Deposition Date
2026-02-12
Release Date
2026-05-06
Last Version Date
2026-05-06
Entry Detail
PDB ID:
28PP
Keywords:
Title:
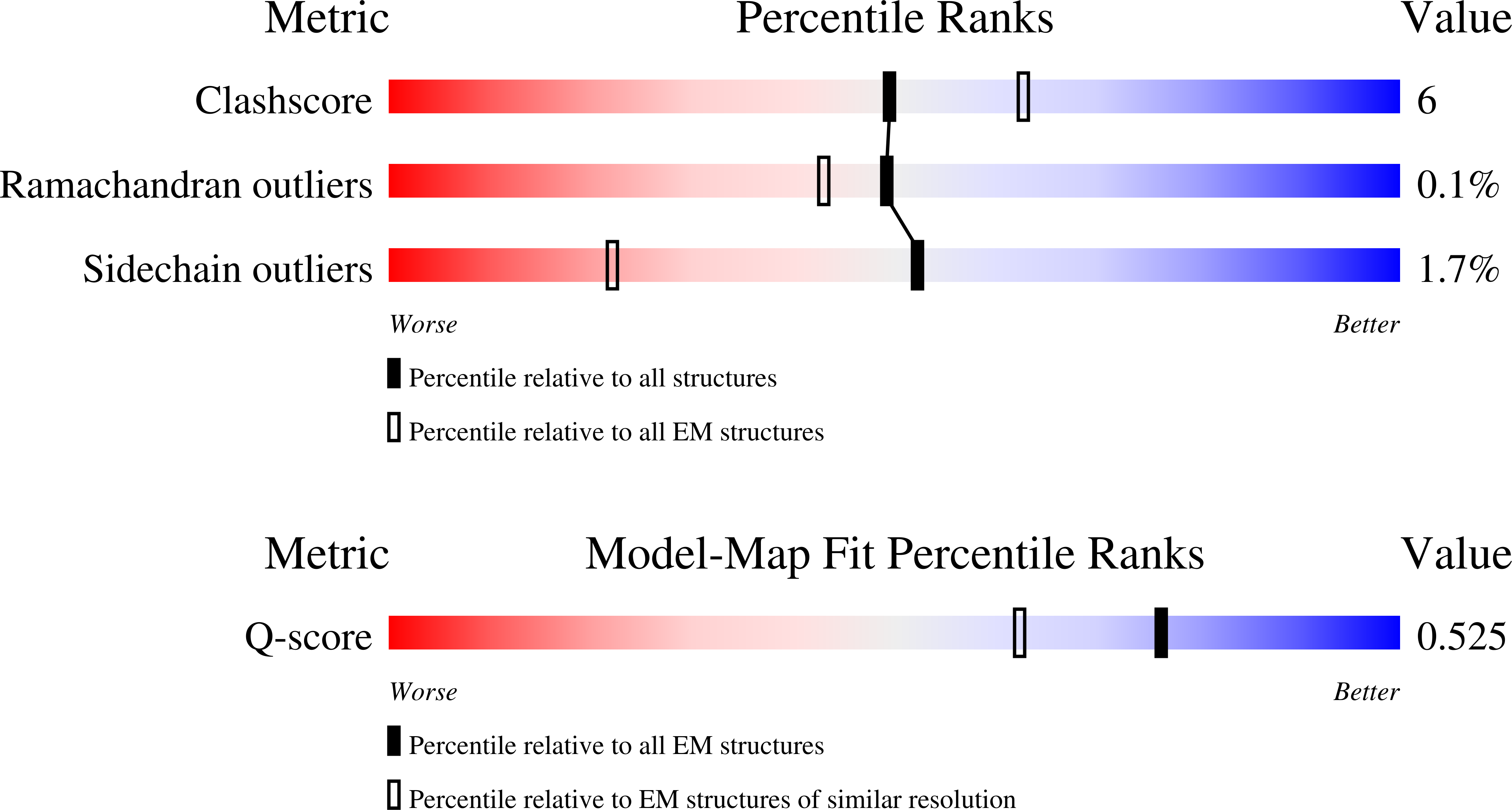
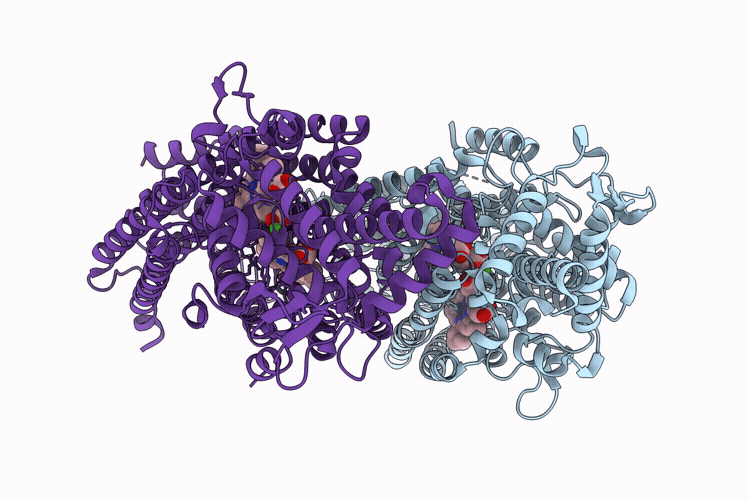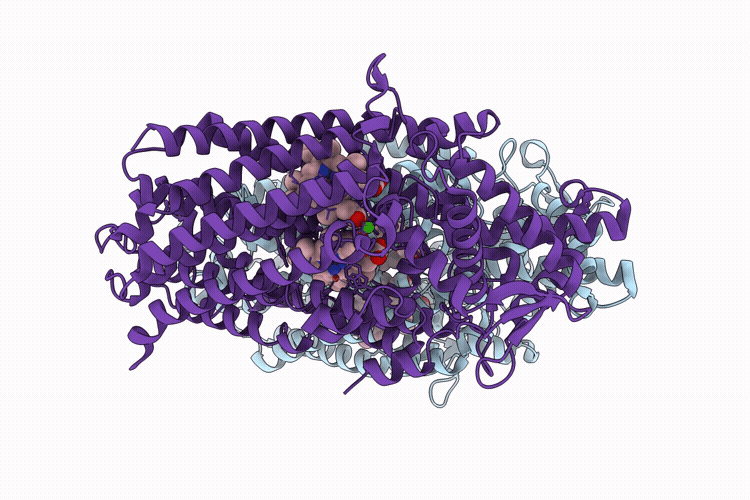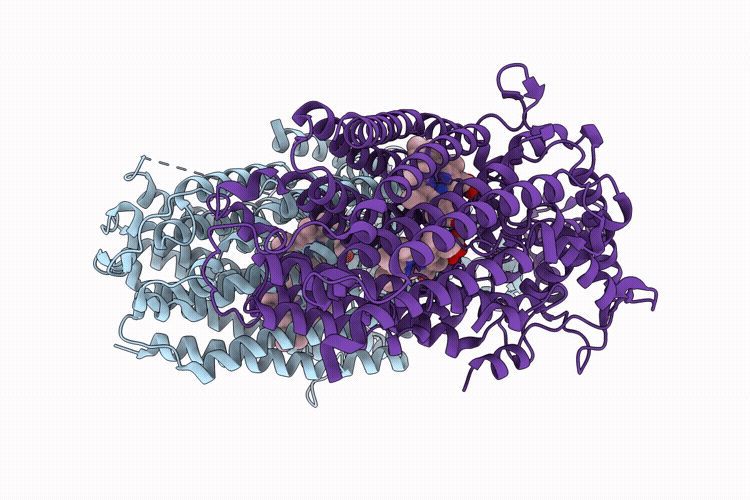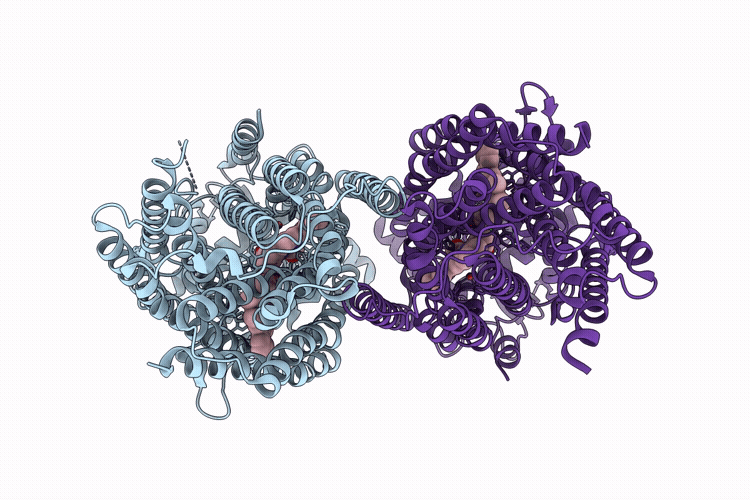
CryoEM structure of native quinol dependent Nitric Oxide Reductase Arg720Ala variant at pH 6.5 on gold grid.
Biological Source:
Source Organism(s):
Achromobacter xylosoxidans (Taxon ID: 85698)
Expression System(s):
Method Details:
Experimental Method:
Resolution:
2.90 Å
Aggregation State:
PARTICLE
Reconstruction Method:
SINGLE PARTICLE