Search Count: 1,984
All
Selected
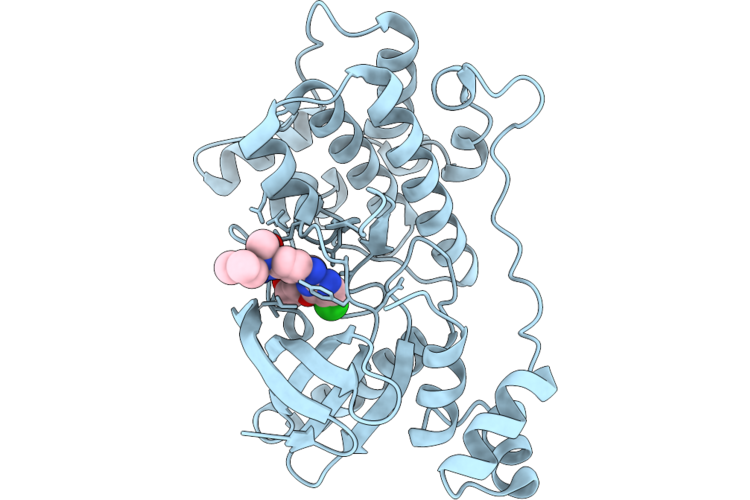 |
Crystal Structure Of The Nuak1-Mark3 Kinase Domain Chimera Bound With Small Molecule Inhibitor 2-Amino-N-(5-((5-Chloro-4-(((3R,3Ar,6R,6Ar)-6-Methoxyhexahydrofuro[3,2-B]Furan-3-Yl)Oxy)Pyrimidin-2-Yl)Amino)-2-((2-(Dimethylamino)Ethyl)(Methyl)Amino)Phenyl)Acetamide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: A1ER0 |
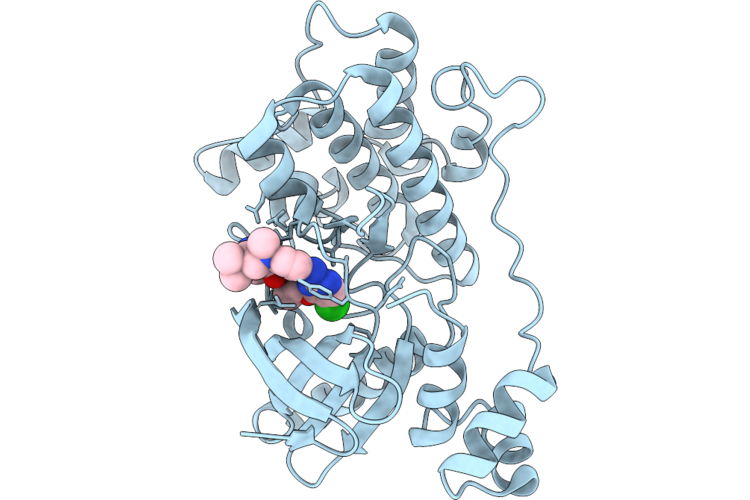 |
Crystal Structure Of The Nuak1-Mark3 Kinase Domain Chimera Bound With Small Molecule Inhibitor 4-((5-((5-Chloro-4-(((3R,3Ar,6R,6Ar)-6-Methoxyhexahydrofuro[3,2-B]Furan-3-Yl)Oxy)Pyrimidin-2-Yl)Amino)-2-((2-(Dimethylamino)Ethyl)(Methyl)Amino)Phenyl)Carbamoyl)-1-Methyl-3H-Pyrazol-1-Ium-3-Ide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: CYTOSOLIC PROTEIN Ligands: A1ER9 |
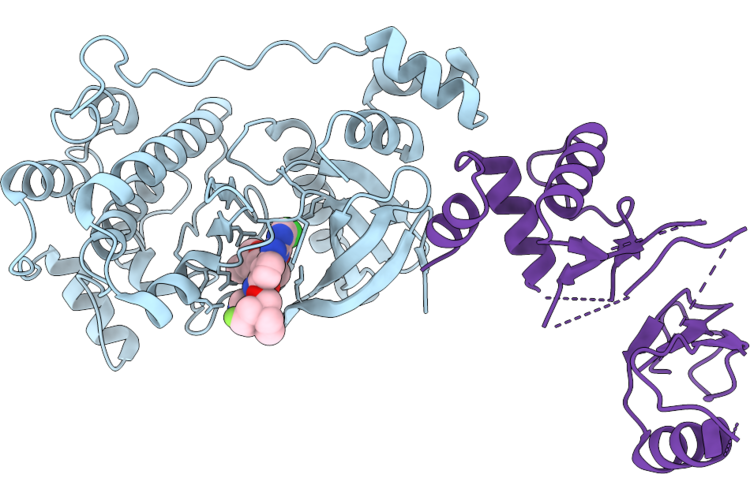 |
Crystal Structure Of The Nuak1-Mark3 Kinase Domain Chimera Bound With Small Molecule Inhibitor N-(5-((5-Chloro-4-(((3As,6R,6Ar)-6-Methoxy-3A,5,6,6A-Tetrahydrofuro[3,2-B]Furan-3-Yl)Oxy)Pyrimidin-2-Yl)Amino)-2-(((2R,7Ar)-2-Fluorotetrahydro-1H-Pyrrolizin-7A(5H)-Yl)Methoxy)Phenyl)-1-Methyl-1H-Pyrazole-4-Carboxamide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-22 Classification: CYTOSOLIC PROTEIN Ligands: A1ESF |
