Search Count: 259
All
Selected
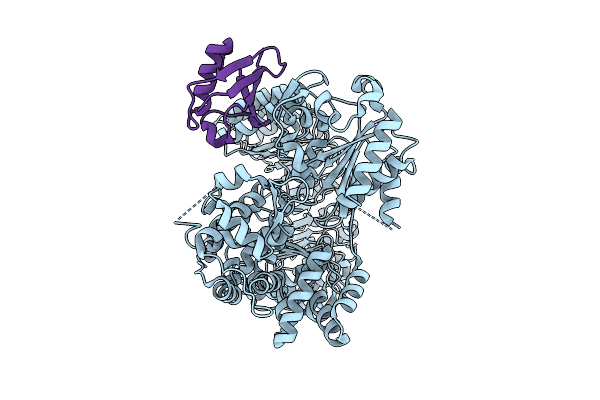 |
Organism: Escherichia coli k-12, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.49 Å Release Date: 2026-03-18 Classification: CYTOSOLIC PROTEIN |
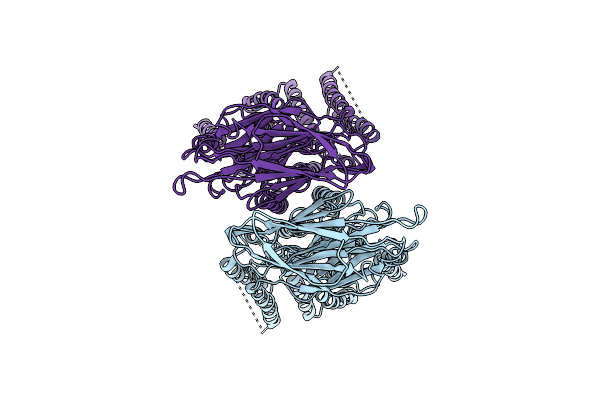 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN |
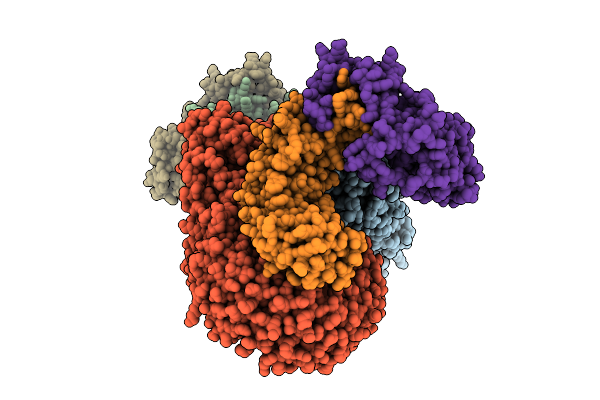 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: CYTOSOLIC PROTEIN |
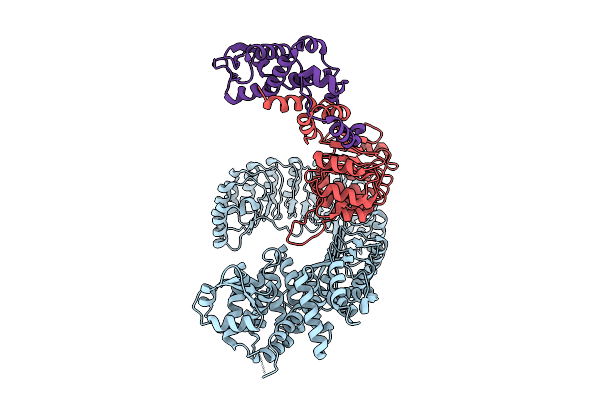 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: CYTOSOLIC PROTEIN |
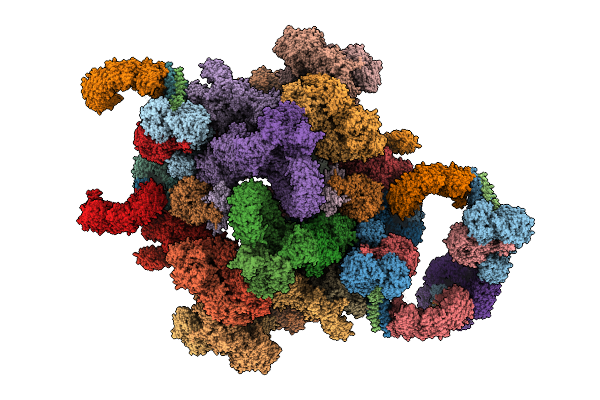 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: CYTOSOLIC PROTEIN Ligands: ZN, GTP, MG, ADP |
 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVAL PROTEIN/RNA/DNA |
 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVIRAL PROTEIN |
 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVIRAL PROTEIN |
