Search Count: 3,153
All
Selected
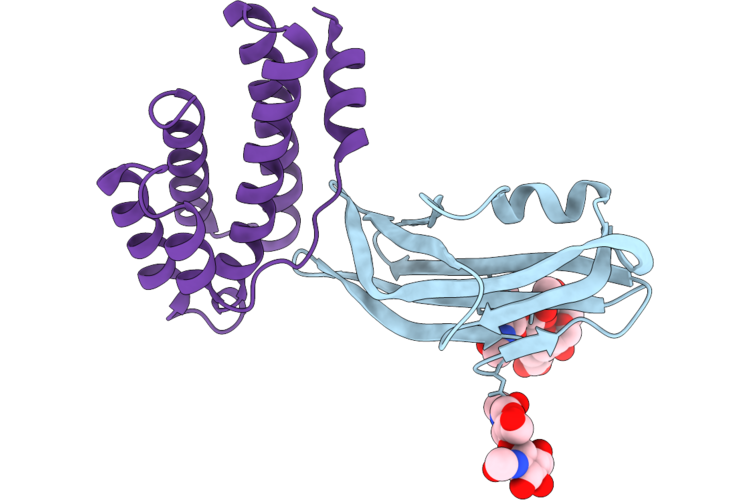 |
Structure Of The Porcine Deltacoronavirus (Pdcov) Receptor-Binding Domain Bound To The Rbd Minibinder 11, The Pd3 Fab, And The Kappa Light Chain Nanobody (Local Refinement)
Organism: Porcine deltacoronavirus, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-05-13 Classification: VIRAL PROTEIN |
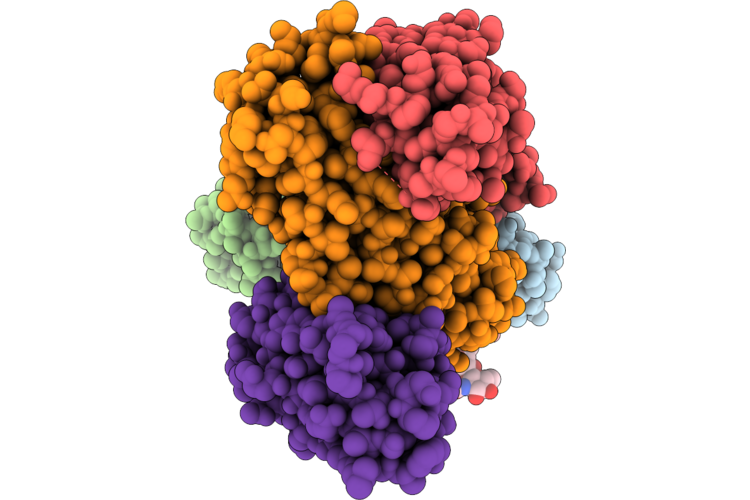 |
Structure Of The Porcine Deltacoronavirus (Pdcov) Receptor-Binding Domain Bound To The Rbd Minibinder 11, The Pd3 Fab, And The Kappa Light Chain Nanobody
Organism: Porcine deltacoronavirus, Synthetic construct, Mus musculus, Lama glama
Method: ELECTRON MICROSCOPY Resolution:2.80 Å Release Date: 2026-05-13 Classification: VIRAL PROTEIN |
