Search Count: 29,254
All
Selected
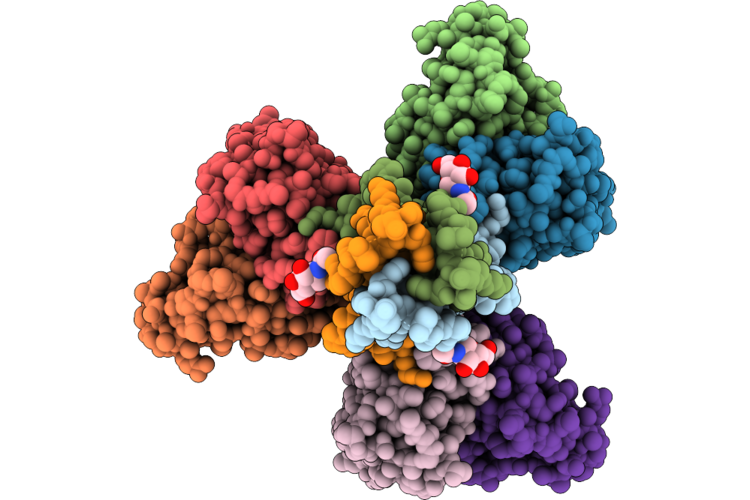 |
Sars-Cov-2 Spike S2 Trimer Stabilized In The Early Fusion Intermediate Conformation (E-Fics-V3) Bound To The Vn01H1 Fab (Fab Local Refinement)
Organism: Saccharomyces cerevisiae s288c, Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
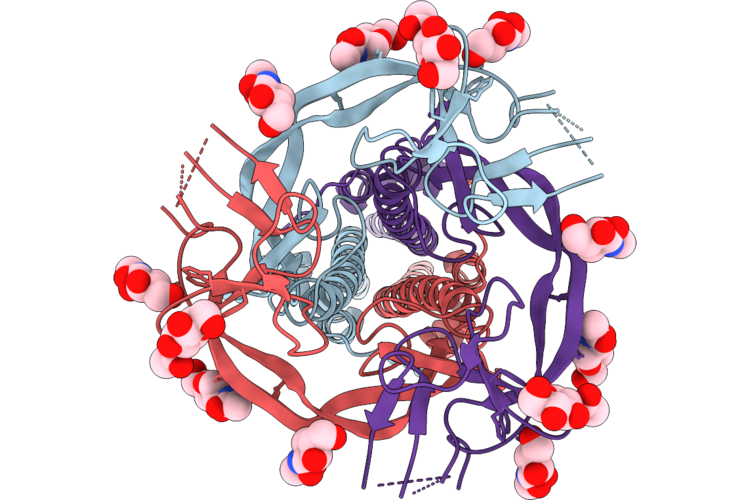 |
Sars-Cov-2 Spike S2 Trimer Stabilized In The Early Fusion Intermediate Conformation (E-Fics-V3) Bound To The Vn01H1 Fab (S2 Local Refinement)
Organism: Saccharomyces cerevisiae s288c, Severe acute respiratory syndrome coronavirus 2
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
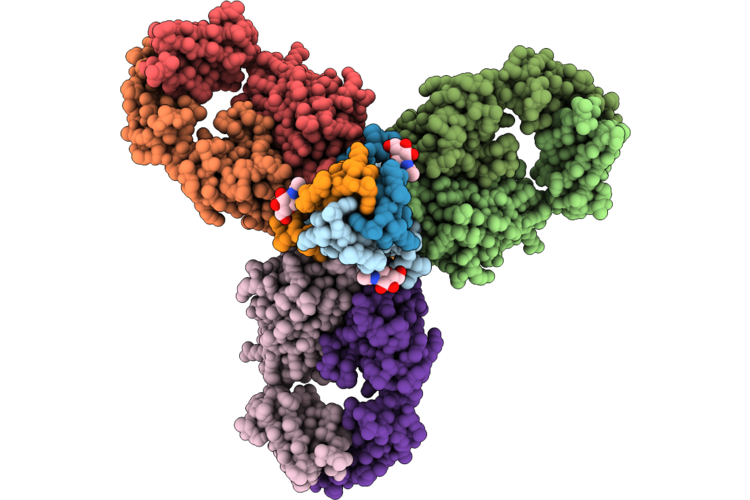 |
Sars-Cov-2 Spike S2 Trimer Stabilized In The Early Fusion Intermediate Conformation (E-Fics-V3) Bound To C77G12 (Fab Local Refinement)
