Search Count: 1,112
All
Selected
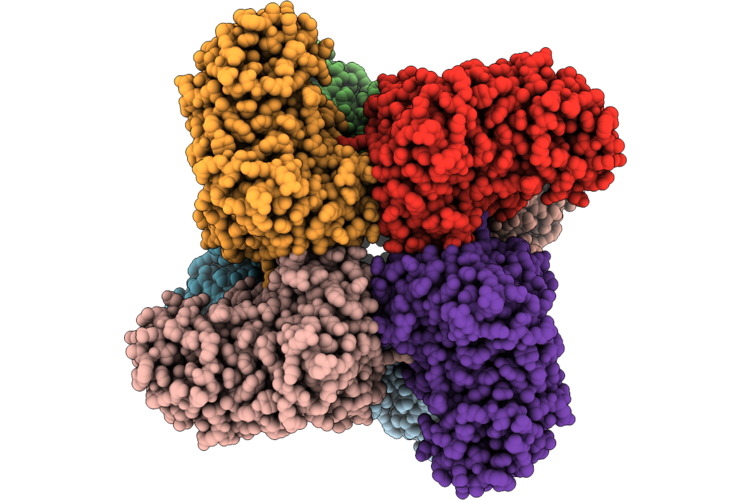 |
Cryo-Electron Microscopic Structure Of A Novel Amidohydrolase Adh3 Triple Mutation
Organism: Stenotrophomonas sp. cw117
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: HYDROLASE Ligands: ZN, 97U |
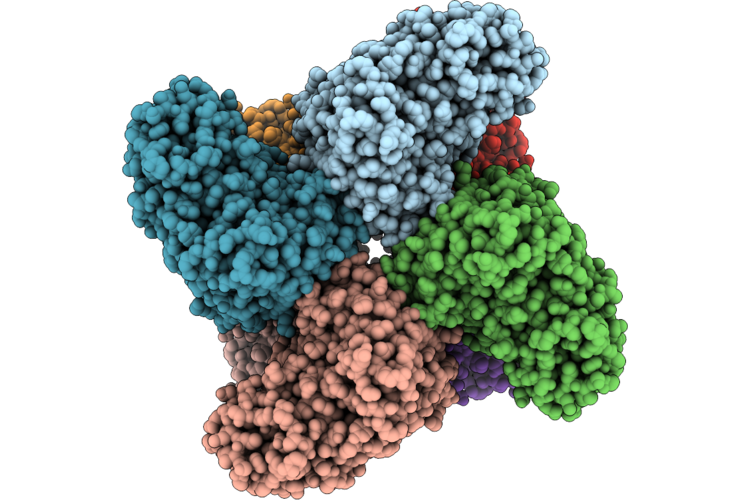 |
Cryo-Em Structure And Rational Engineering Of A Novel Efficient Ochratoxin A-Detoxifying Amidohydrolase
Organism: Pseudoxanthomonas wuyuanensis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN, 97U |
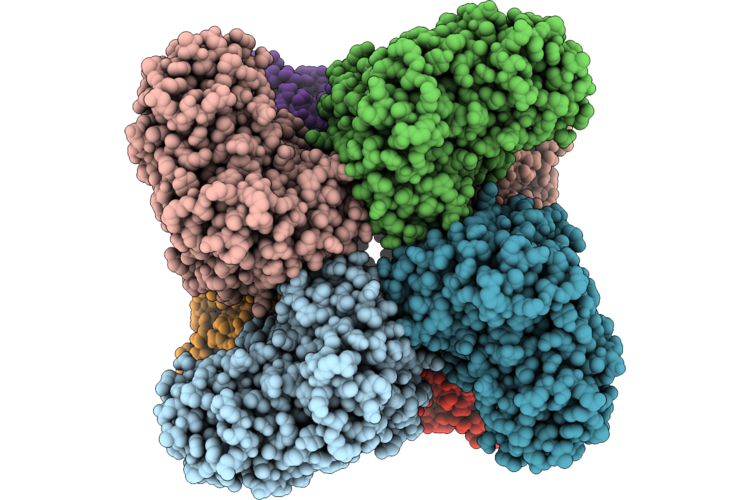 |
Cryo-Electron Microscopic Structure Of A Novel Amidohydrolase With Three Mutations
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN, 97U |
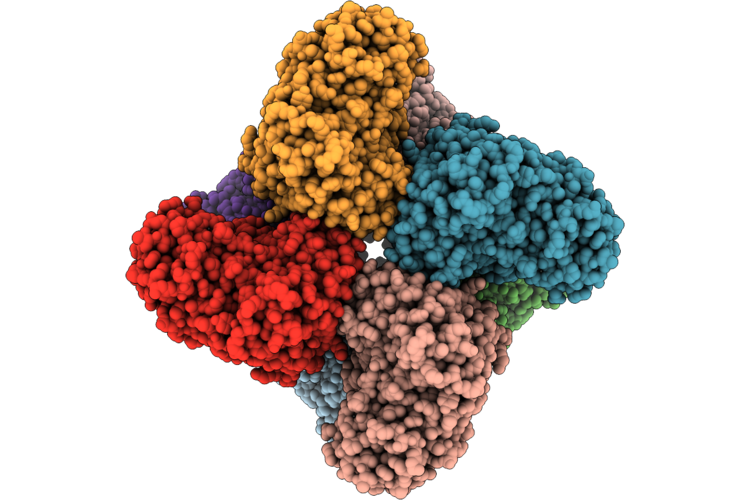 |
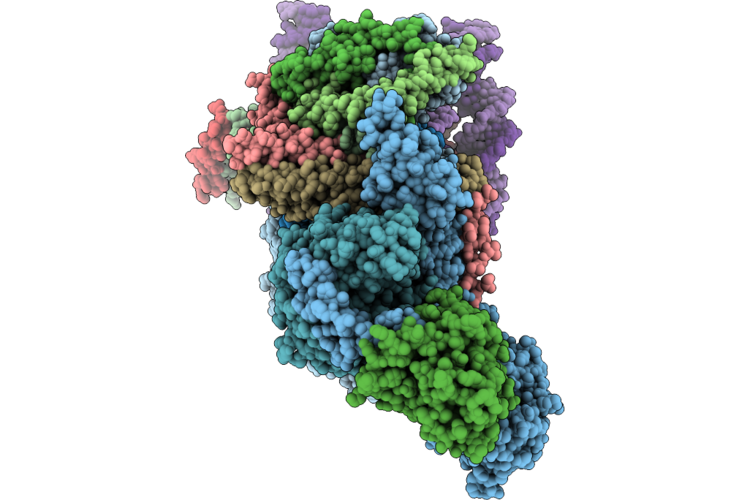
Cryo-Electron Microscopic Structure Of A Highly Efficient Ochratoxin Detoxification Enzyme Lladh
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN |
 |
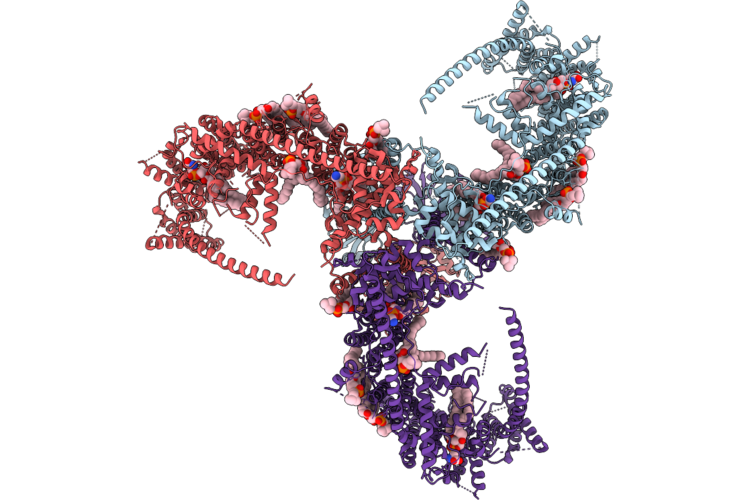
Cryo-Em Structure Of The Histone Deacetylase Complex Rpd3L In Complex With Mono-Nucleosome