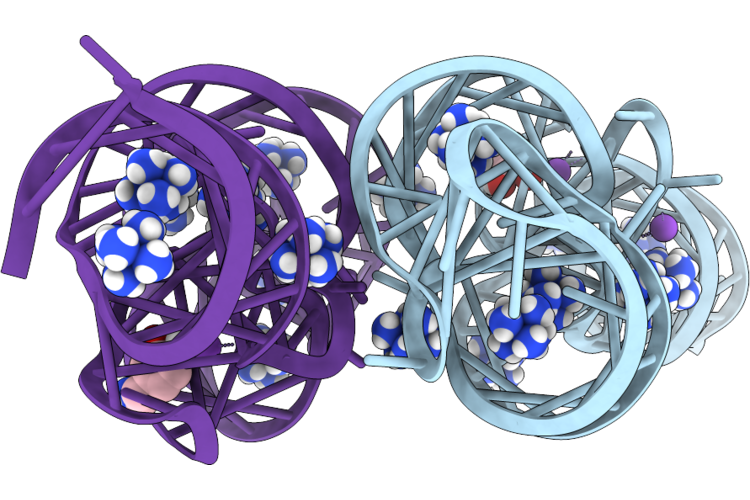 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2026-05-13 Classification: RNA Ligands: K, NCO, LDP |
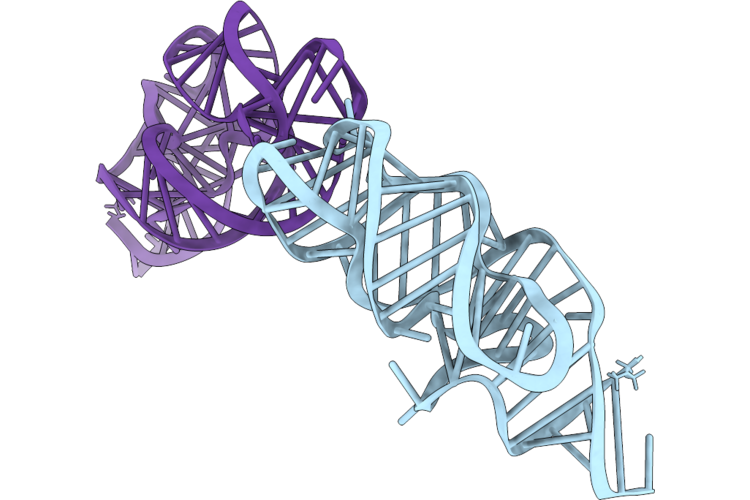 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.93 Å Release Date: 2026-05-13 Classification: RNA Ligands: SR |
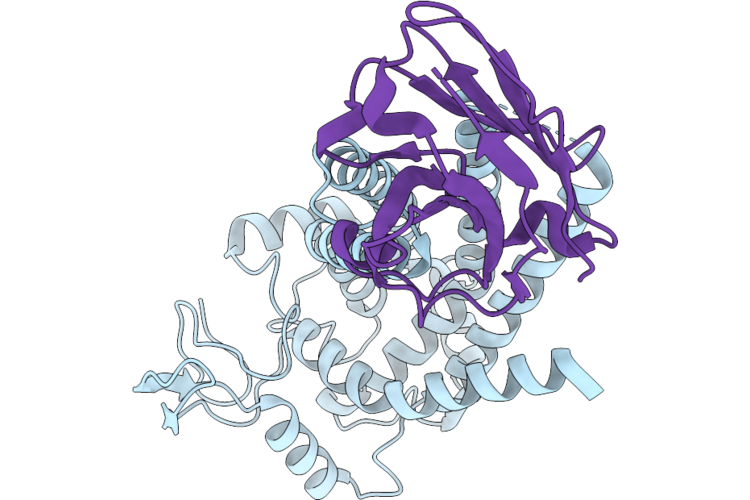 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-05-13 Classification: MEMBRANE PROTEIN |
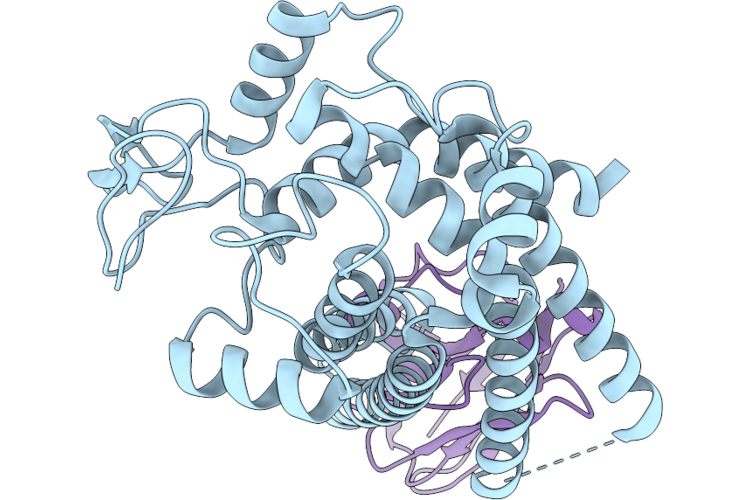 |
Single Particle Cryo-Em Structure Of Human Mtch2 (Hyperactive Mutant F285N F286N)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-05-13 Classification: MEMBRANE PROTEIN |
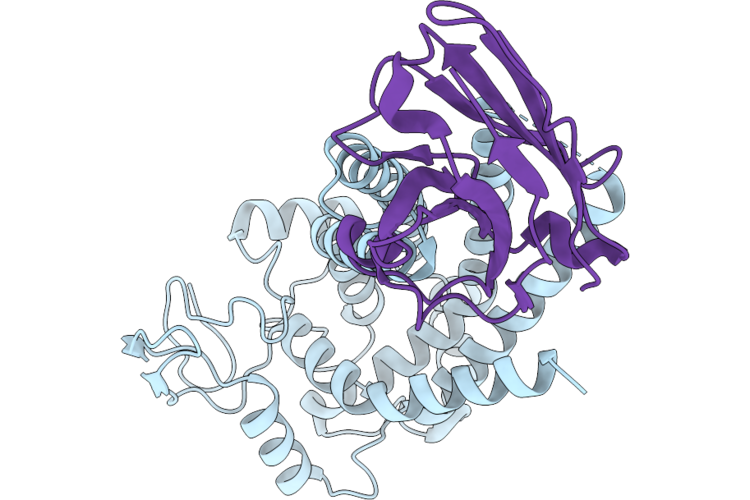 |
Single Particle Cryo-Em Structure Of Human Mtch2 (Hyperactive Mutant K25E Y235A V238D)
