Search Count: 1,766
All
Selected
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
 |
Organism: Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
 |
Organism: Bacillus sp. tsa-4
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-04-15 Classification: TRANSPORT PROTEIN Ligands: ADP |
 |
Organism: Bacillus sp. tsa-4
Method: ELECTRON MICROSCOPY Resolution:3.90 Å Release Date: 2026-04-15 Classification: TRANSPORT PROTEIN Ligands: ADP |
 |
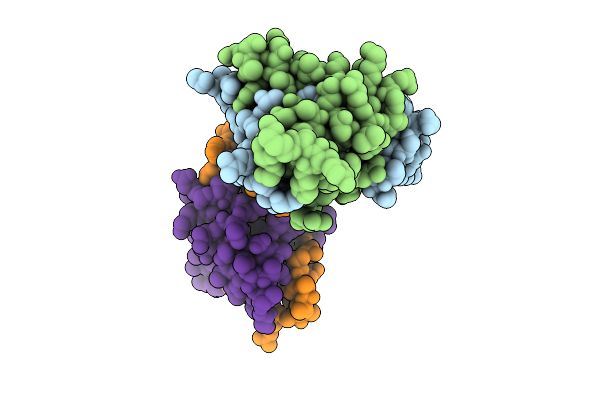
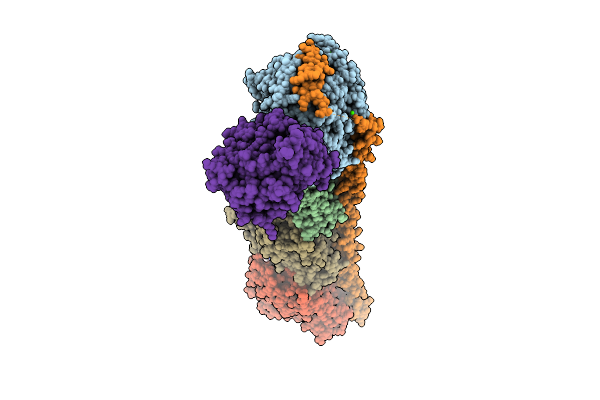
Cryo-Em Structure Of Integrin Alpha V Beta 6 Complex With A Bicyclic Inhibitory Peptide
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: CELL ADHESION Ligands: CA, NAG, MAN, MN |
 |
The Structural Basis Of The Recognition Of The Histone Variant H2A.Z By The Srcap Catalytic Subunit
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.53 Å Release Date: 2026-04-08 Classification: DNA BINDING PROTEIN |
 |
Organism: Rattus norvegicus, Gallus gallus, Sus scrofa
