Search Count: 1,046
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: CU, ZN |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.37 Å Release Date: 2026-04-01 Classification: OXIDOREDUCTASE Ligands: CU, ZN |
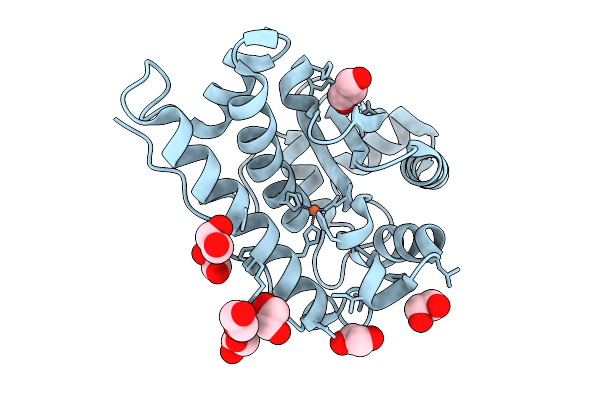 |
Crystal Structure Of A Fe Superoxide Dismutase From The Acenetobacter Baumannii (Ab) At 1.45 A
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-03-11 Classification: METAL BINDING PROTEIN Ligands: FE, GOL, EDO |
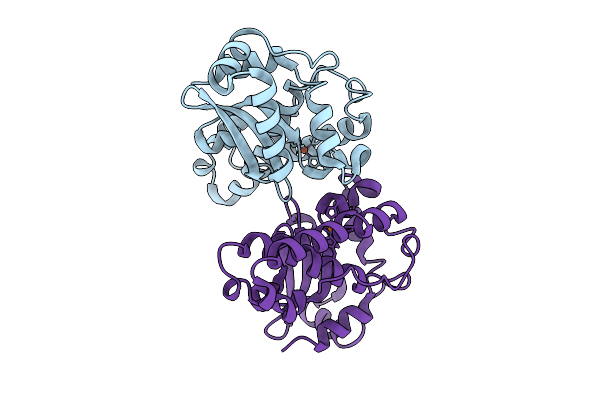 |
Organism: Listeria monocytogenes
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: FE |
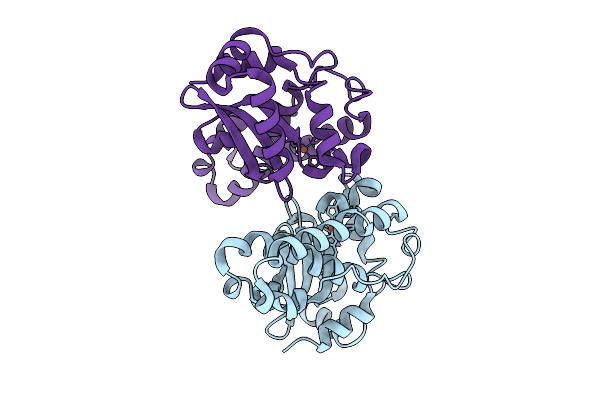 |
Organism: Listeria monocytogenes
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: FE |
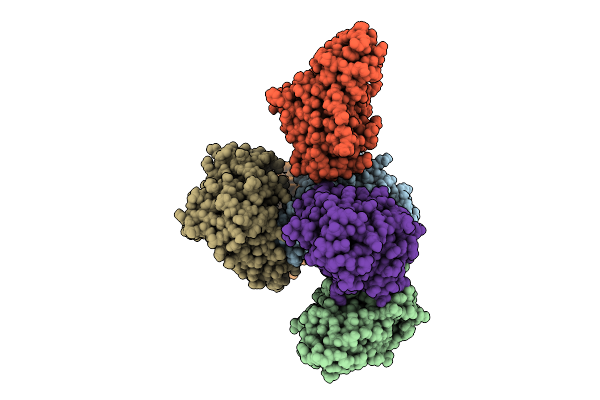 |
Organism: Neisseria gonorrhoeae
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-11 Classification: OXIDOREDUCTASE Ligands: FE |
 |
Cryo-Em Structure Of The Chromera Velia Psi Supercomplex At 1.84 Angstrom Resolution
Organism: Chromera velia
Method: ELECTRON MICROSCOPY Resolution:1.84 Å Release Date: 2026-01-28 Classification: PHOTOSYNTHESIS Ligands: FE, CLA, CL0, SF4, PQN, XAT, LMG, DGD, LMT, BCR, AV0, A1L6D, SQD, A1I05 |
