Search Count: 55,446
All
Selected
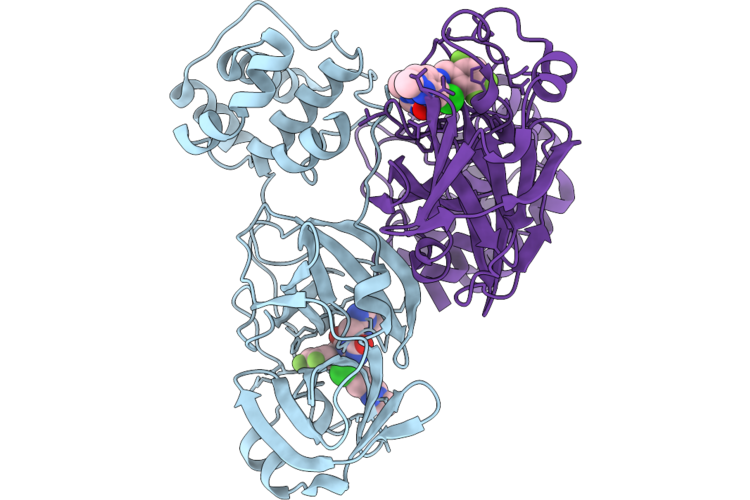 |
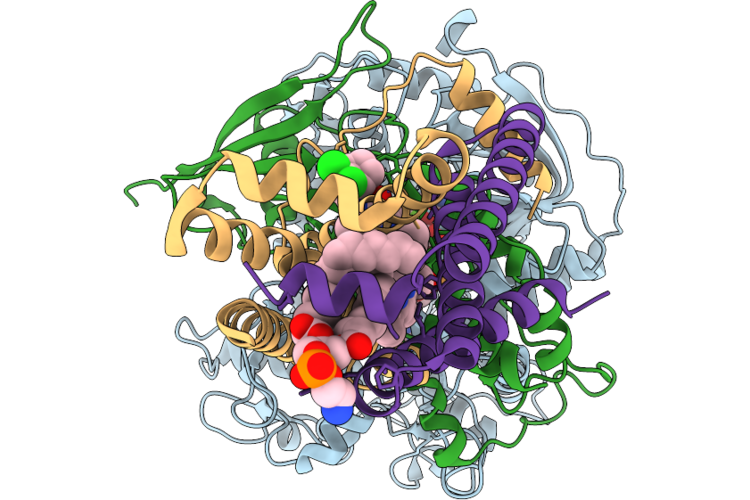
Room Temperature X-Ray Structure Of Sars Cov-2 Main Protease Intermediate Precursor With Ensitrelvir (Esv)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-05-06 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: 7YY |
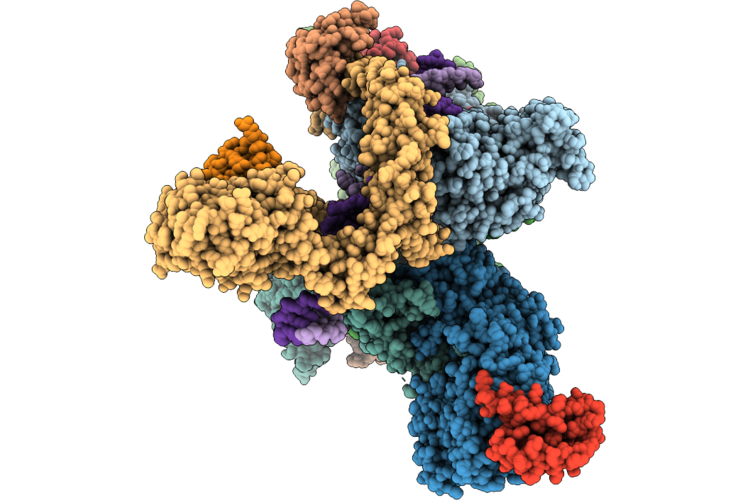 |
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Composite Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.32 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
