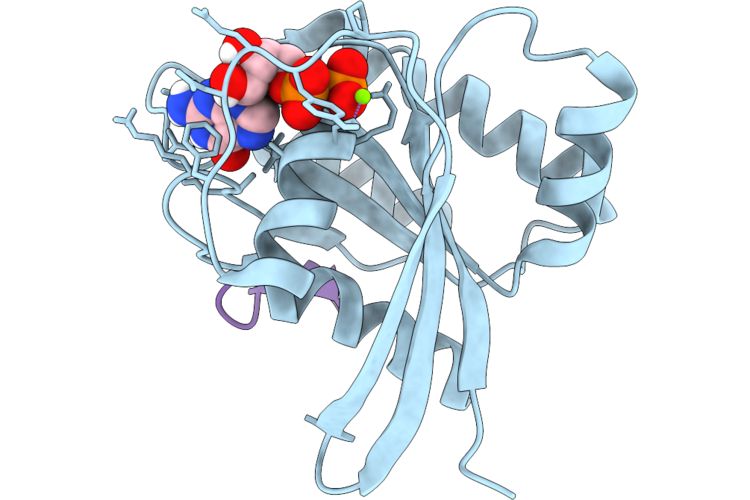
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
 |
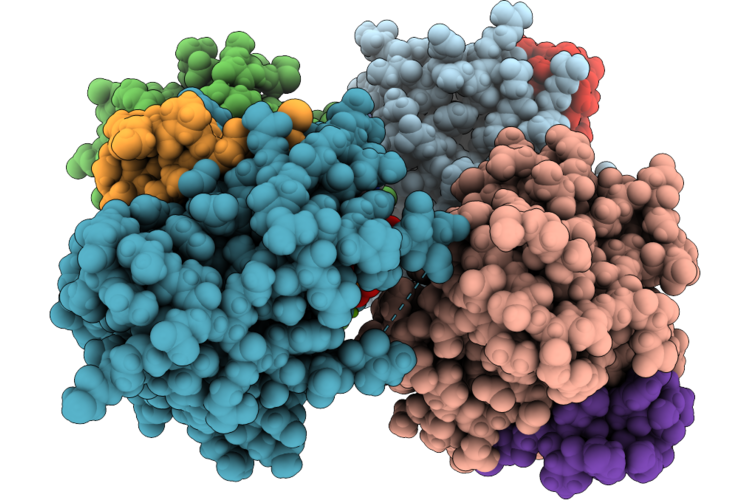
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
 |
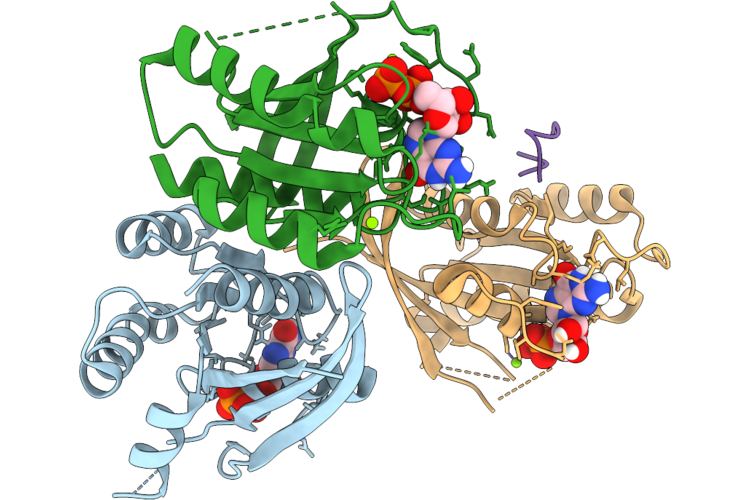
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: MG, GDP |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
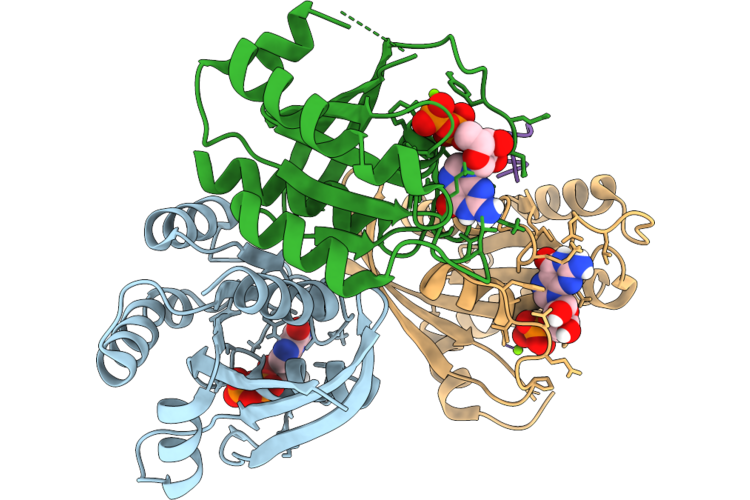 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
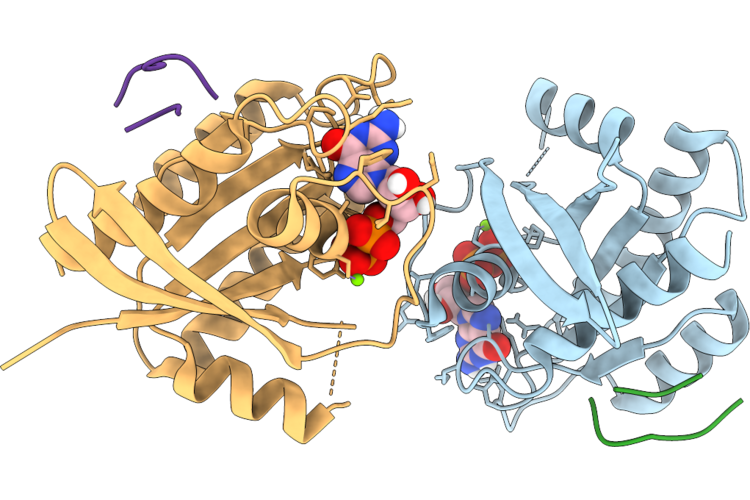 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: MG, GDP, PO4 |
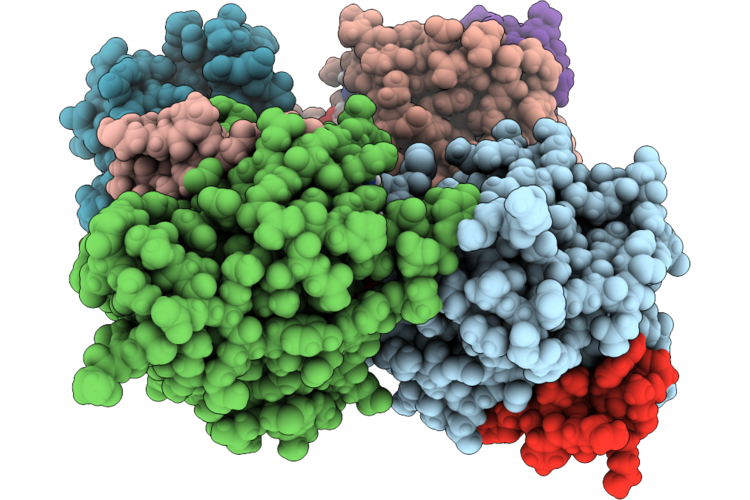 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP |