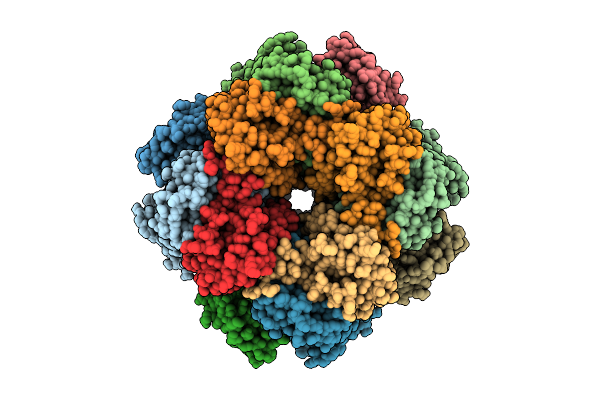 |
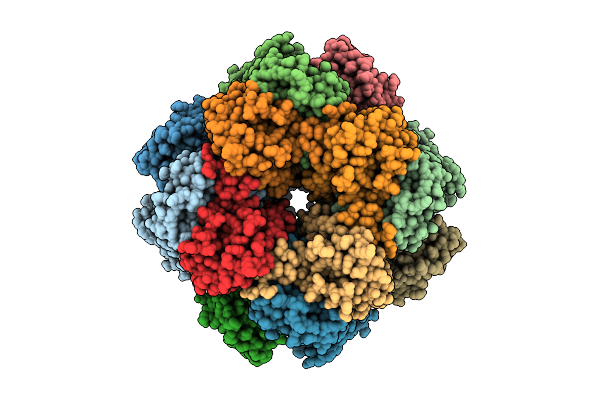
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.87 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN |
 |
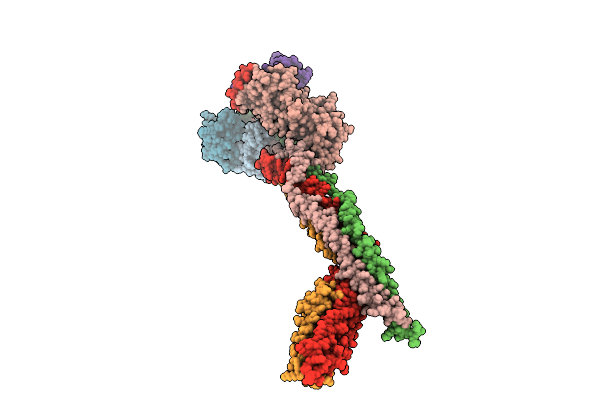
Organism: Arabidopsis thaliana
Method: ELECTRON MICROSCOPY Resolution:2.97 Å Release Date: 2026-04-08 Classification: PLANT PROTEIN |
 |
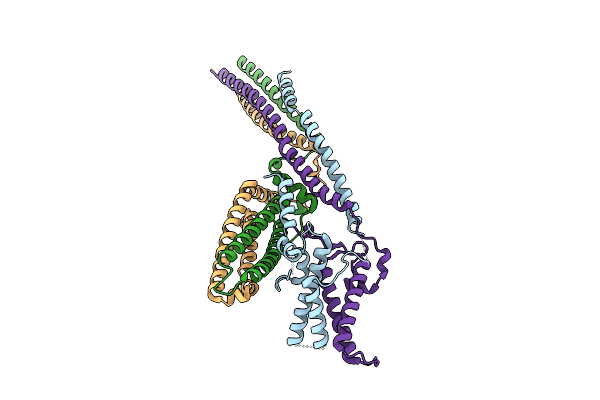
Cryo-Em Structure Of The Saccharomyces Cerevisiae Kmn Junction Complex Lacking The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
 |
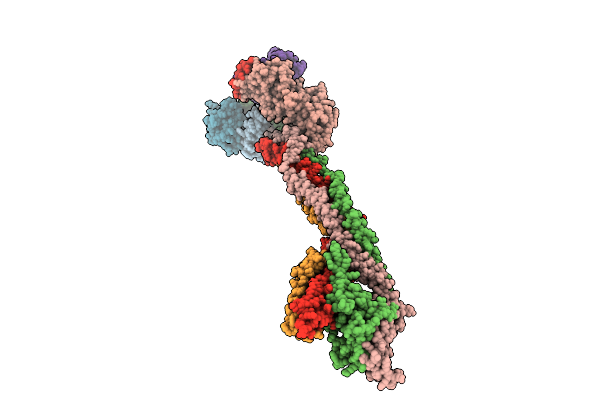
Cryo-Em Structure Of The Base Of The Saccharomyces Cerevisiae Kmn Junction Complex Containing The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
 |
Cryo-Em Structure Of The Saccharomyces Cerevisiae Kmn Junction Complex Containing The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae, Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