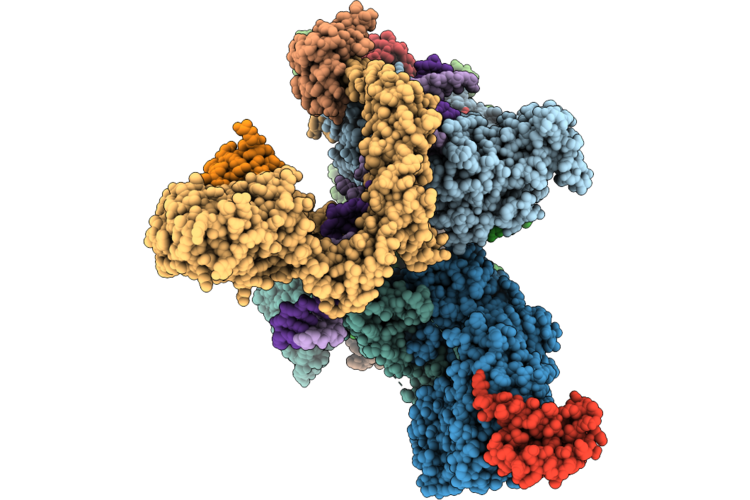 |
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Consensus Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.91 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
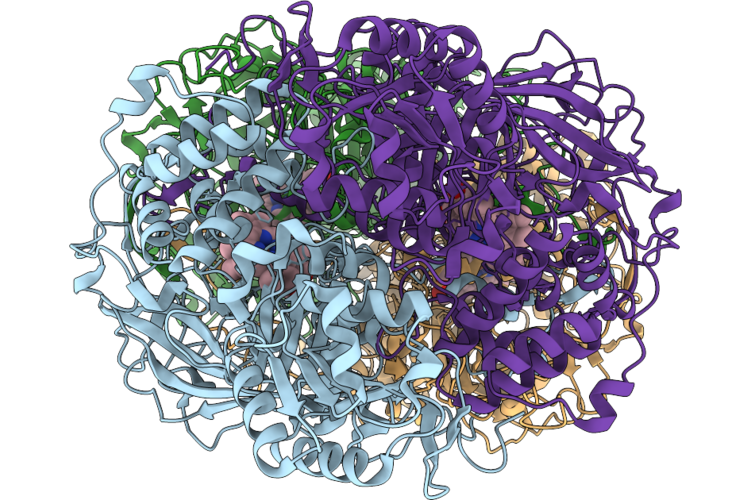 |
Organism: Escherichia coli bl21(de3)
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
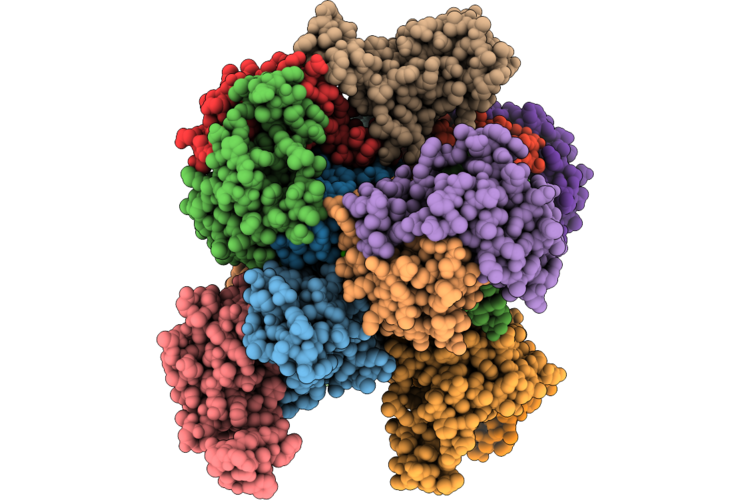
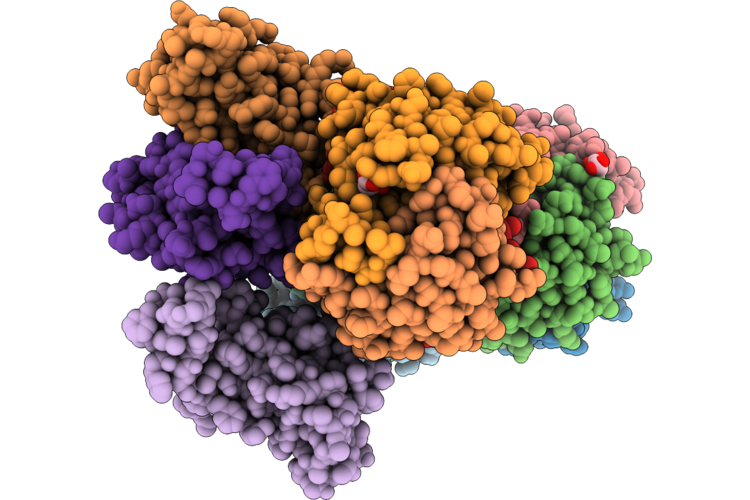
The Cryo-Em Structure Of Human Succinate Dehydrogenase In Complex With Benzovindiflupyr
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FAD, FES, SF4, F3S, A1EGM, HEM, PEV |
