Search Count: 43,357
All
Selected
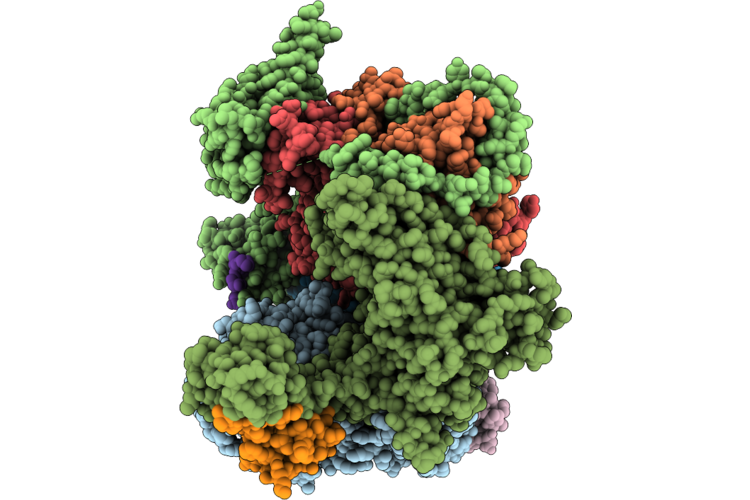 |
Cryo-Em Structure Of The Human Holo-Tfiih And Xpc Initial Encounter Complex
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN |
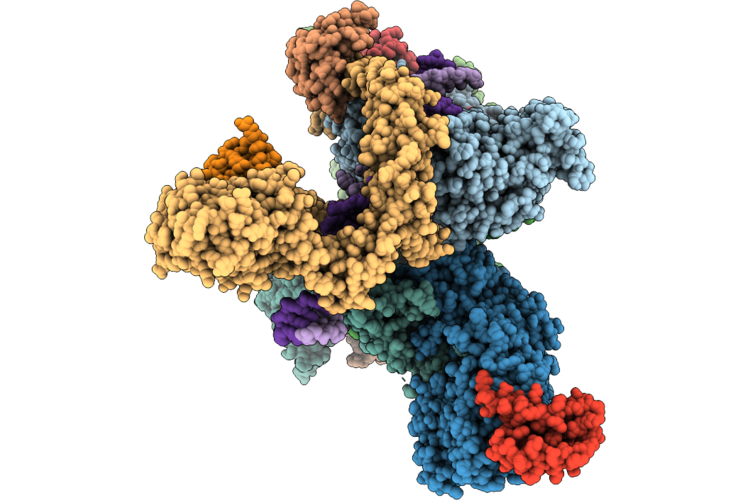 |
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Composite Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.32 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
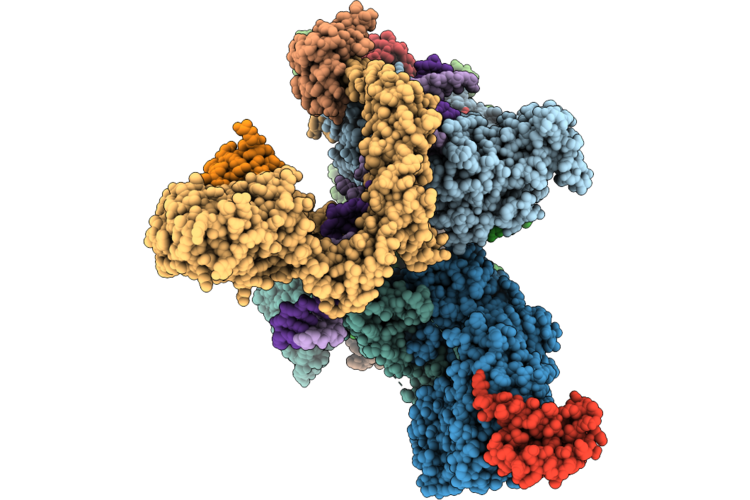 |
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Consensus Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.91 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
