Search Count: 46,868
All
Selected
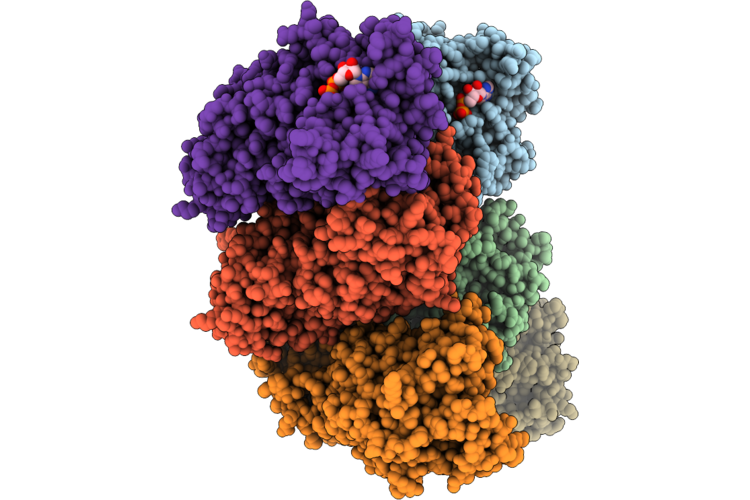 |
Crystal Structure Of Hera Like Helicae Yjgr With Ntpase Fold From Escherchia Coli
Organism: Escherichia coli str. k-12 substr. mg1655
Method: X-RAY DIFFRACTION Resolution:2.90 Å Release Date: 2026-05-06 Classification: GENE REGULATION Ligands: ATP, MG |
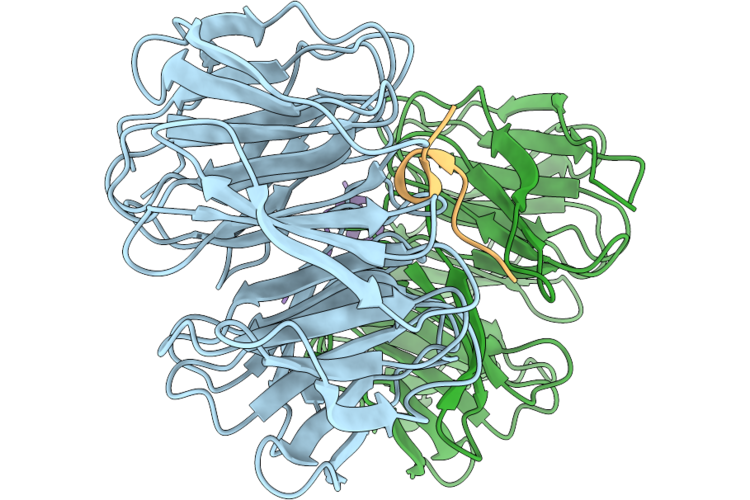 |
Organism: Homo sapiens, Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.57 Å Release Date: 2026-05-06 Classification: NUCLEAR PROTEIN |
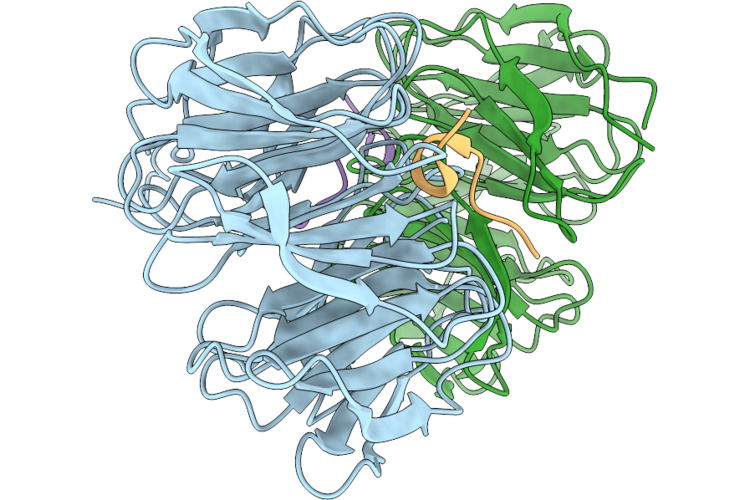 |
Organism: Homo sapiens, Mus musculus
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-05-06 Classification: NUCLEAR PROTEIN |
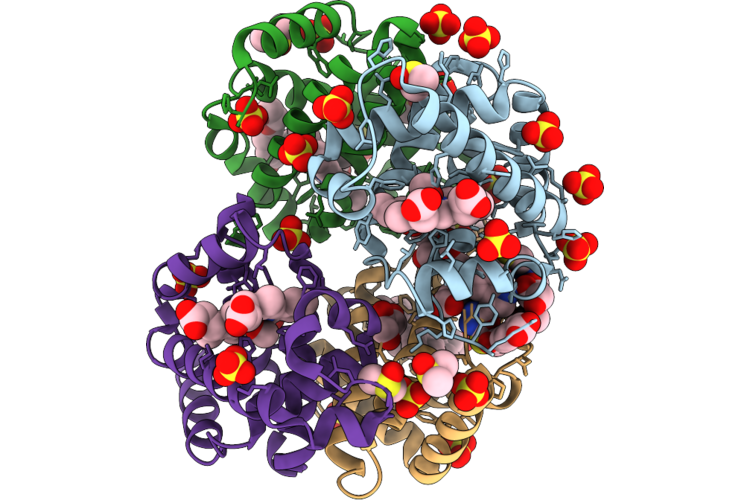 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: HEM, GOL, A1ITR, DMS, CMO, ACT, SO4, PGE |
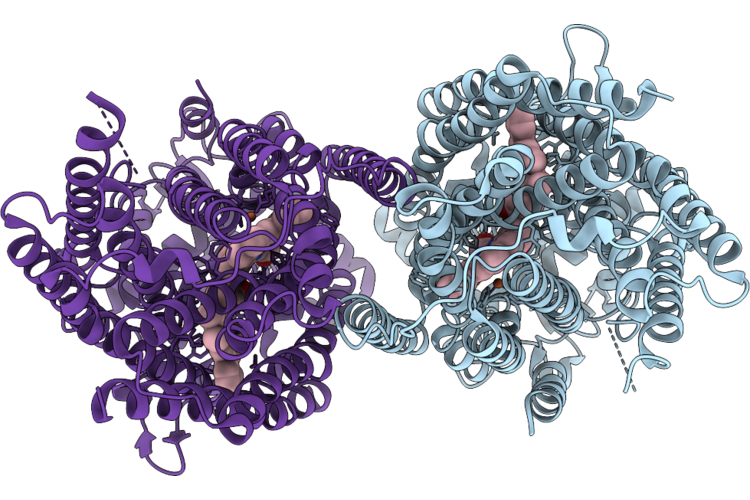 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase At Ph 8.0 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