 |
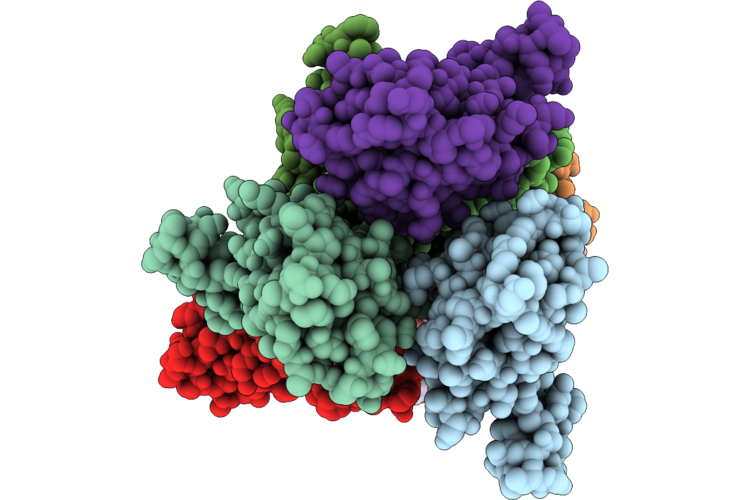
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2026-05-06 Classification: CYTOKINE Ligands: CA |
 |
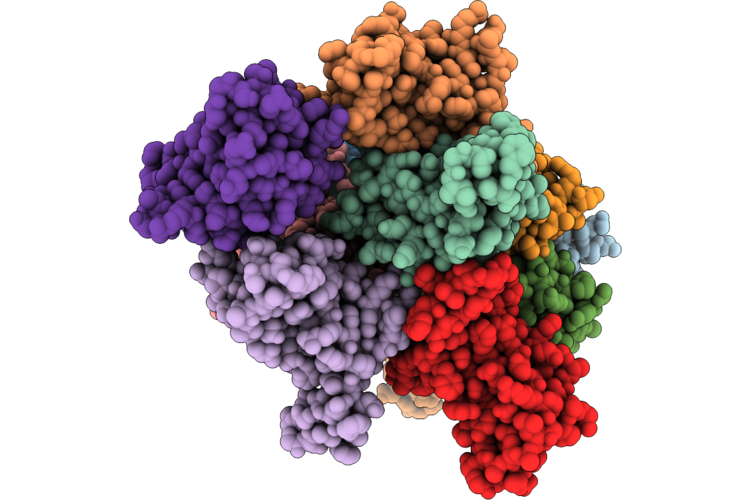
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2026-05-06 Classification: CYTOKINE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.53 Å Release Date: 2026-05-06 Classification: CYTOKINE Ligands: TRS |
 |
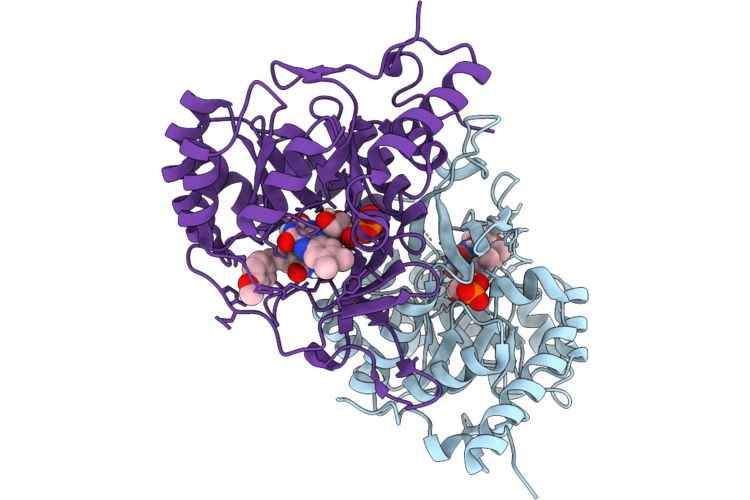
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-05-06 Classification: CYTOKINE Ligands: MLI |
 |
