Search Count: 837
All
Selected
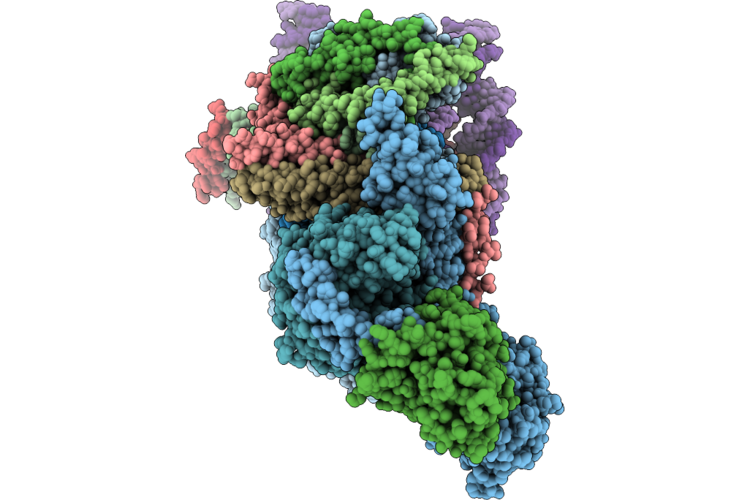 |
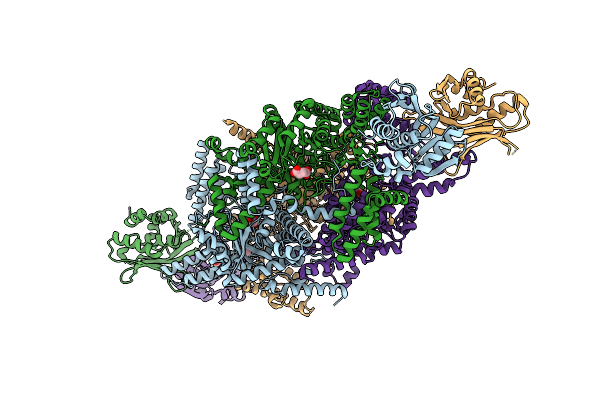
Cryo-Em Structure Of The Histone Deacetylase Complex Rpd3L In Complex With Mono-Nucleosome
Organism: Saccharomyces cerevisiae s288c, Xenopus laevis, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: GENE REGULATION/DNA Ligands: ZN |
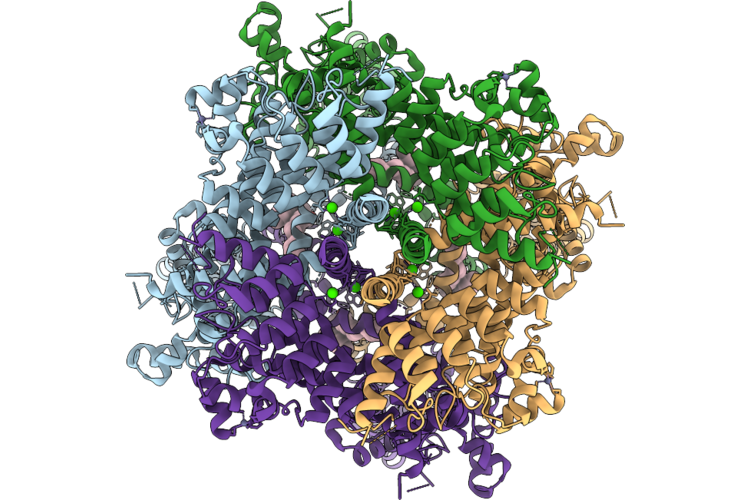 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: METAL TRANSPORT Ligands: CA, ZN, A1L5I |
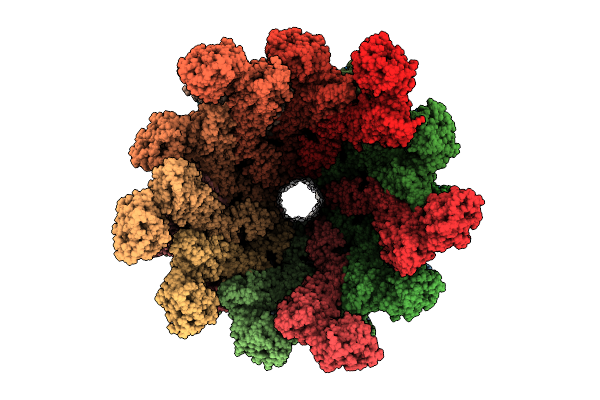 |
Organism: Vibrio alginolyticus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: STRUCTURAL PROTEIN |
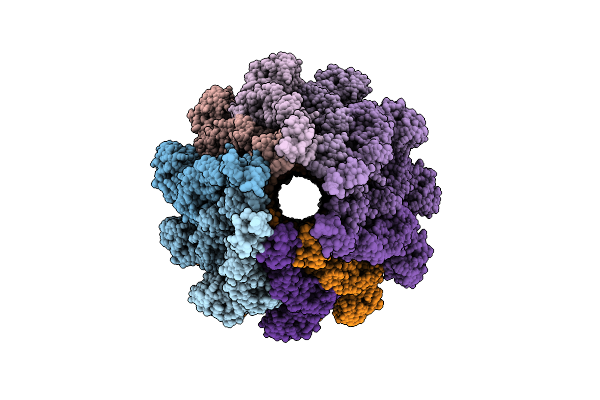 |
Organism: Vibrio alginolyticus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: STRUCTURAL PROTEIN |
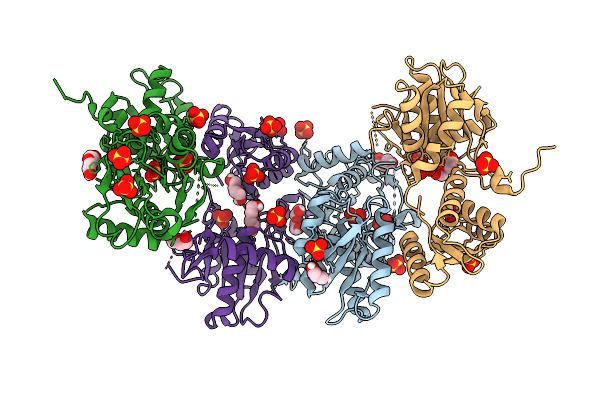 |
Organism: Neisseria gonorrhoeae fa 1090
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-01 Classification: BIOSYNTHETIC PROTEIN Ligands: SO4, PEG, GOL |
