Search Count: 741
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: A1ES7 |
 |
Organism: Chromobacterium violaceum
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
Organism: Streptomyces sp. nl15-2k
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-08 Classification: LIGASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-08 Classification: LIGASE Ligands: A1C4E, GOL |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2026-04-08 Classification: LIGASE Ligands: A1C4F, GOL |
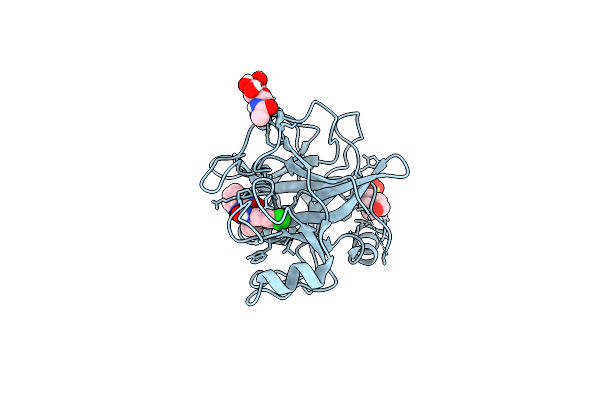 |
N-Alkyl &Amp; N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasma Kallikrein Inhibitors Complex With Compound 15 ((3'R)-1'-(5-Amino-1-Phenyl-1H-Pyrazole-4-Carbonyl)-6-Chloro-5-Fluorospiro[[3,1]Benzoxazine-4,3'-Piperidin]-2(1H)-One)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-04-01 Classification: HYDROLASE/INHIBITOR Ligands: MES, A1C6X, NAG |
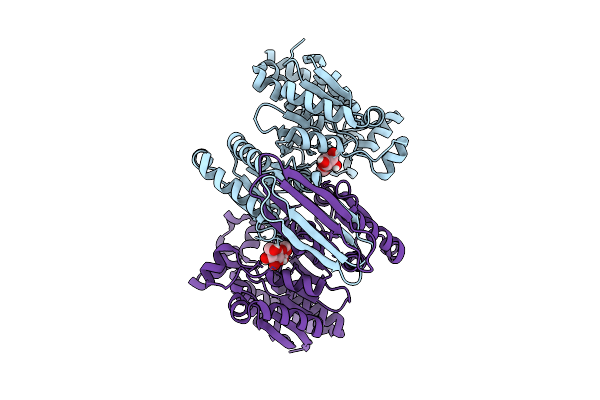 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:3.08 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: A1EOQ |
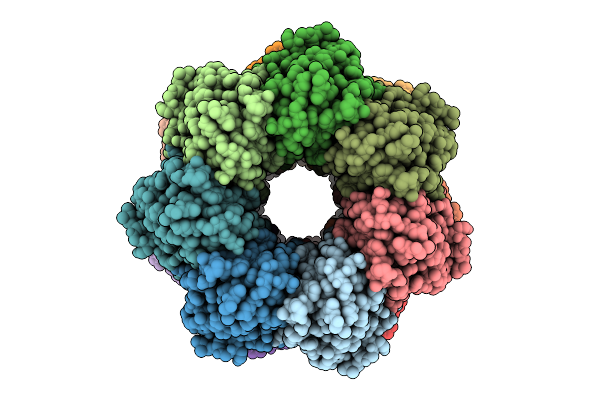 |
Organism: Pseudomonas plecoglossicida nbrc 103162 = dsm 15088
Method: ELECTRON MICROSCOPY Resolution:3.07 Å Release Date: 2026-03-18 Classification: HYDROLASE |
 |
N-Alkyl & N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasma Kallikrein Inhibitors
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.78 Å Release Date: 2026-03-11 Classification: HYDROLASE/INHIBITOR Ligands: A1C6O |
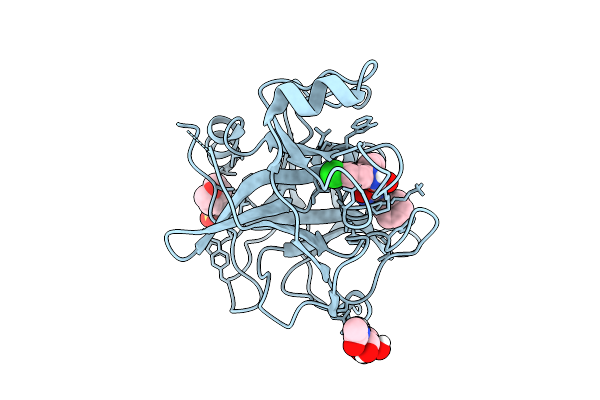 |
N-Alkyl &Amp; N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasmakallikrein Inhibitors Complex With Compound 4 ((3'R)-1'-(5-Amino-1-Benzyl-1H-Pyrazole-4-Carbonyl)-6-Chloro-5-Fluorospiro[[3,1]Benzoxazine-4,3'-Piperidin]-2(1H)-One)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-03-11 Classification: HYDROLASE/INHIBITOR Ligands: MES, A1C6Q, NAG |
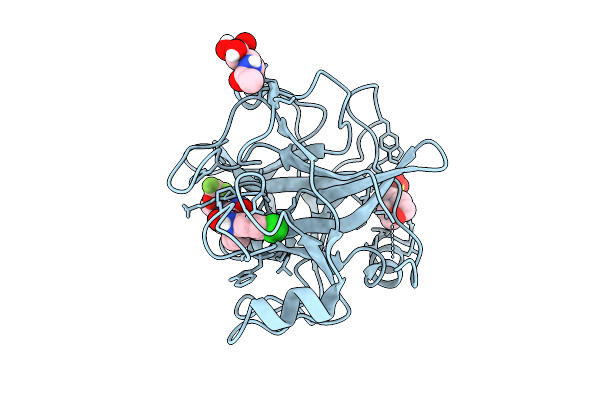 |