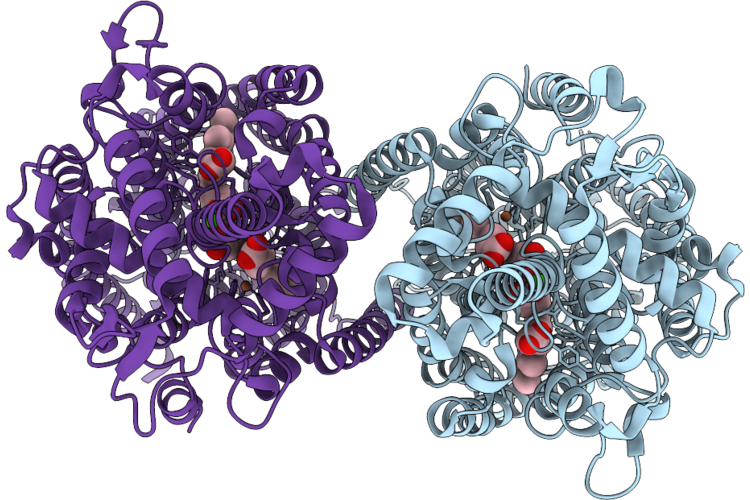 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Arg720Ala Variant At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE |
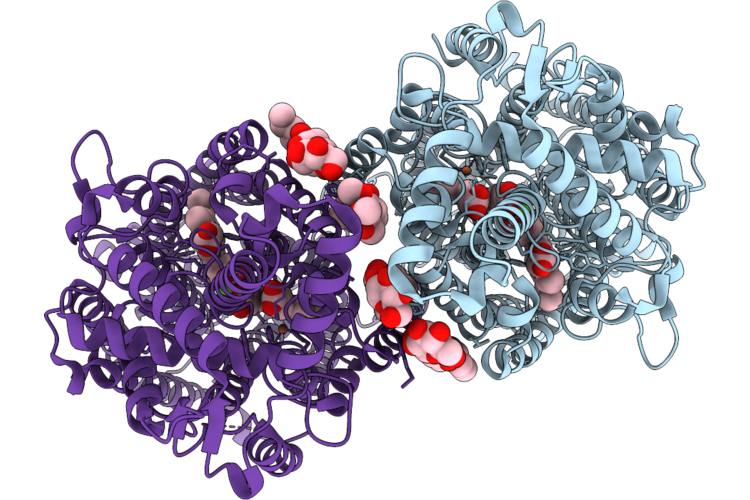 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Trp718Ala Variant At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, LMT, UQ5 |
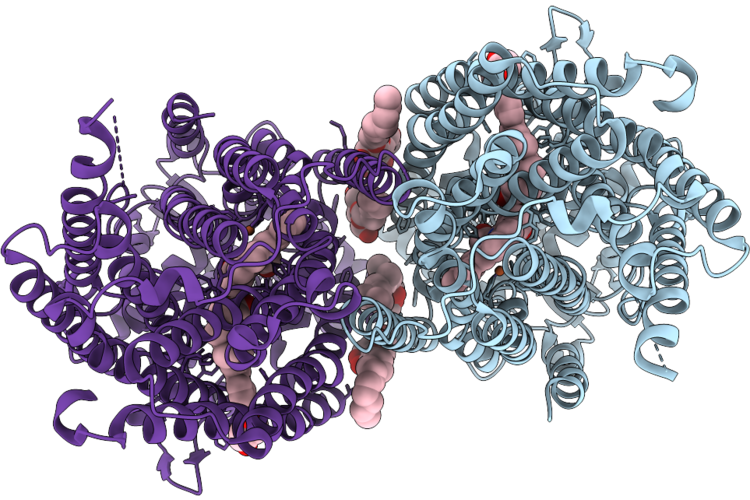 |
Cryoem Structure Of Native Quinol Dependent Nitric Oxide Reductase Trp718Ala Variant With Quino At Ph 6.5 On Gold Grid.
Organism: Achromobacter xylosoxidans
Method: ELECTRON MICROSCOPY Resolution:2.40 Å Release Date: 2026-05-06 Classification: MEMBRANE PROTEIN Ligands: HEM, CA, FE, UQ5, HQE, LMT |
 |
Crystal Structure Of Udp-N-Acetylmuramate-L-Alanine Ligase (Murc) From Pseudomonas Aeruginosa In Complex With Compound Osa_001175 (Wyh77)
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-05-06 Classification: LIGASE Ligands: PO4, MG, A1J52, EDO, DMS, PEG |
 |
Aetf, A Single-Component Flavin-Dependent Tryptophan Halogenase, In Complex With Tryptoline
Organism: Aetokthonos hydrillicola thurmond2011