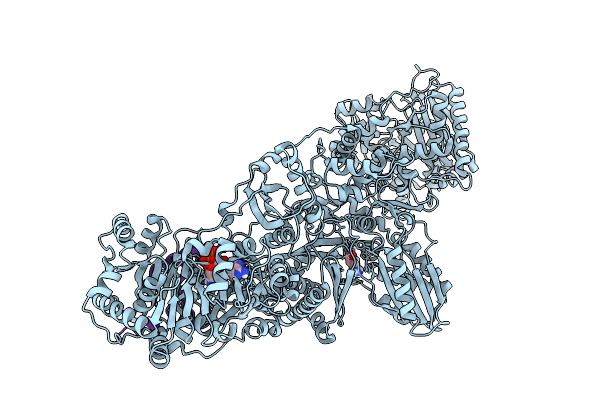 |
Organism: Pseudomonas aeruginosa pao1
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: BIOSYNTHETIC PROTEIN Ligands: DAB, ATP |
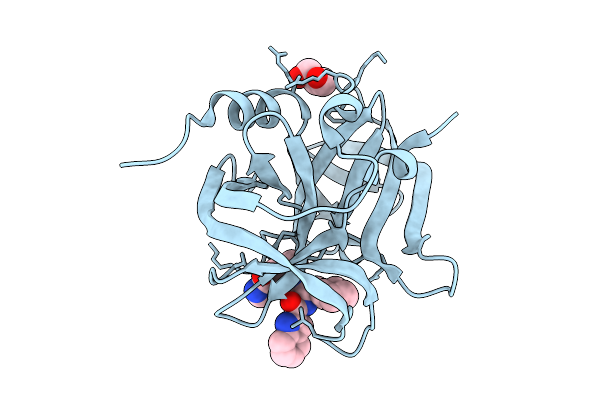 |
Organism: Human enterovirus 71 (strain brcr)
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-01 Classification: HYDROLASE Ligands: GOL, FHR |
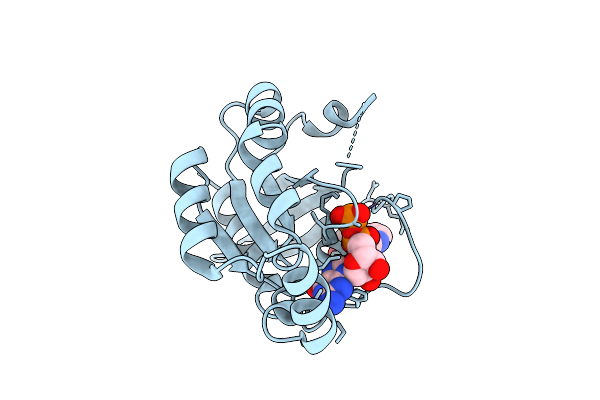 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.43 Å Release Date: 2026-04-01 Classification: CELL INVASION Ligands: GDP, A1D9L, MG |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-01 Classification: CELL INVASION Ligands: GDP, MG |
 |
Structure Of The Rhoa Y42C Mutant Featuring Thr37 Coordination Of Magnesium Ion
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-01 Classification: CELL INVASION Ligands: GDP, MG |
 |
Crystal Structure Of Mycobacterium Tuberculosis Isocitrate Lyase 2 Fixed In The Apo Form With Disulfide Bonds
Organism: Mycobacterium tuberculosis cdc1551