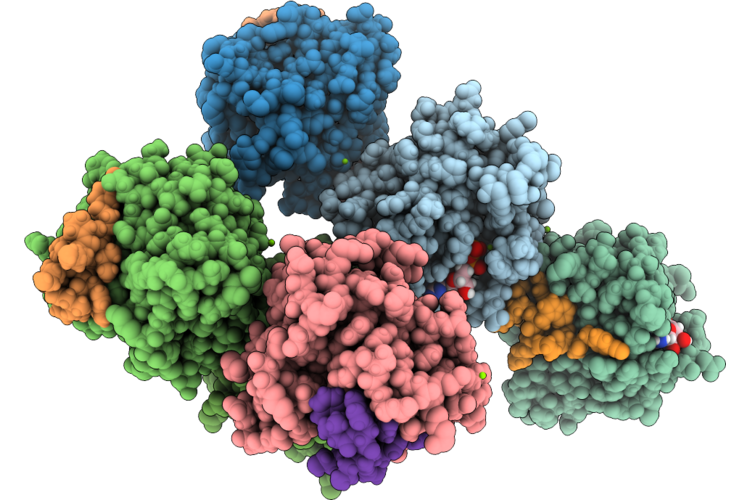 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE/INHIBITOR Ligands: GDP, MG |
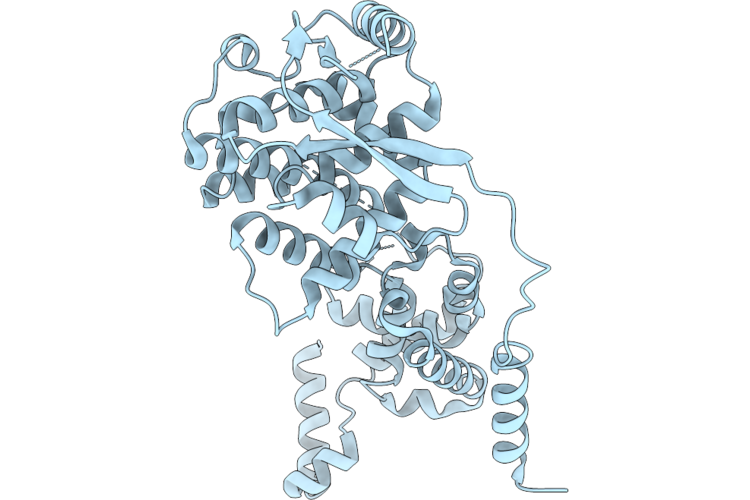 |
Organism: Langya virus
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-22 Classification: VIRAL PROTEIN |
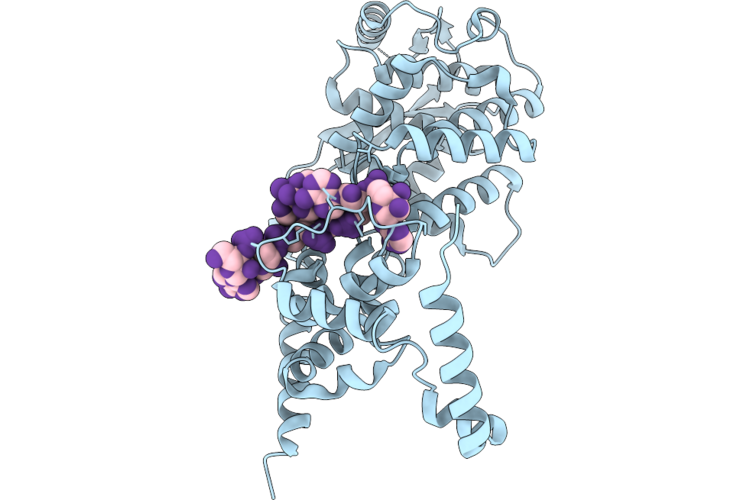 |
Helical Reconstruction Of Langya Henipavirus N-Core Nucleocapsid-Like Complex
Organism: Langya virus, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-04-22 Classification: VIRAL PROTEIN |