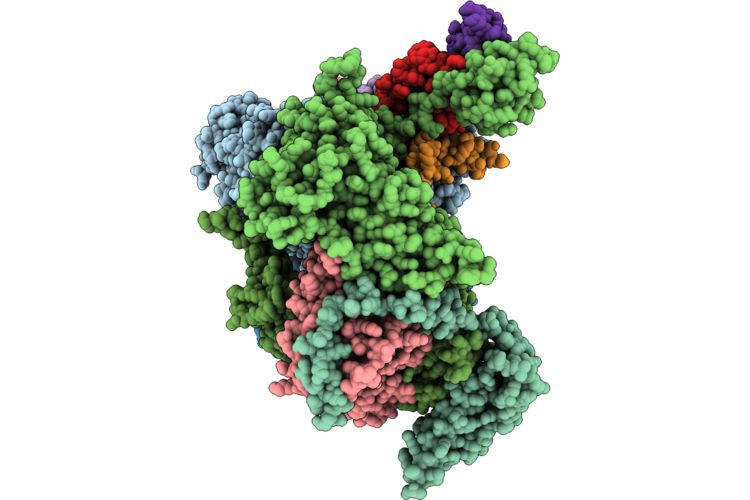 |
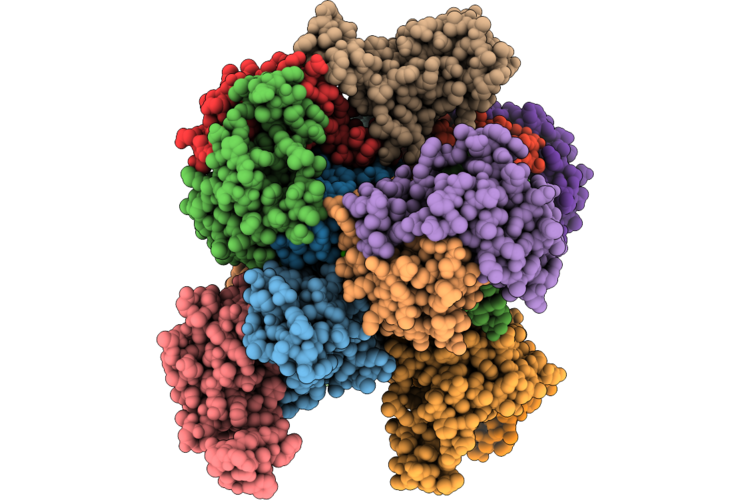
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc-Xpa Complex Bound To Bulky Lesion-Mimic Dna
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN |
 |
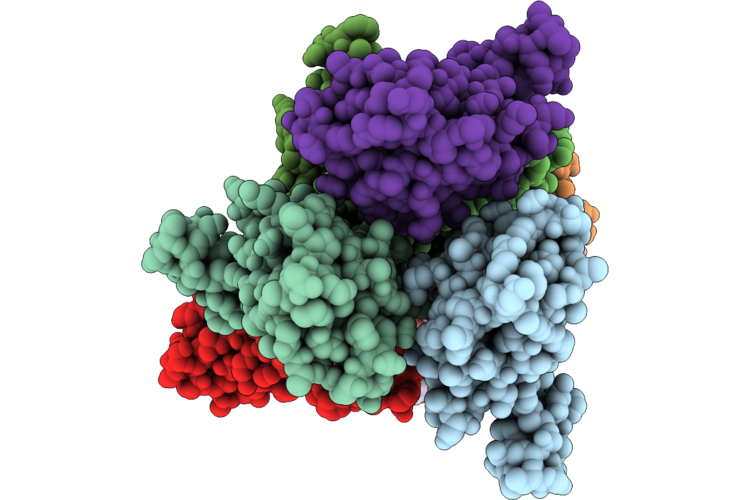
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.54 Å Release Date: 2026-05-06 Classification: CYTOKINE Ligands: CA |
 |
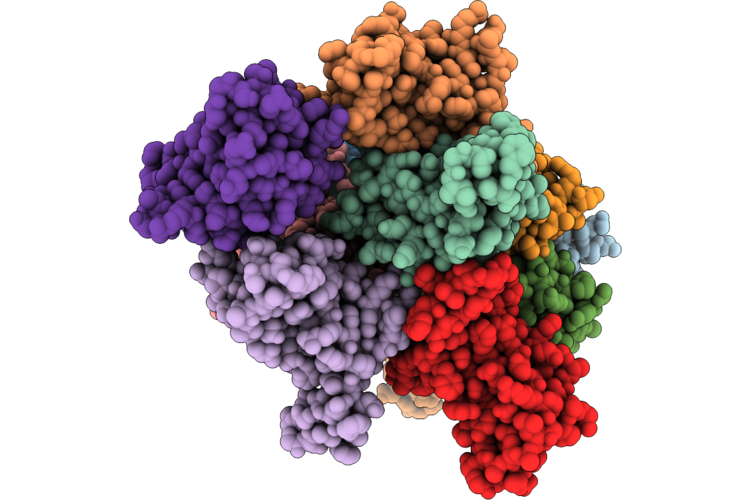
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2026-05-06 Classification: CYTOKINE |
 |
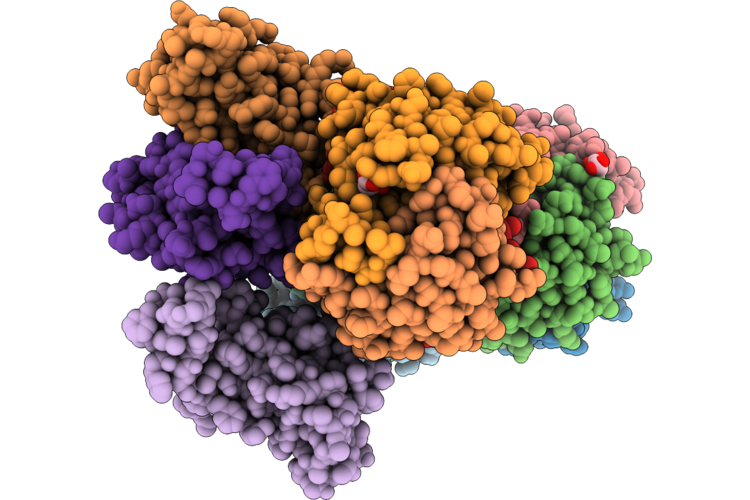