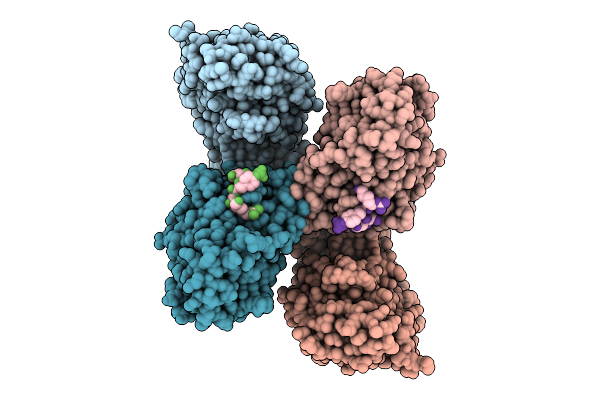 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-08 Classification: HYDROLASE |
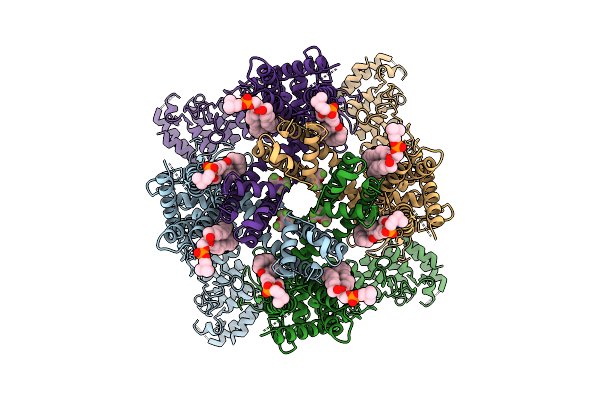 |
Cryo-Em Structure Of Human Trpv3 In Complex With Sevoflurane Determined In Msp2N2 Nanodisc
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: A1IPI, POV |
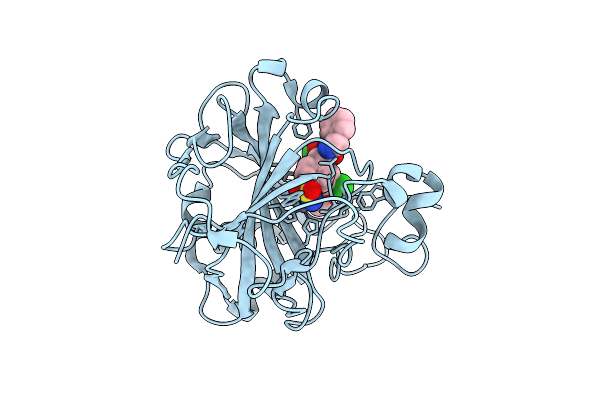 |
Crystal Structure Of Human Carbonic Anhydrase I In Complex With N-Benzyl-2-(2-Chloro-N-(3-Chloro-4-Methoxyphenyl)Acetamido)-2-(4-Sulfamoylphenyl)Acetamide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.31 Å Release Date: 2026-04-01 Classification: LYASE Ligands: ZN, A1JD1 |
 |
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Trp185Tyr/His280Ala With 2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-04-01 Classification: TRANSFERASE |
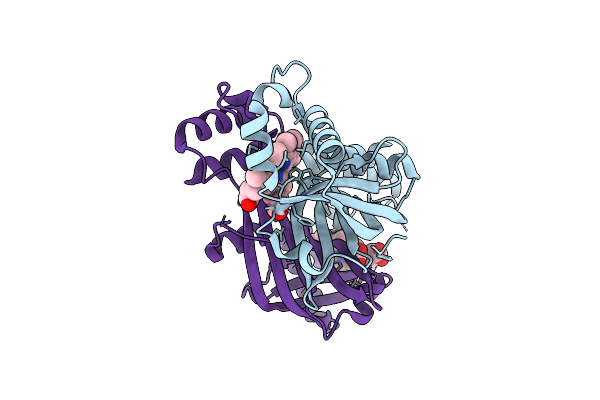 |
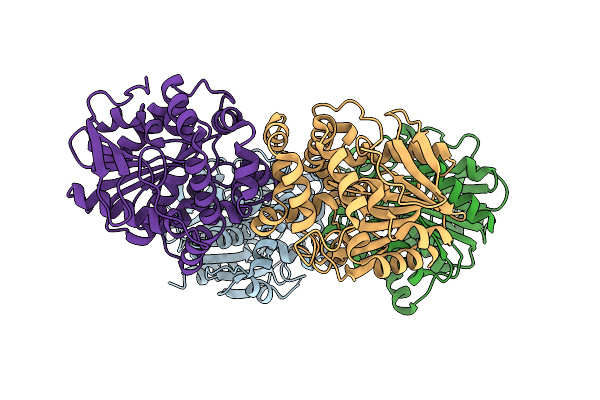
The Crystal Structure Of Paib From Bacillus Stearothermophilus Bound To Hem
Organism: Geobacillus kaustophilus hta426
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2026-04-01 Classification: TRANSCRIPTION Ligands: HEM |