 |
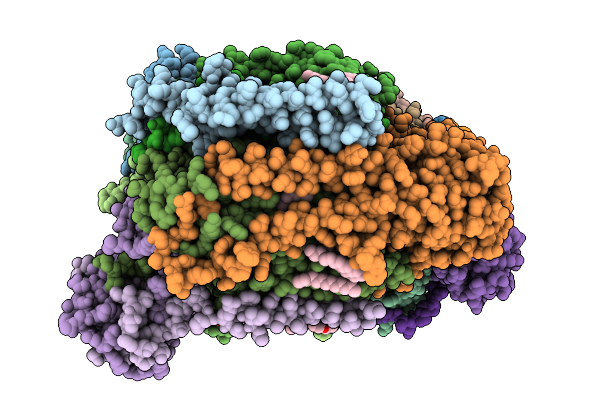
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:4.74 Å Release Date: 2026-01-28 Classification: PROTEIN FIBRIL |
 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: PROTEIN FIBRIL |
 |
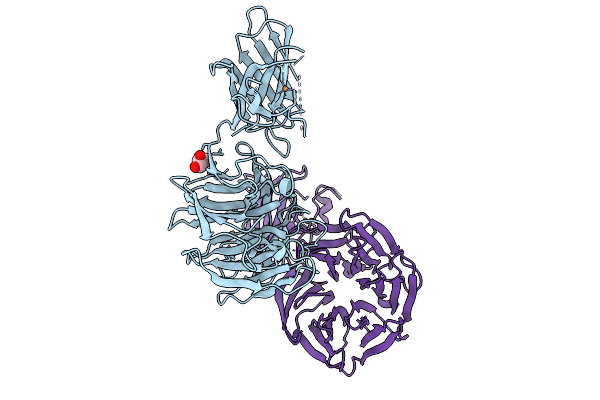
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-10-29 Classification: PROTEIN FIBRIL |
 |
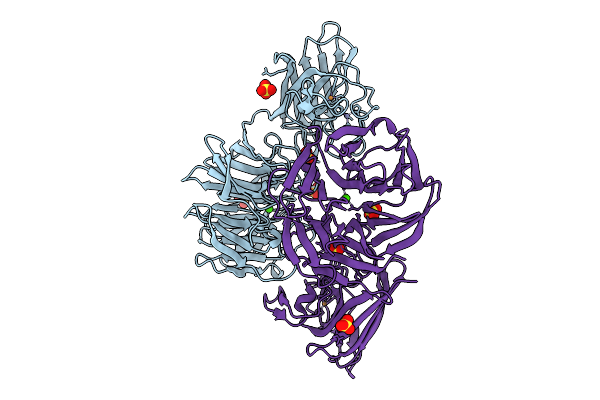
Organism: Deinococcus radiodurans r1 = atcc 13939 = dsm 20539
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-09-24 Classification: METAL BINDING PROTEIN Ligands: CU, GOL |
 |
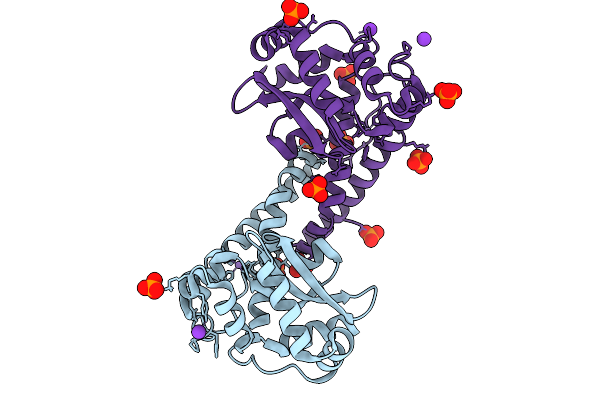
Organism: Deinococcus radiodurans r1 = atcc 13939 = dsm 20539
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-09-24 Classification: METAL BINDING PROTEIN Ligands: ZN, CU, CA, SO4, CL |
 |
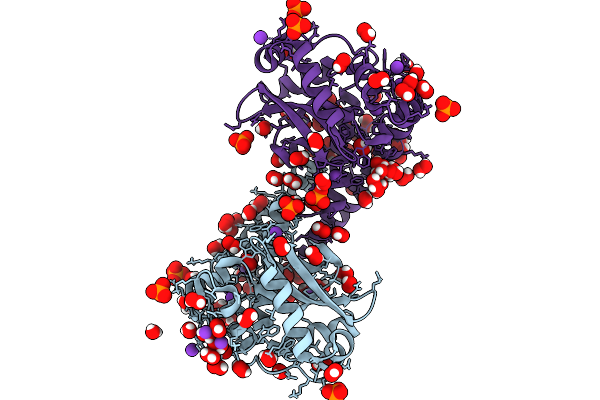
Finding The Exit Route Of Hydrogen Peroxide From The Manganese Superoxide Dismutase (Mnsod) Active Site
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: K, PO4, MN |
 |
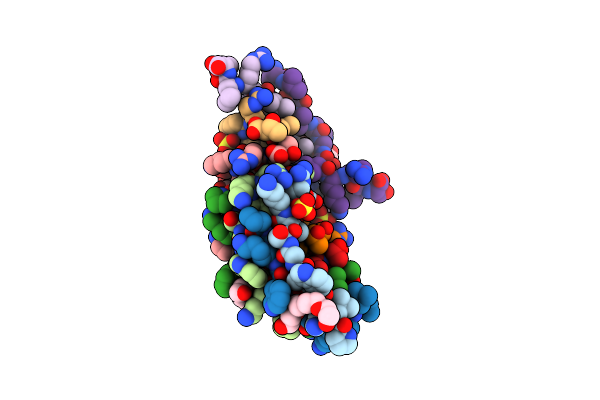