Organism: Saccharomyces cerevisiae s288c, Xenopus laevis, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: GENE REGULATION/DNA Ligands: ZN |
 |
Organism: Drosophila
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: PLX, PEE, P5S |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-15 Classification: STRUCTURAL PROTEIN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2026-04-01 Classification: CELL INVASION Ligands: GDP, A1D9L, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-01 Classification: CELL INVASION Ligands: GDP, MG |
 |
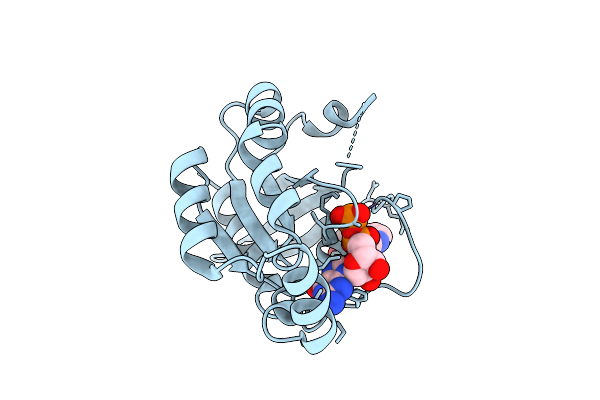
Structure Of The Rhoa Y42C Mutant Featuring Thr37 Coordination Of Magnesium Ion
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-01 Classification: CELL INVASION Ligands: GDP, MG |
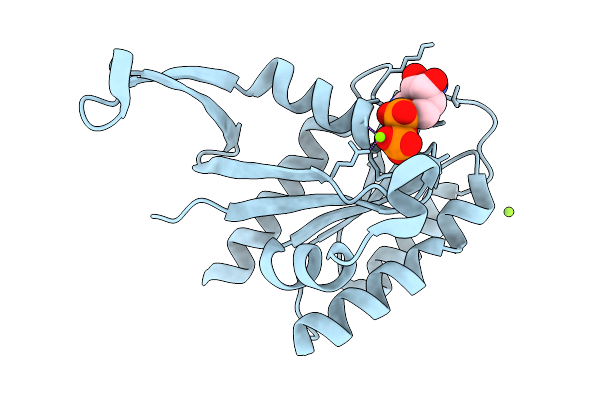 |
Crystal Structure Of Gdp-Bound Kras G12D/I55E: Suppressing G12D Oncogenicity Via Second-Site I55E Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.23 Å Release Date: 2026-03-25 Classification: ONCOPROTEIN Ligands: GDP, MG, CL |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.87 Å Release Date: 2026-03-18 Classification: TRANSLATION Ligands: AR6 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-03-18 Classification: RNA BINDING PROTEIN Ligands: AR6 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-18 Classification: IMMUNE SYSTEM Ligands: GOL, NAG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-03-18 Classification: IMMUNE SYSTEM Ligands: GOL, NAG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2026-03-18 Classification: IMMUNE SYSTEM Ligands: GOL, NAG, SO4 |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2026-03-18 Classification: IMMUNE SYSTEM Ligands: MLI, GOL, NA, NAG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2026-03-18 Classification: IMMUNE SYSTEM Ligands: GOL, MLI, NAG |
 |
Cryoem Structure Of Toxin B (Tcdb) From Clostridioides Difficile Complexed With Taurochenodeoxycholic Acid (Tcdca)
Organism: Clostridioides difficile
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: TOXIN Ligands: TUD |
 |
Cryoem Structure Of Toxin B (Tcdb) From Clostridioides Difficile Complexed With Methyl Cholate
Organism: Clostridioides difficile
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: TOXIN Ligands: A1CBA |