Organism: Saccharomyces cerevisiae s288c, Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: VIRAL PROTEIN Ligands: NAG |
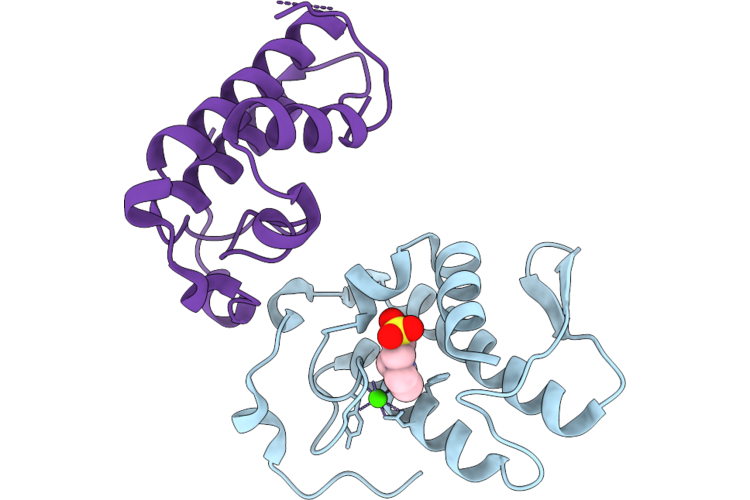 |
Organism: Lachesis muta
Method: X-RAY DIFFRACTION Resolution:2.36 Å Release Date: 2026-05-13 Classification: TOXIN,HYDROLASE Ligands: MES, CA |
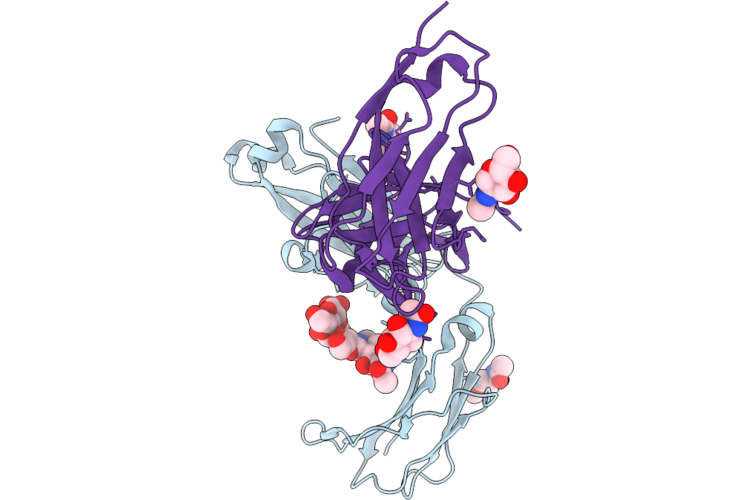 |
Crystal Structure Of The Human Iga2M2 Fc Fragment-Fc-Alpha Receptor (Cd89) Complex
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2026-05-13 Classification: IMMUNE SYSTEM Ligands: NAG, CL |
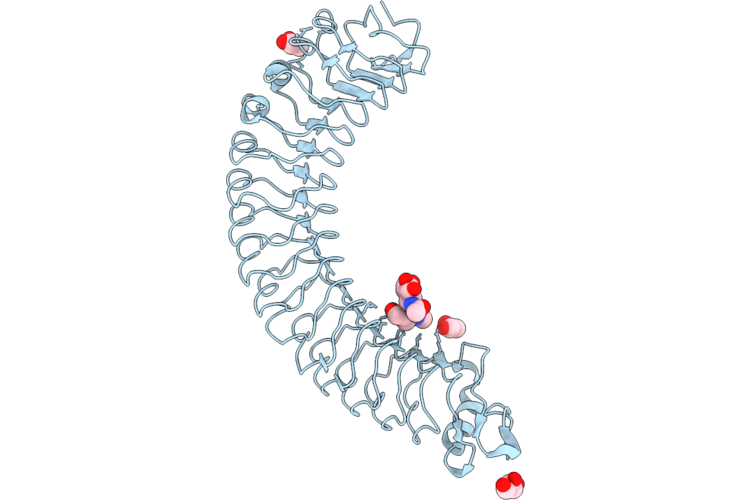 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2026-05-13 Classification: SIGNALING PROTEIN Ligands: GOL |
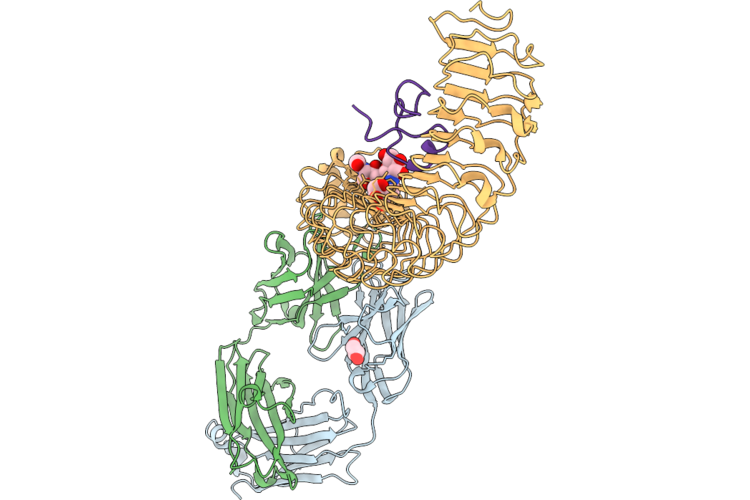 |
Mouse Lrrc15 Extracellular Domain In Complex With Disulfide Constrained Peptide Ml-Ysd-07 And Fab-E1