Finding The Exit Route Of Hydrogen Peroxide From The Manganese Superoxide Dismutase (Mnsod) Active Site
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.33 Å Release Date: 2025-08-27 Classification: OXIDOREDUCTASE Ligands: PEO, PO4, K, MN |
 |
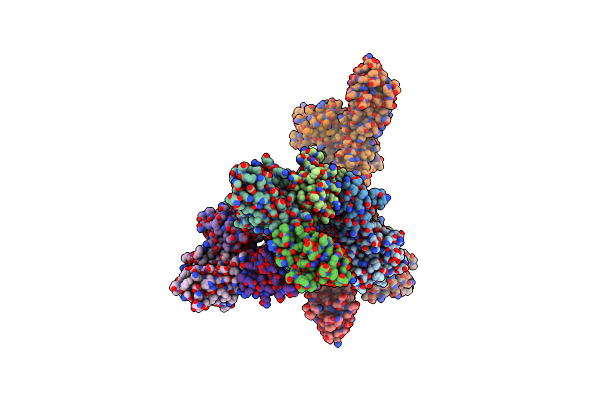
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2025-06-04 Classification: PROTEIN FIBRIL Ligands: SO4, PG4, PEG |
 |
Organism: Homo sapiens, Vicugna pacos, Camelus dromedarius
Method: X-RAY DIFFRACTION Resolution:3.28 Å Release Date: 2025-05-21 Classification: OXIDOREDUCTASE/IMMUNE SYSTEM Ligands: ZN |
 |
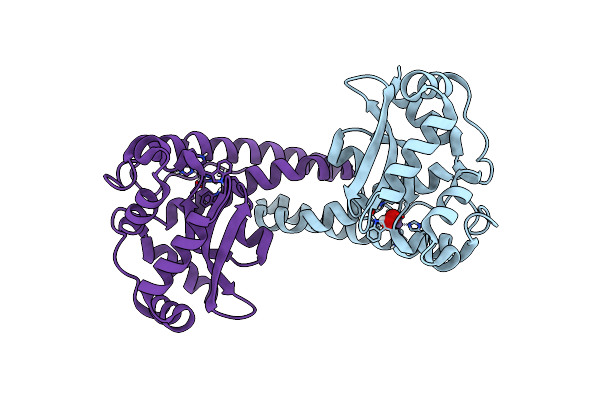
Organism: Mycolicibacterium smegmatis mc2 155
Method: ELECTRON MICROSCOPY Release Date: 2025-04-09 Classification: OXIDOREDUCTASE Ligands: FES, 9YF, PLM, HEM, CDL, A1IRE, HEC, MQ9, 9Y0, CU, HEA, CA, MQ7 |
 |
Organism: Homo sapiens
Method: NEUTRON DIFFRACTION Resolution:2.30 Å Release Date: 2025-03-12 Classification: OXIDOREDUCTASE Ligands: MN, PEO |
 |
Organism: Homo sapiens
Method: NEUTRON DIFFRACTION Resolution:2.50 Å Release Date: 2025-03-12 Classification: OXIDOREDUCTASE Ligands: MN |