 |
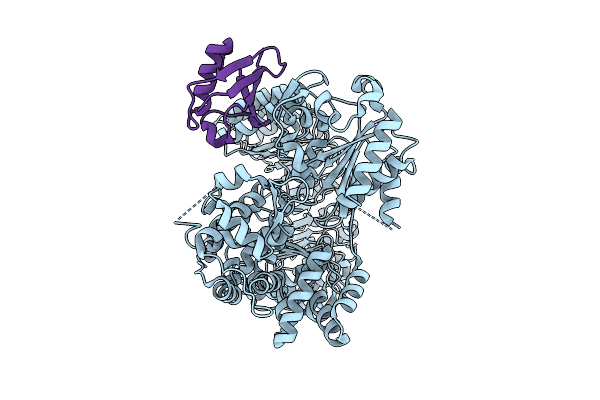
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Met/Trp185Tyr/His280Ala With 2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-03-25 Classification: TRANSFERASE |
 |
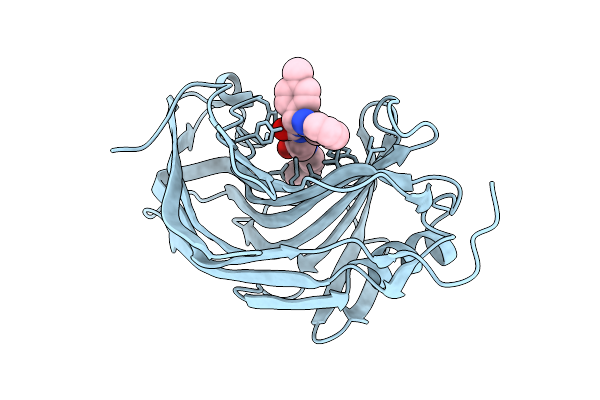
Organism: Escherichia coli k-12, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.49 Å Release Date: 2026-03-18 Classification: CYTOSOLIC PROTEIN |
 |
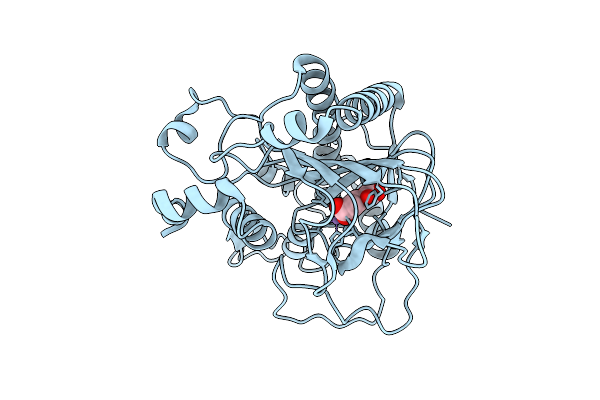
Crystal Structure Of The Pathogen-Secreted Apoplastic Gh12 Xyloglucan-Specific Endoglucanase Xeg1
Organism: Phytophthora sojae strain p6497
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: A1ENU |
 |
Fe(Ii)-2-Oxoglutarate-Dependent Pseudomonas Savastanoi Pv Phaseolicola 1449B In Complex With 2-Oxoglutarate
Organism: Pseudomonas savastanoi
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: MN, AKG |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: CYTOSOLIC PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: CYTOSOLIC PROTEIN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: CYTOSOLIC PROTEIN Ligands: ZN, GTP, MG, ADP |