Method: X-RAY DIFFRACTION Resolution:1.87 Å Release Date: 2026-05-06 Classification: FLAVOPROTEIN Ligands: FAD, WYH, PEG, CL |
 |
Organism: Oryza sativa subsp. japonica
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSFERASE |
 |
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSFERASE |
 |
Organism: Sorghum bicolor
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: A1EE9, MG |
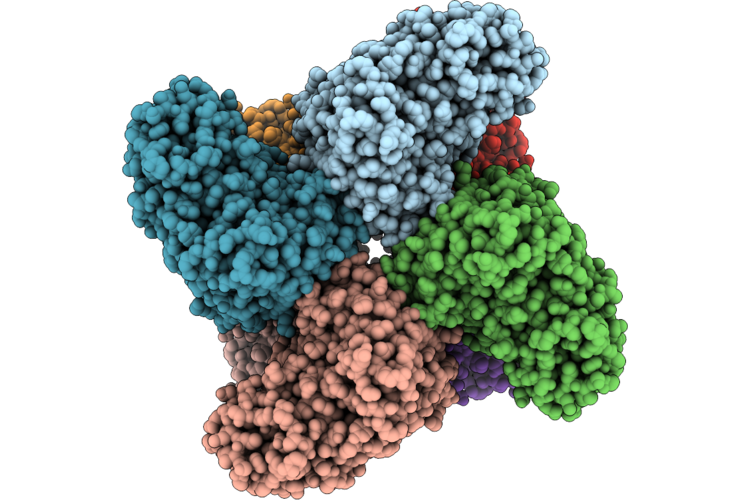 |
Cryo-Em Structure And Rational Engineering Of A Novel Efficient Ochratoxin A-Detoxifying Amidohydrolase
Organism: Pseudoxanthomonas wuyuanensis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN, 97U |
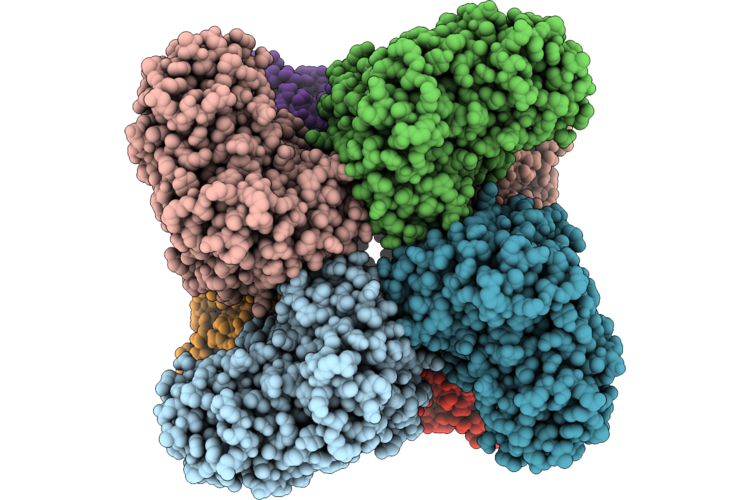 |
Cryo-Electron Microscopic Structure Of A Novel Amidohydrolase With Three Mutations
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.69 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN, 97U |
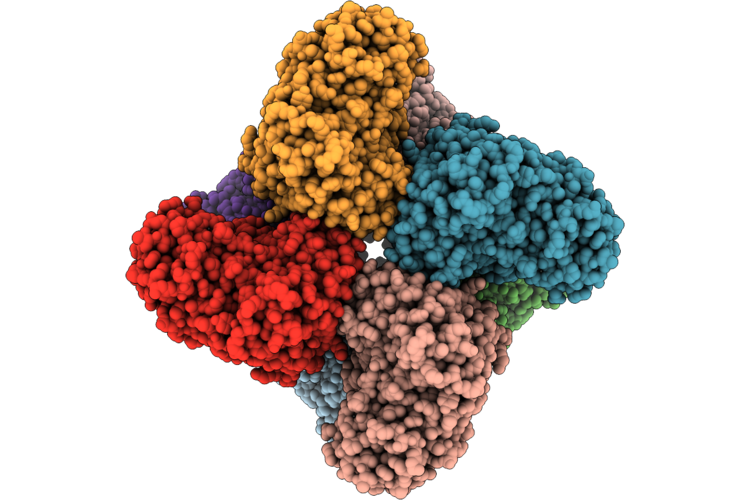 |
Cryo-Electron Microscopic Structure Of A Highly Efficient Ochratoxin Detoxification Enzyme Lladh
Organism: Novilysobacter luteus