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.53 Å Release Date: 2026-05-06 Classification: CYTOKINE Ligands: TRS |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.49 Å Release Date: 2026-05-06 Classification: CYTOKINE Ligands: MLI |
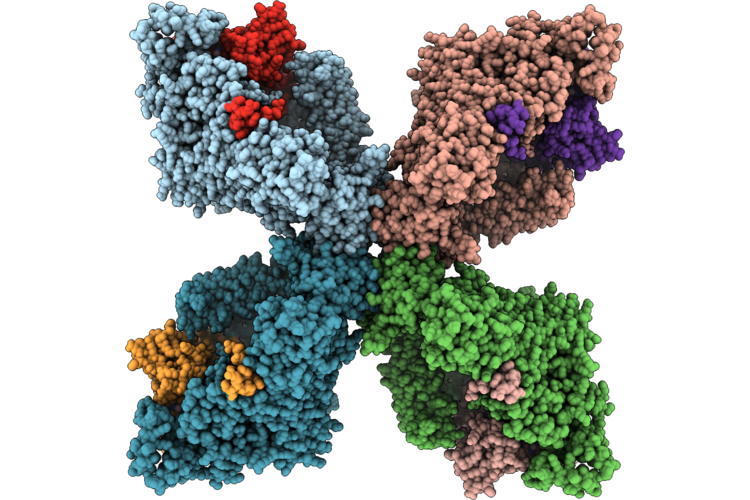 |
Organism: Escherichia coli, Escherichia phage phiv-1
Method: ELECTRON MICROSCOPY Resolution:2.33 Å Release Date: 2026-05-06 Classification: ANTIVIRAL PROTEIN Ligands: ATP, MG |
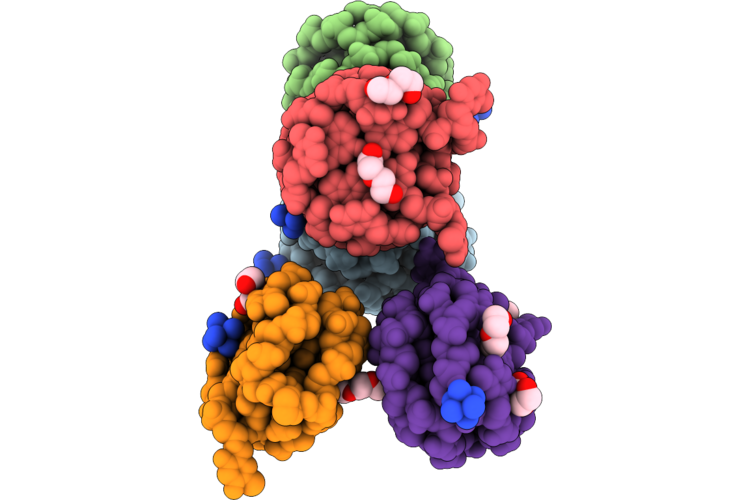 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2026-05-06 Classification: DNA Ligands: K, NCO, PEG, PG4, PGE, MES |
 |
Hybrid-1 Form Of Human Telomere Dna Quadruplex With Wild-Type 5'-Flanking In K+ Solution
|