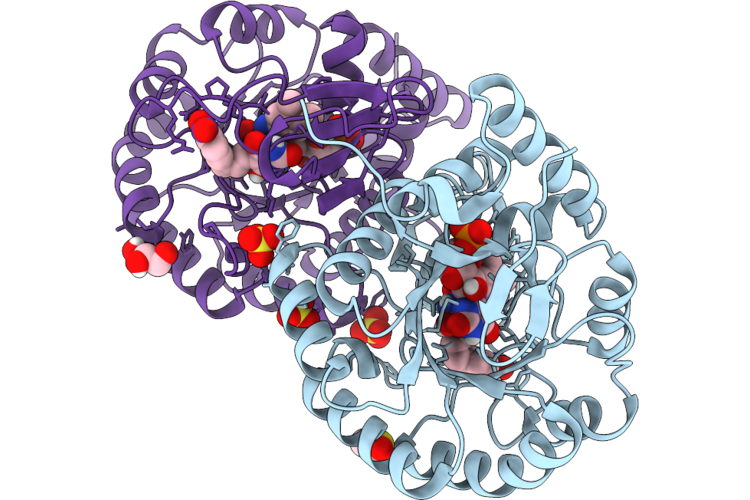
Crystal Structure Of Dihydroorotate Dehydrogenase From Leishmania Brasiliensis In Complex With 5-[(E)-3-(P-Methoxyphenyl)-2-Propenylidene]-2,4,6(1H,3H,5H)-Pyrimidinetrione
Organism: Leishmania braziliensis
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FMN, 5TI |
 |
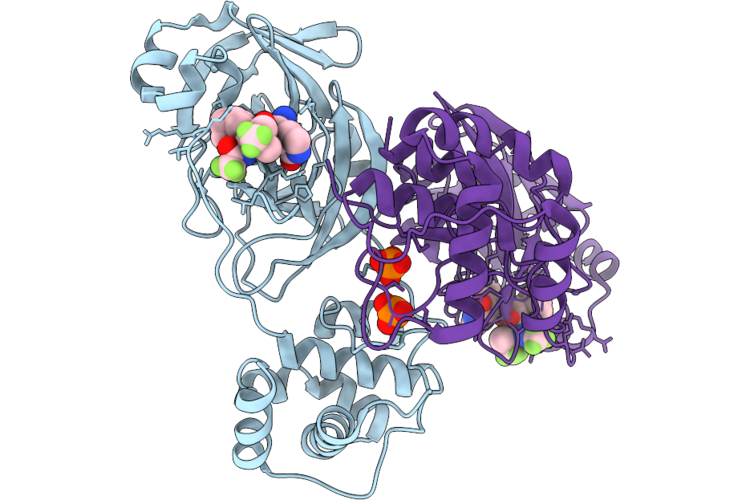
Crystal Structure Of Dihydroorotate Dehydrogenase From Leishmania Brasiliensis In Complex With (E)-5-(3-(4-Nitrophenyl)Allylidene)Pyrimidine-2,4,6(1H,3H,5H)-Trione
Organism: Leishmania braziliensis