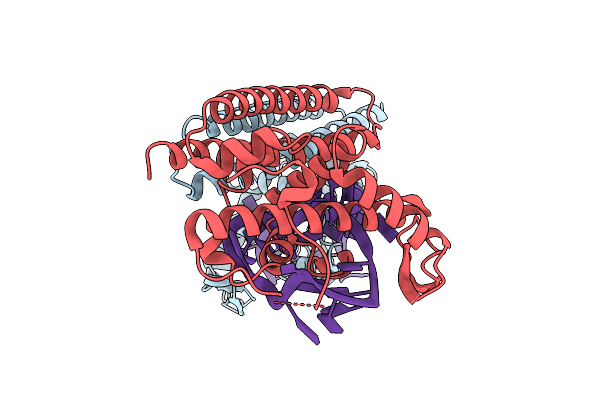 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVIRAL PROTEIN |
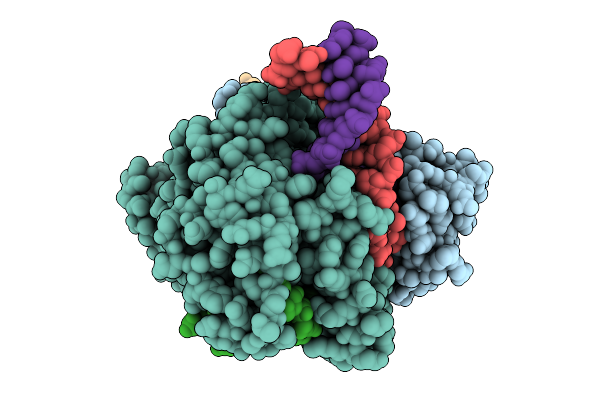 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVIRAL PROTEIN Ligands: MG |
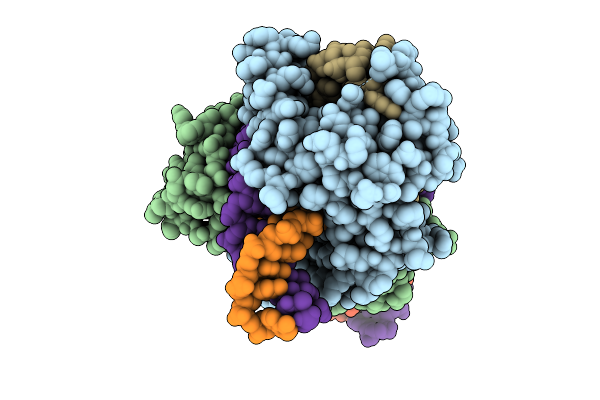 |
Organism: Salicola phage cgphi29
Method: ELECTRON MICROSCOPY Release Date: 2026-03-11 Classification: ANTIVIRAL PROTEIN Ligands: MG |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2026-02-04 Classification: METAL BINDING PROTEIN Ligands: A1BVO, FE2 |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.61 Å Release Date: 2026-02-04 Classification: METAL BINDING PROTEIN Ligands: A1BVO, FE2 |
 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.86 Å Release Date: 2026-02-04 Classification: METAL BINDING PROTEIN Ligands: FE2 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.03 Å Release Date: 2026-01-28 Classification: LIGASE/HYDROLASE |