Organism: Homo sapiens, Mus musculus, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.57 Å Release Date: 2026-05-13 Classification: SIGNALING PROTEIN/Immune System Ligands: GOL |
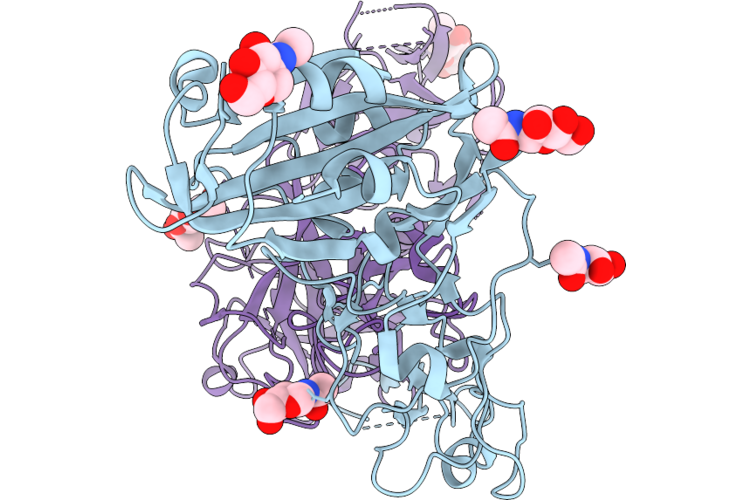 |
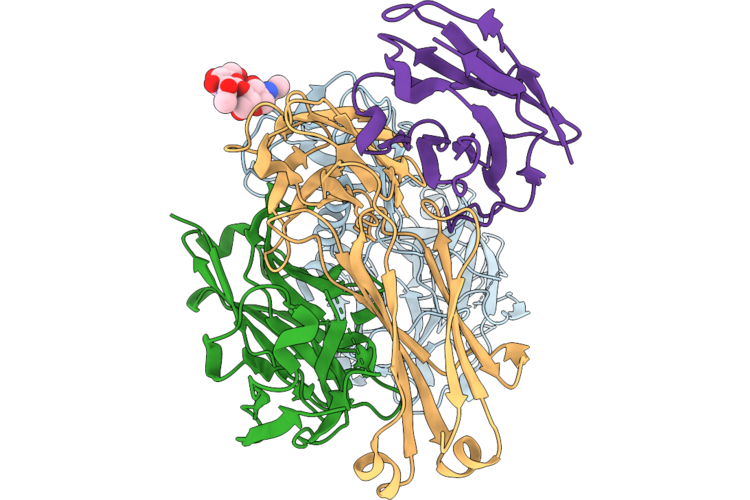
Tmprss2 (S441A) Bound To The Hcov-Nl63 S2'Region Genetically Fused To The Hcov-Hku1 Rbd
Organism: Human coronavirus hku1, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-05-13 Classification: VIRAL PROTEIN/HYDROLASE Ligands: NAG |
 |
Tmprss2 S441A In Complex With The H1H7 Fab And Anti-Kappa Light Chain Nanobody
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-05-13 Classification: HYDROLASE/IMMUNE SYSTEM |
 |
Organism: Lotus japonicus
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-05-13 Classification: OXYGEN BINDING Ligands: HEM, CYN, PO4 |