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-03-18 Classification: LYASE Ligands: SIN, EDO |
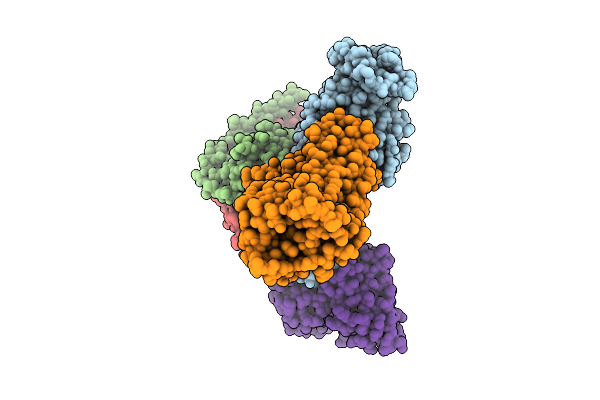 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: AKG |
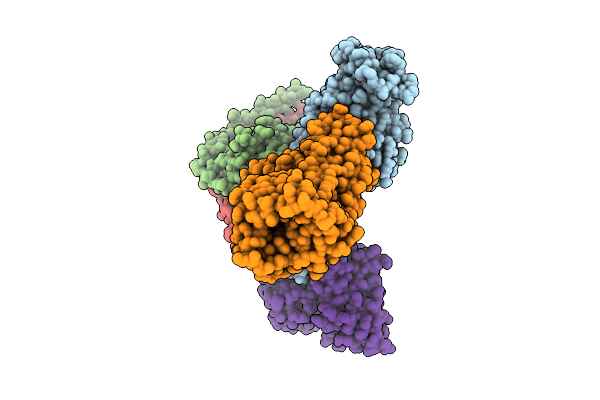 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: ITN |
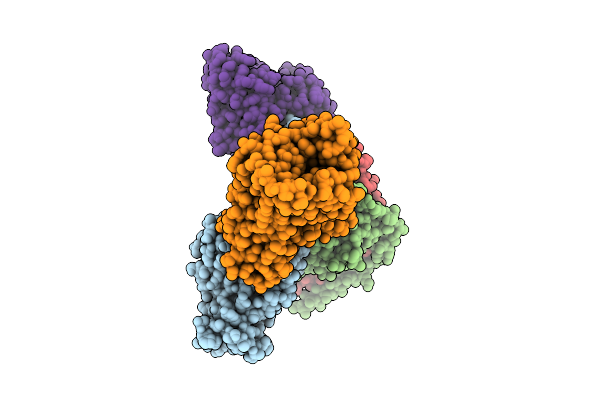 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: A1E2R |
 |
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM |
 |
Structure Of Mammalian Rna Polymerase Ii Stalled By Actinomycin D At The N-1 Position.
Organism: Synthetic construct, Bos taurus
Method: ELECTRON MICROSCOPY Resolution:2.53 Å Release Date: 2026-02-11 Classification: TRANSCRIPTION Ligands: ZN, MG |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-02-04 Classification: PROTEIN FIBRIL |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN Ligands: PIO, A1LVR |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN Ligands: PIO, A1LWZ |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: A1LVR |