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FMN, DMS, A1CAU, SO4, GOL |
 |
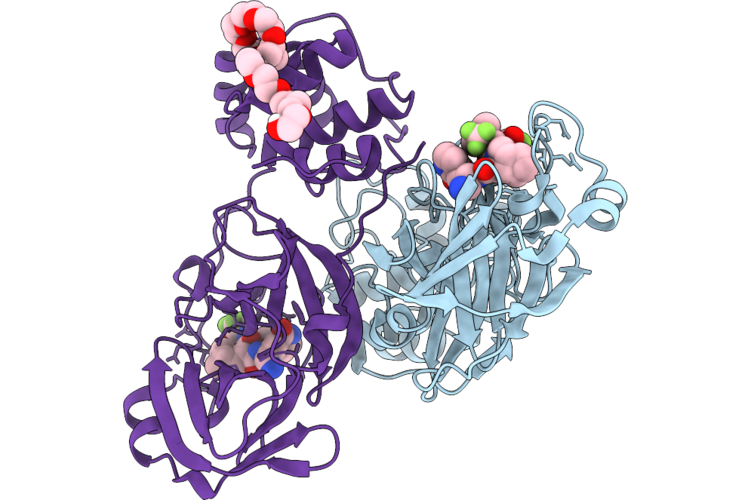
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-05-06 Classification: VIRAL PROTEIN Ligands: A1CHJ, PO4 |
 |
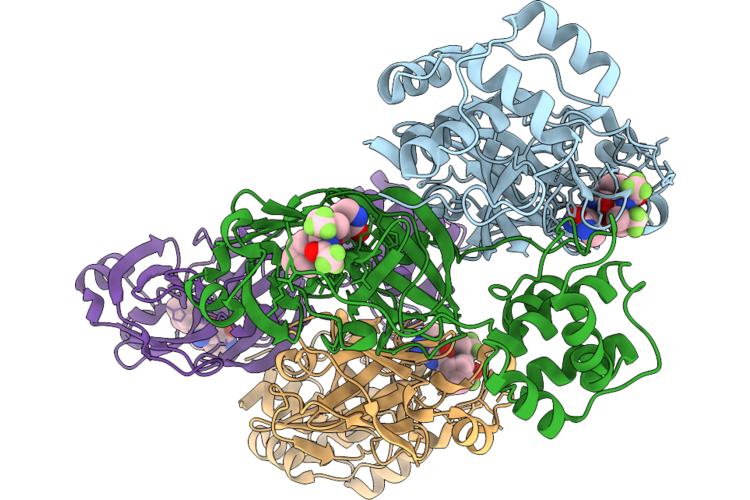
Organism: Human coronavirus 229e
Method: X-RAY DIFFRACTION Resolution:1.32 Å Release Date: 2026-05-06 Classification: VIRAL PROTEIN Ligands: A1CHJ, 12P |
 |
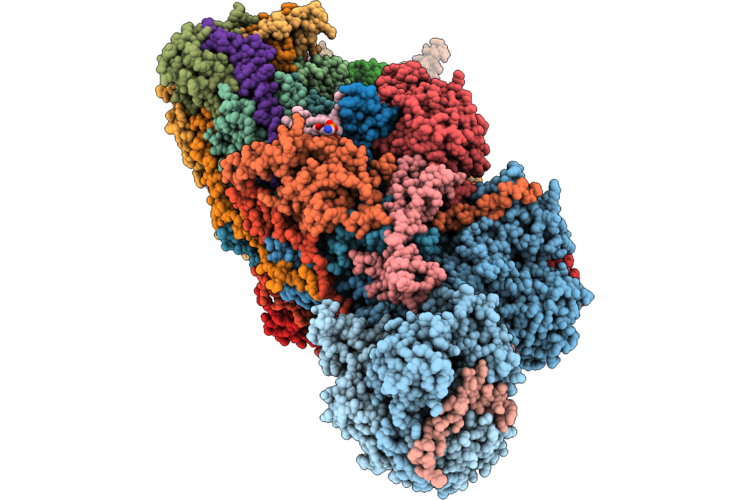
Organism: Middle east respiratory syndrome-related coronavirus
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-05-06 Classification: VIRAL PROTEIN Ligands: A1CHJ |