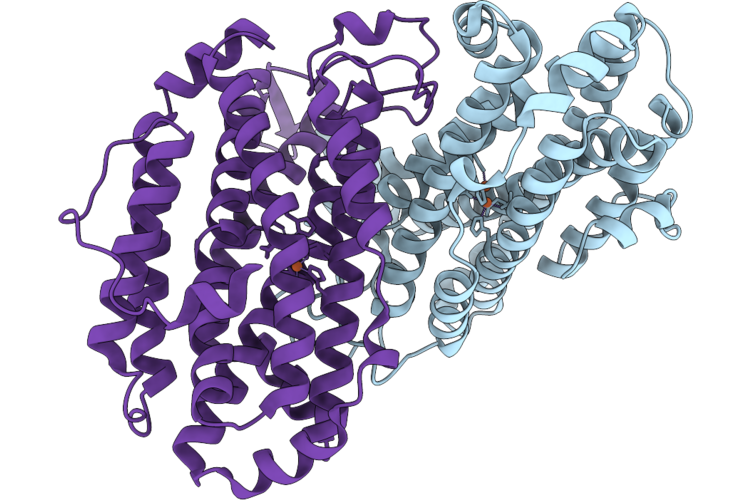 |
Xfel Structure Of Oxidised Ribonucleotide Reductase R2A Y122F Mutant From E. Coli
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE |
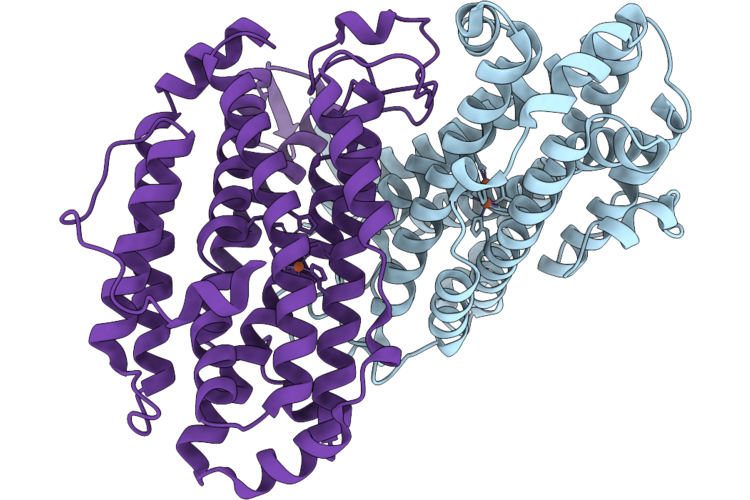 |
Xfel Structure Of Ribonucleotide Reductase R2A Y122F Mutant From E. Coli,Reduced Form
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE2 |
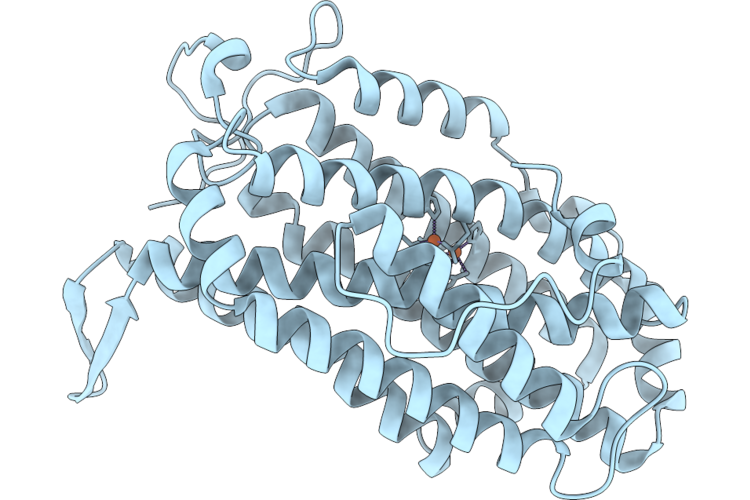 |
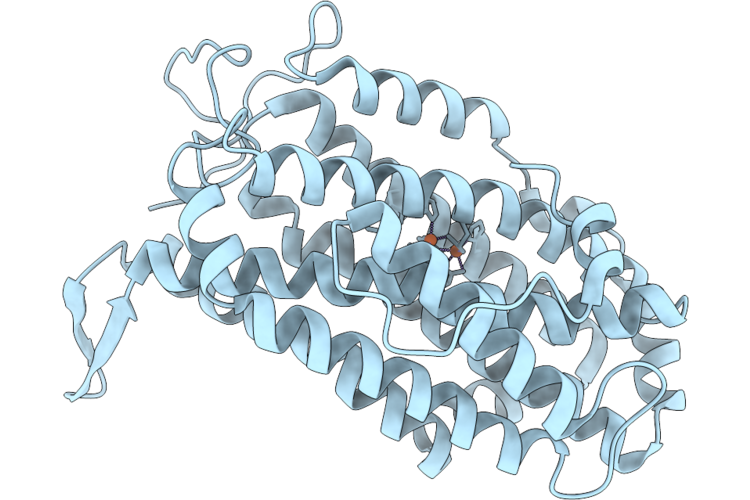
Xfel Structure Of Oxidised Ribonucleotide Reductase R2A Y122F Mutant From E. Coli, Hexagonal P6122 Form
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE |
 |
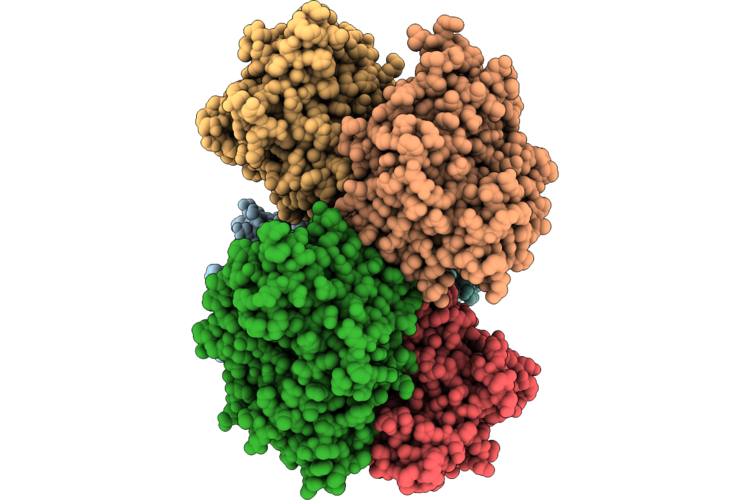
Xfel Structure Of Ribonucleotide Reductase R2A Y122F Mutant From E. Coli,Reduced Form, Hexagonal P6122