Organism: Synthetic construct, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.10 Å Release Date: 2026-05-13 Classification: MEMBRANE PROTEIN |
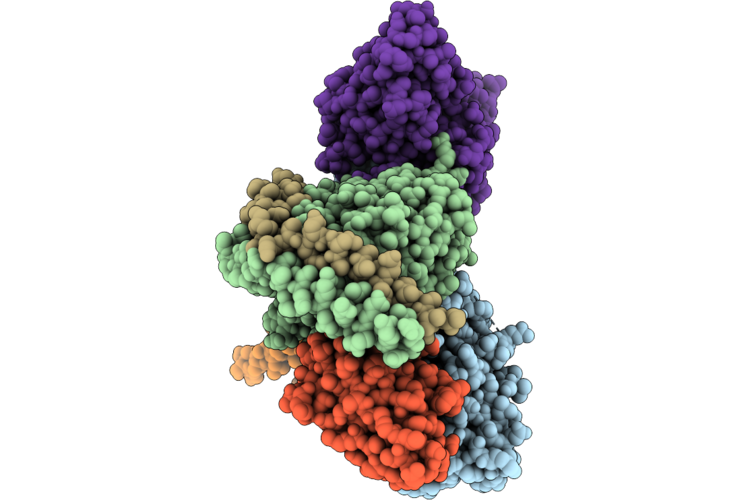 |
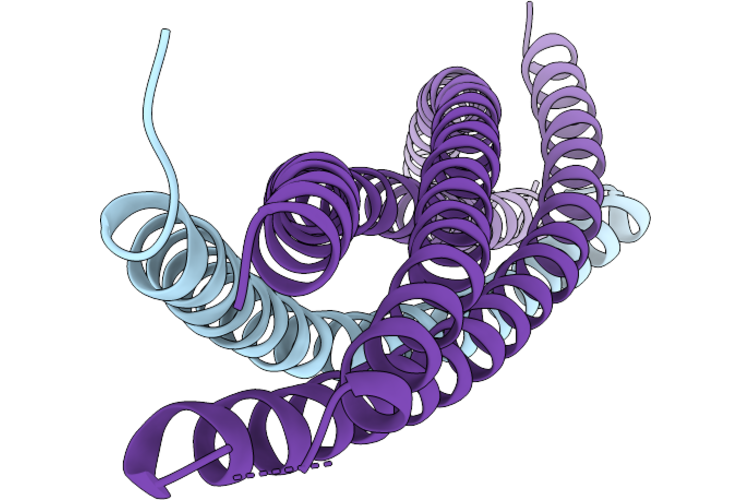
Organism: Petromyzon marinus, Homo sapiens, Lama glama, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: SPM |
 |
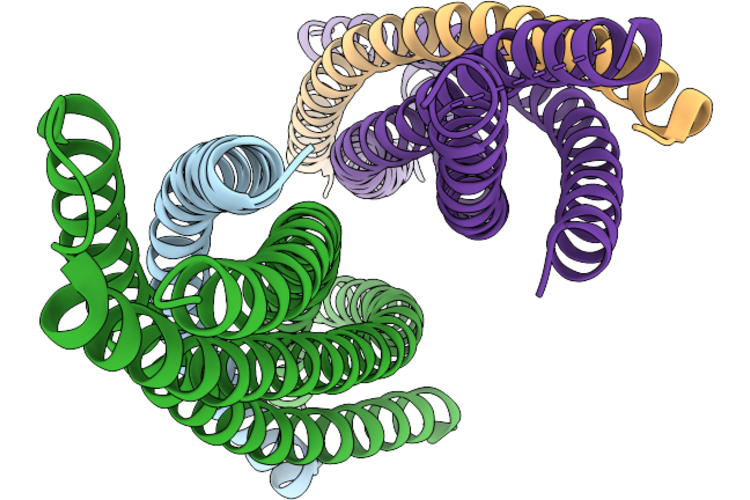
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.45 Å Release Date: 2026-05-13 Classification: MEMBRANE PROTEIN |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-05-13 Classification: MEMBRANE PROTEIN |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.52 Å Release Date: 2026-05-13 Classification: MEMBRANE PROTEIN |
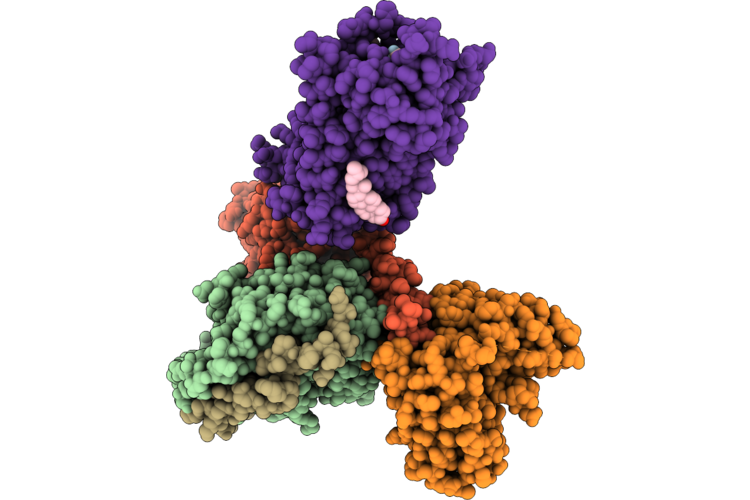 |
Organism: Rattus norvegicus, Bos taurus, Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: SIGNALING PROTEIN/IMMUNE SYSTEM Ligands: CLR |