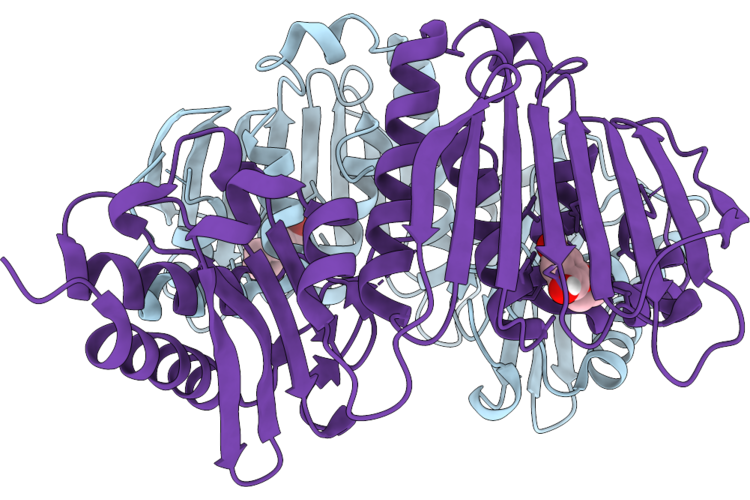 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: A1CKR |
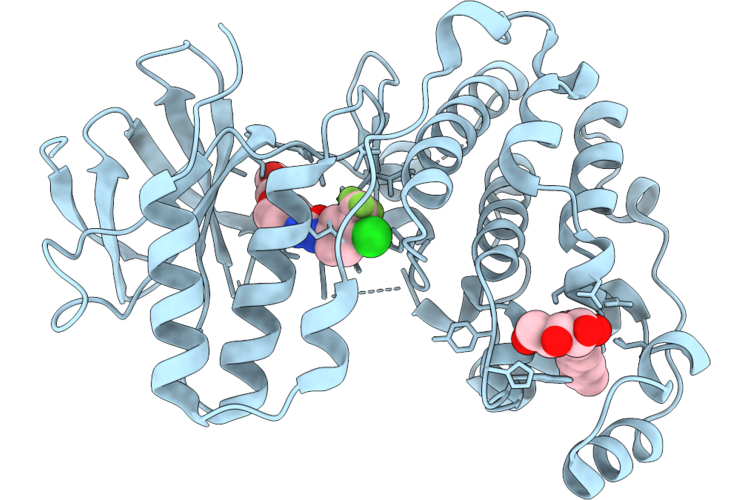 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: BAX, BOG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: BOG, A1JCR |
 |
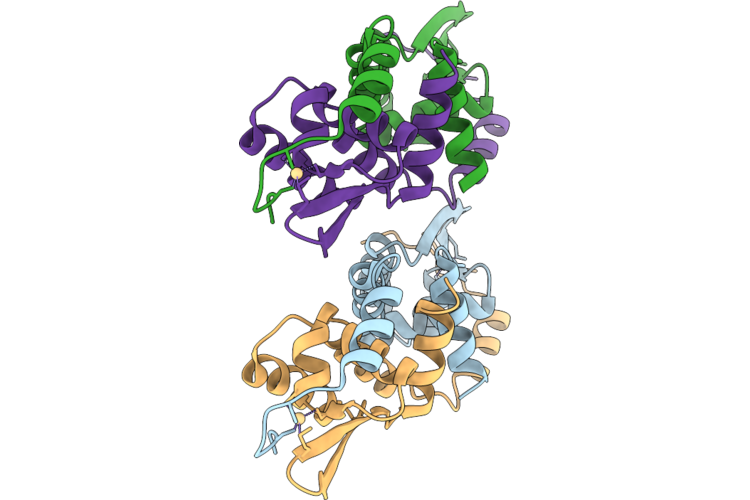
Organism: Listeria monocytogenes egd-e
Method: X-RAY DIFFRACTION Resolution:3.01 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN |
 |
Organism: Listeria monocytogenes egd-e
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: CD |
 |
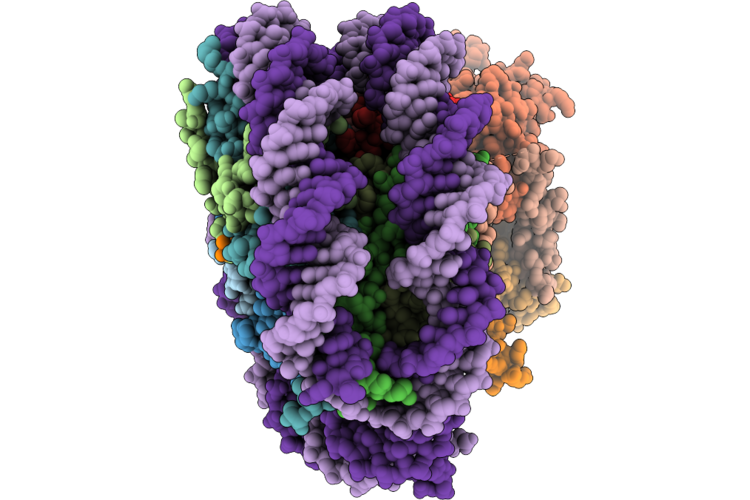
Organism: Listeria monocytogenes egd-e
Method: X-RAY DIFFRACTION Resolution:1.69 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: CR |