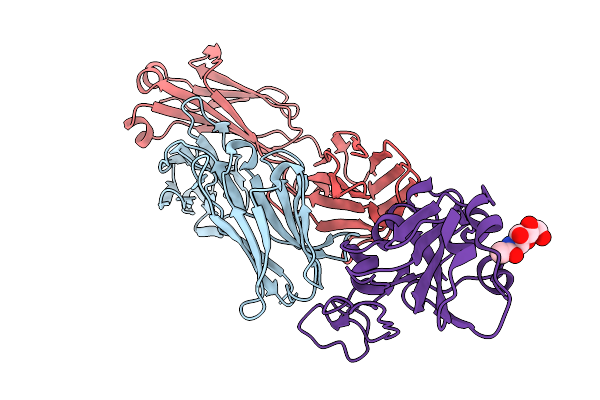 |
Organism: Severe acute respiratory syndrome coronavirus 2, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2026-01-07 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
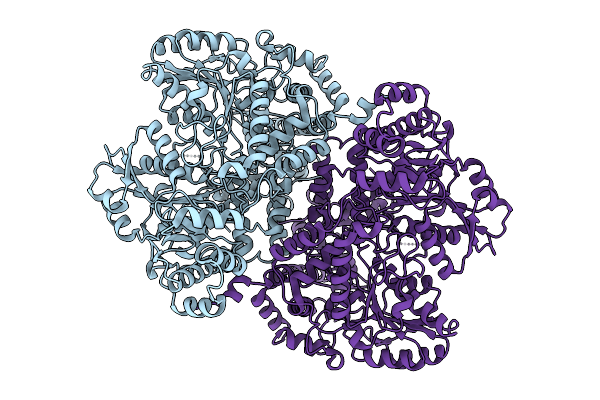 |
Organism: Escherichia coli str. k-12 substr. dh10b
Method: ELECTRON MICROSCOPY Resolution:2.32 Å Release Date: 2025-12-31 Classification: OXIDOREDUCTASE |
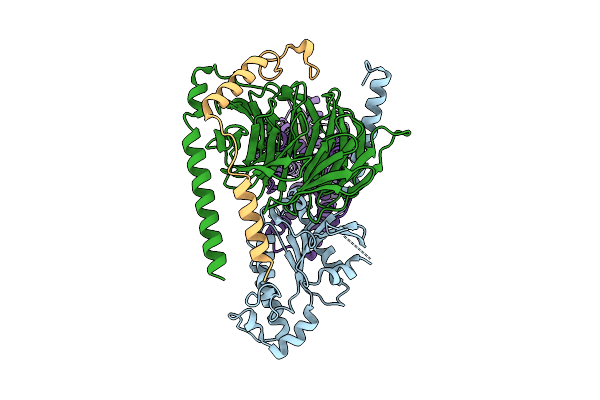 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.01 Å Release Date: 2025-12-10 Classification: SIGNALING PROTEIN Ligands: P9X |
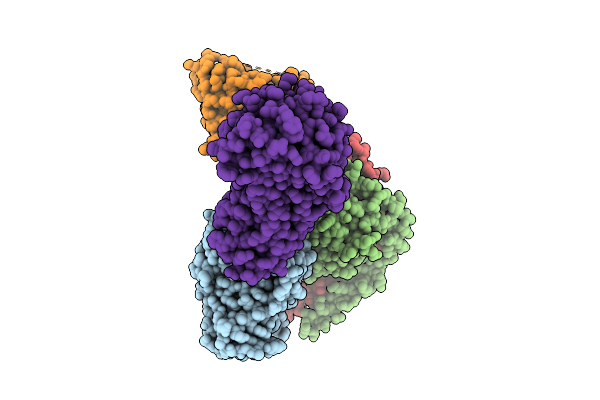 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: SIGNALING PROTEIN/IMMUNE SYSTEM Ligands: FI7 |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.73 Å Release Date: 2025-12-10 Classification: SIGNALING PROTEIN/IMMUNE SYSTEM Ligands: P9X |