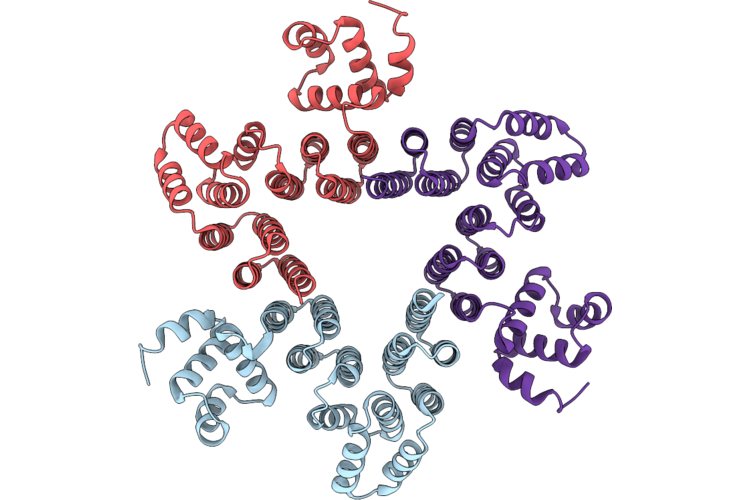 |
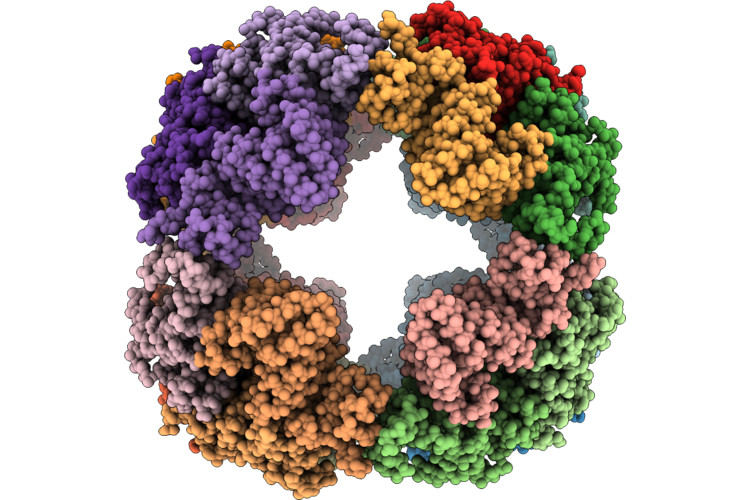
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-05-06 Classification: DE NOVO PROTEIN |
 |
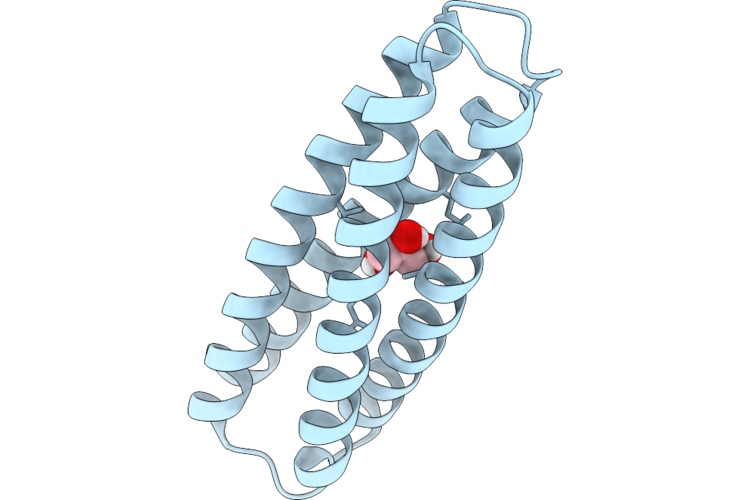
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: DE NOVO PROTEIN |
 |
X-Ray Crystal Structure Of A De Novo Designed Nile Red Binder, Sc-Apcc-4-Nr-1
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-05-06 Classification: DE NOVO PROTEIN Ligands: GOL |
 |
X-Ray Crystal Structure Of A De Novo Designed Arcyriaflavin A Binder, Sc-Apcc-4-A7F-1
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-05-06 Classification: DE NOVO PROTEIN Ligands: EDO |
 |
X-Ray Crystal Structure Of A De Novo Designed Sn38 Binder, Sc-Apcc-4-Sn38-1