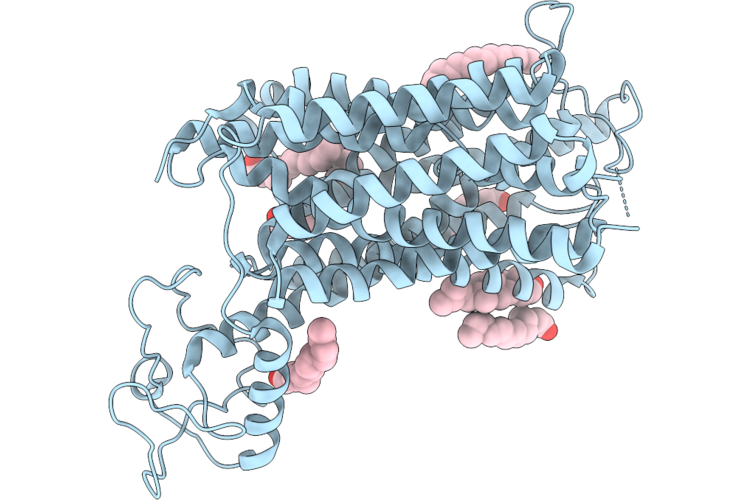 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:3.54 Å Release Date: 2026-04-22 Classification: TRANSPORT PROTEIN Ligands: POV, CLR |
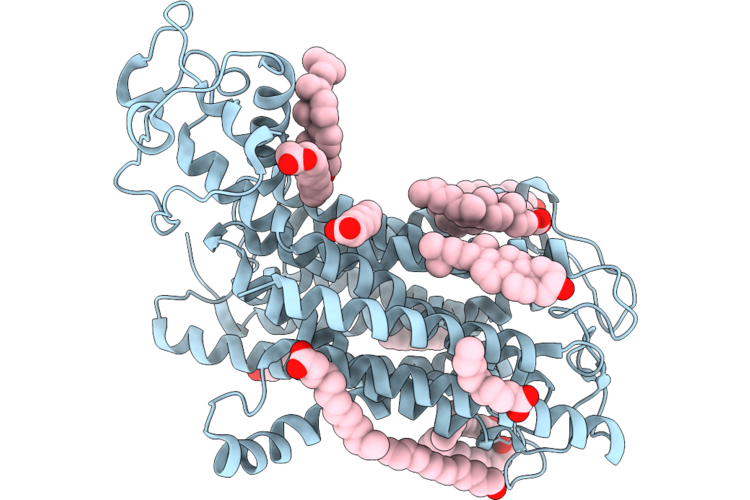 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:3.22 Å Release Date: 2026-04-22 Classification: TRANSPORT PROTEIN Ligands: CLR, POV |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:3.27 Å Release Date: 2026-04-22 Classification: TRANSPORT PROTEIN Ligands: CLR, POV, A1EP2 |
 |
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:3.14 Å Release Date: 2026-04-22 Classification: TRANSPORT PROTEIN Ligands: CLR, A1EPL, POV |