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-06 Classification: HYDROLASE Ligands: ZN |
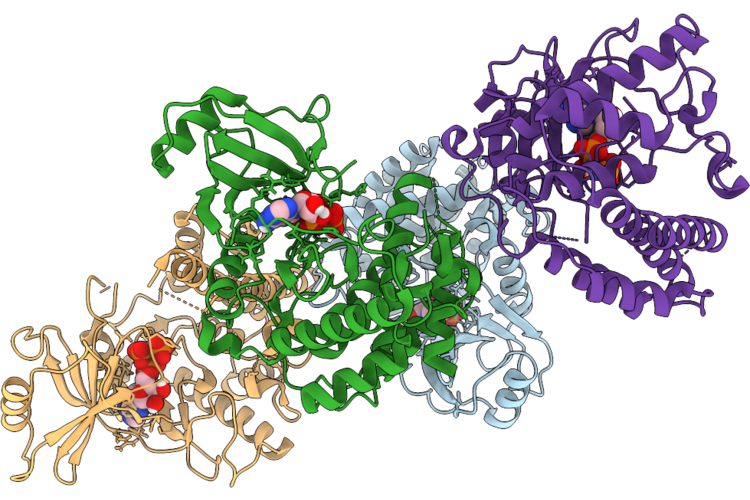 |
Caldalkalibacillus Thermarum Acyl-Coa Dehydrogenase Member 10 Bound To Anp And Magnesium
Organism: Caldalkalibacillus thermarum
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: ANP |
 |
Crystal Structure Of Human Adenosine Kinase (Adk) In Complex With Inhibitor Bki-1817
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-05-06 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: CL, SO4, A1B96 |
 |
Crystal Structure Of Human Adenosine Kinase (Adk) In Complex With Inhibitor Bki-1676
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.81 Å Release Date: 2026-05-06 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: A1B95, SO4, CL |
 |
Crystal Structure Of Human Adenosine Kinase (Adk) In Complex With Inhibitor Bki-1553
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.41 Å Release Date: 2026-05-06 Classification: TRANSFERASE/TRANSFERASE INHIBITOR Ligands: CL, UW5, SO4 |