 |
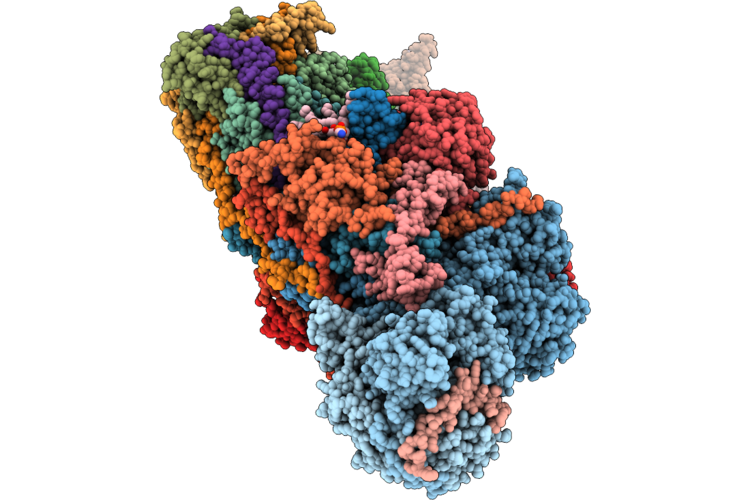
Organism: Ovis aries
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: 3PE, PC1, DGT, MG, MYR, ZMP, SF4, FMN, NAI, FES, K, A1JBT, ZN, NDP |
 |
Organism: Ovis aries
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: 3PE, PC1, A1JBT, SF4, DGT, MG, MYR, ZMP, FMN, NAI, FES, K, ZN, NDP |
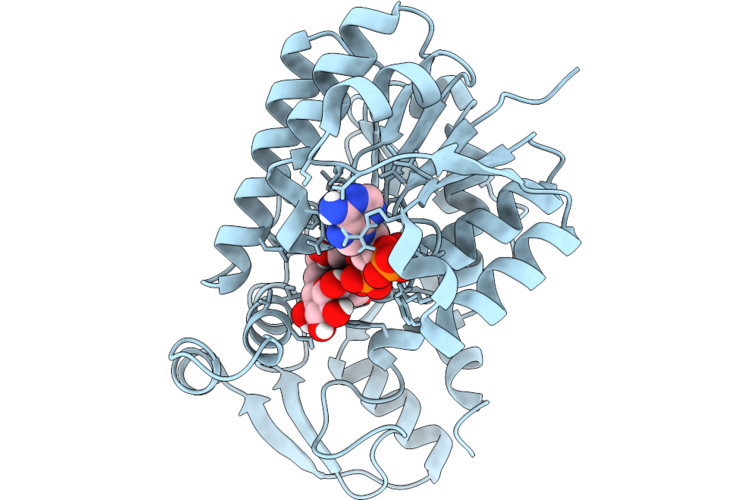 |
Crystal Structure Of An Nta622L Variant In Complex With Nadph And Nicotinic Acid N-Glucoside
Organism: Nicotiana tabacum