N-Alkyl &Amp; N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasma Kallikrein Inhibitors Compound 25 ((3'R)-1'-{(1P)-5-Amino-1-[2-(Trifluoromethoxy)Phenyl]-1H-Pyrazole-4-Carbonyl}-6-Chloro-5-Fluorospiro[[3,1]Benzoxazine-4,3'-Piperidin]-2(1H)-One)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-03-11 Classification: HYDROLASE/INHIBITOR Ligands: MES, A1C6Z, NAG |
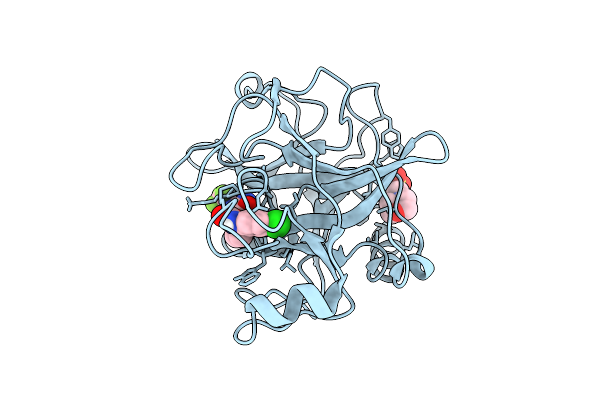 |
N-Alkyl &Amp; N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasma Kallikrein Inhibitors Compound 13 ((3'R)-1'-{5-Amino-1-[(2S)-1,1,1-Trifluorobutan-2-Yl]-1H-Pyrazole-4-Carbonyl}-6-Chloro-5-Fluorospiro[[3,1]Benzoxazine-4,3'-Piperidin]-2(1H)-One)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-03-11 Classification: HYDROLASE/INHIBITOR Ligands: MES, A1C7M |
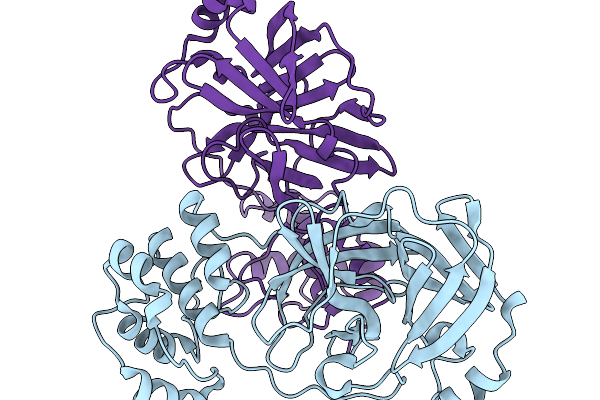 |
Organism: Swine acute diarrhea syndrome coronavirus
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2026-02-18 Classification: VIRAL PROTEIN |
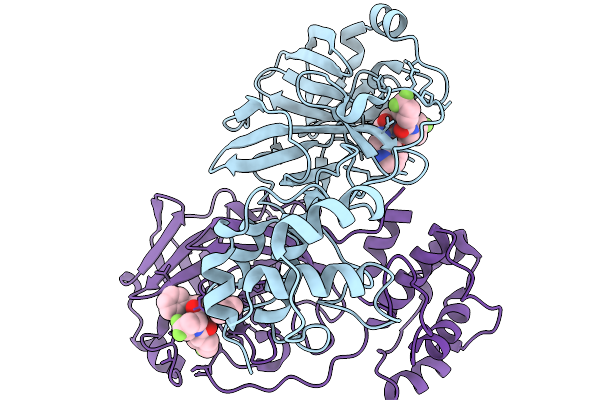 |
Organism: Swine acute diarrhea syndrome coronavirus
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-02-18 Classification: VIRAL PROTEIN Ligands: OF9 |
 |
Crystal Structure Of Sads-Cov Main Protease (Lys35Val/Cys224Ser) In Complex With Sy110
Organism: Swine acute diarrhea syndrome coronavirus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-02-18 Classification: VIRAL PROTEIN Ligands: LVX, P33, PG4 |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-02-18 Classification: VIRAL PROTEIN |
 |
Organism: Swine acute diarrhea syndrome coronavirus
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2026-02-18 Classification: VIRAL PROTEIN |
 |
Organism: Mycobacterium tuberculosis h37rv
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-02-11 Classification: TRANSFERASE Ligands: MPD |
 |
Organism: Legionella pneumophila
Method: X-RAY DIFFRACTION Resolution:2.23 Å Release Date: 2026-01-21 Classification: TRANSFERASE Ligands: IHP |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-01-14 Classification: MEMBRANE PROTEIN Ligands: CLR, A1EK5 |