 |
Organism: Xenopus laevis, Homo sapiens, Artificial sequences
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: GENE REGULATION/DNA |
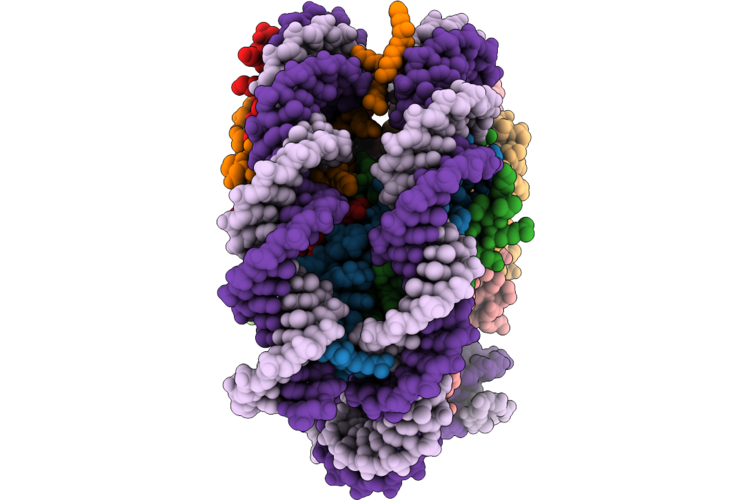 |
Cryo-Em Structure Of The Nucleosome Core Particle With Site-Specific Dna-Histone Crosslinking
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN/DNA |
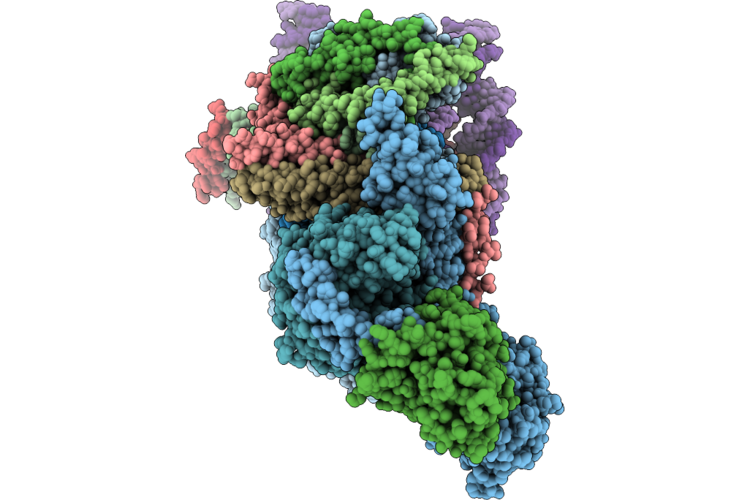 |
Cryo-Em Structure Of The Histone Deacetylase Complex Rpd3L In Complex With Mono-Nucleosome
Organism: Saccharomyces cerevisiae s288c, Xenopus laevis, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: GENE REGULATION/DNA Ligands: ZN |