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-06 Classification: PLANT PROTEIN Ligands: A1JBN, NDP |
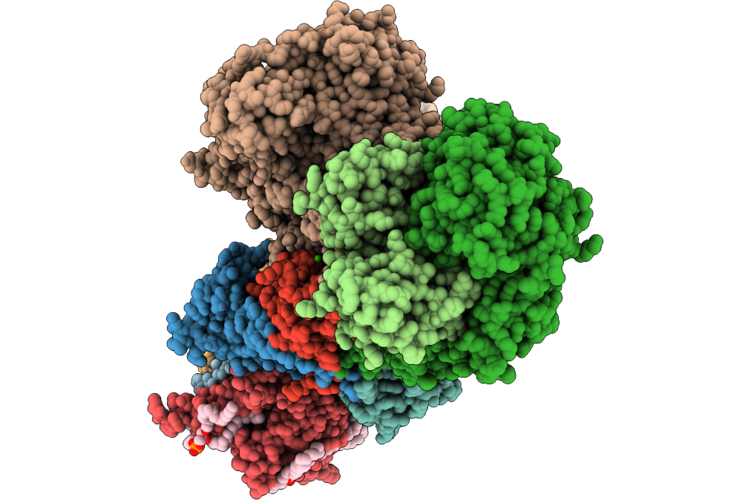 |
Organism: Escherichia coli bw25113, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: FES, CA, FMN, SF4, 3PE, LFA, CDL, TRD, UQ8 |
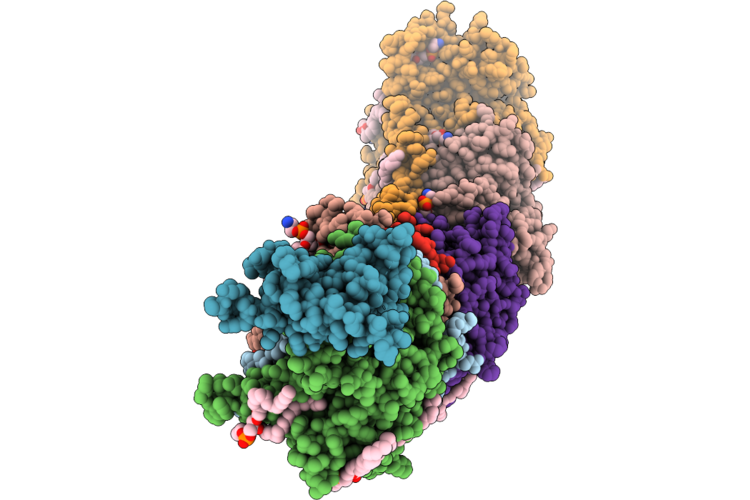 |
Organism: Escherichia coli bw25113
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: 3PE, LFA, CDL, TRD, UQ8 |
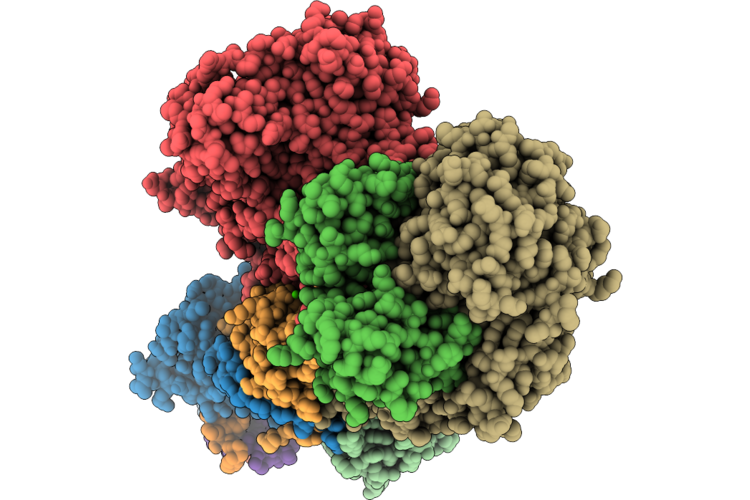 |
Organism: Escherichia coli bw25113
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: FES, CA, FMN, SF4 |