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-05-06 Classification: DE NOVO PROTEIN Ligands: PEG, ETA |
 |
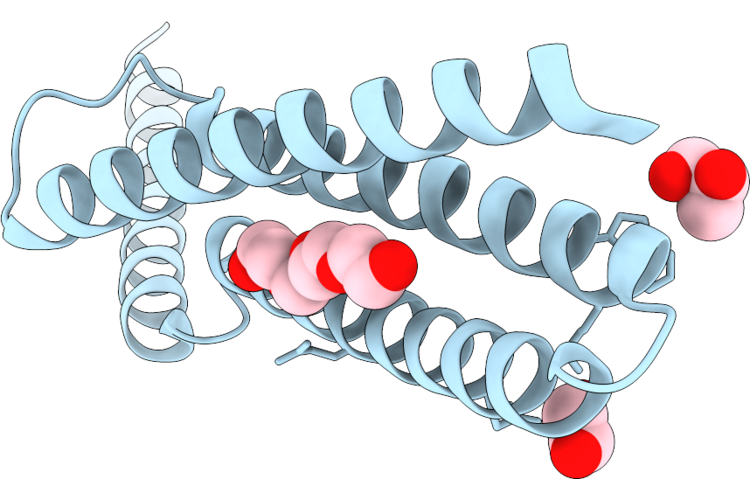
X-Ray Crystal Structure Of A De Novo Designed Lumiflavin Binder, Sc-Apcc-4-Lmf-1
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-05-06 Classification: DE NOVO PROTEIN |
 |
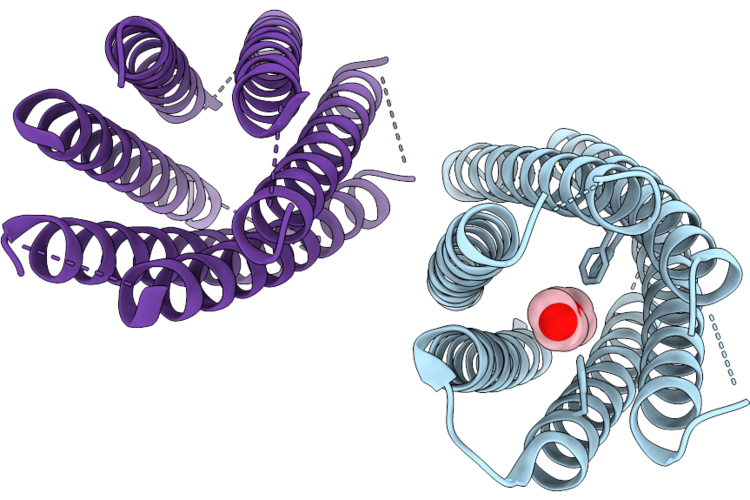
X-Ray Crystal Structure Of A De Novo Designed 1,6-Diphenylhexatriene Binder, Sc-Apcc-4-Dph-2
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-05-06 Classification: DE NOVO PROTEIN Ligands: 1PE, GOL, EDO |
 |
X-Ray Crystal Structure Of A De Novo Designed 1,6-Diphenylhexatriene And Nile Red Binder, Sc-Apcc-6-Dph-Nr
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:3.23 Å Release Date: 2026-05-06 Classification: DE NOVO PROTEIN Ligands: 1PE |