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE2 |
 |
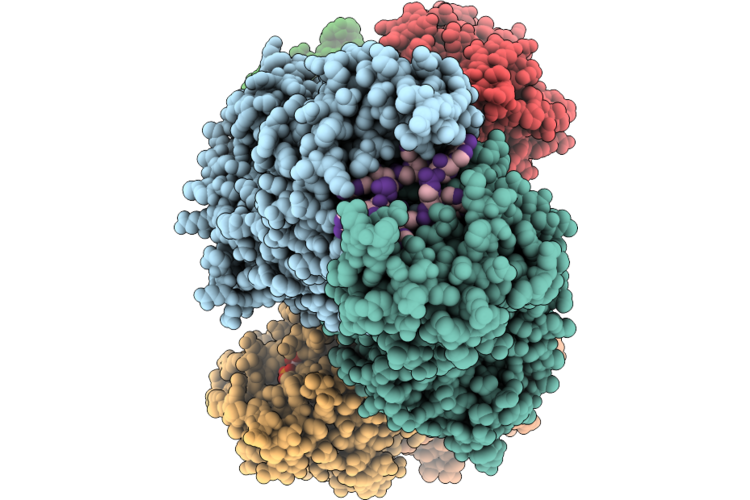
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.74 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
 |
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
 |
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, MG, ZN |
 |
Structure Of The Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 4
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
 |
Structure Of The Type Iii Crispr-Associated Deaminase In Complex Ca6 And Atp, State 5
Organism: Thermoanaerobaculum aquaticum
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-13 Classification: HYDROLASE/RNA Ligands: ATP, ZN, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-05-13 Classification: IMMUNE SYSTEM Ligands: Q87, ACT, CL, GOL, NA |