Method: X-RAY DIFFRACTION Resolution:2.93 Å Release Date: 2026-04-01 Classification: CELL CYCLE Ligands: GTP, MG, CA, MES, A1L80, GDP, ACP |
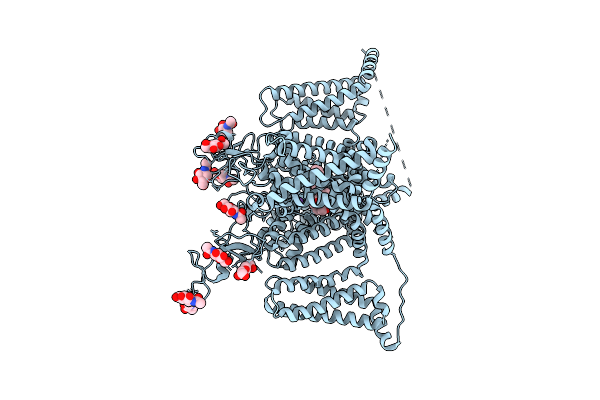 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: MEMBRANE PROTEIN Ligands: NAG, A1EQU, NA |
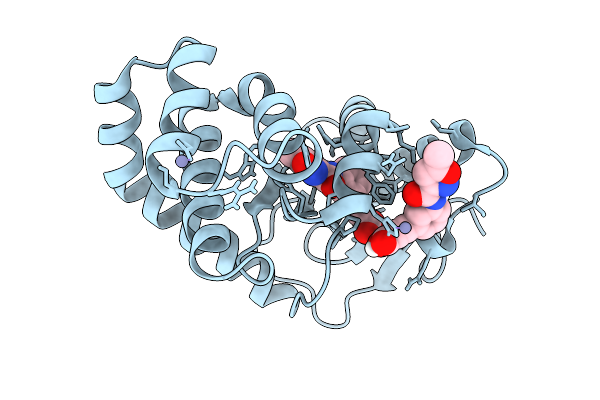 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: ZN, A1CJH |
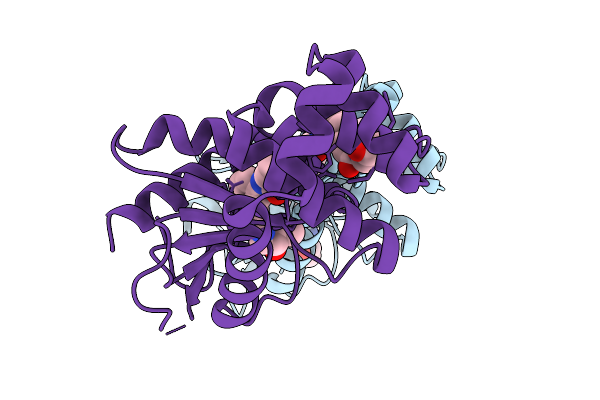 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.61 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CQW, A1CJJ |
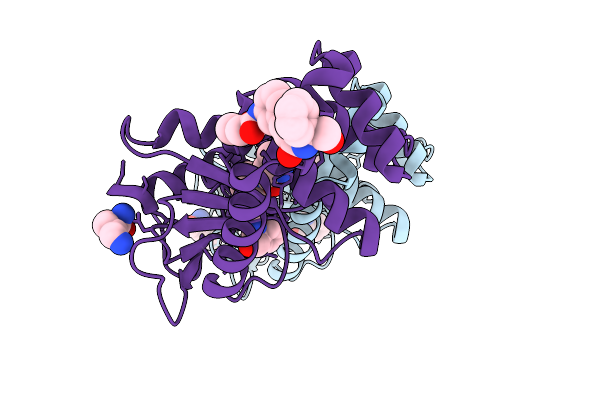 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: LYS, A1CJK |
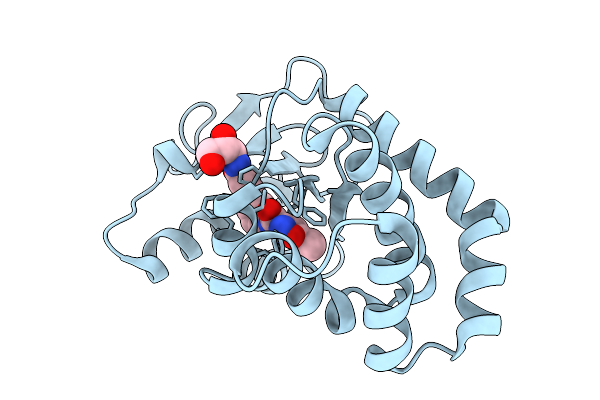 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJL |
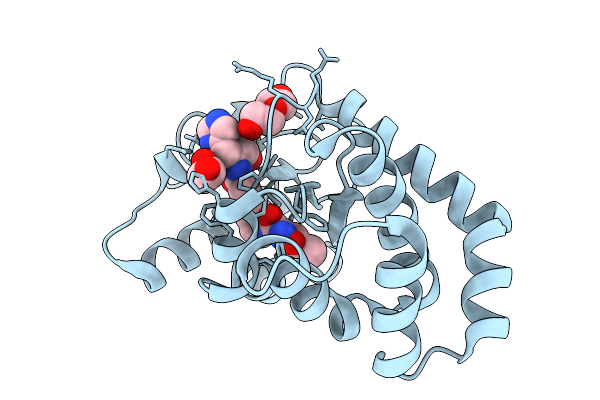 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJM, 1PE, EDO |
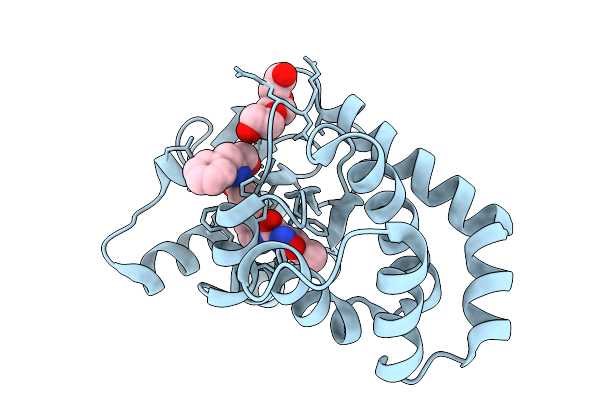 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.81 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJN, PGE |
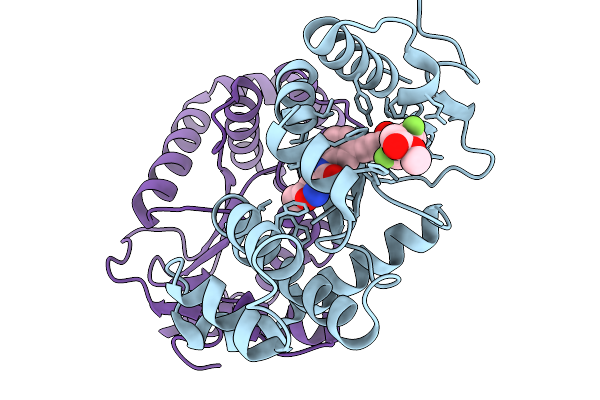 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJO |
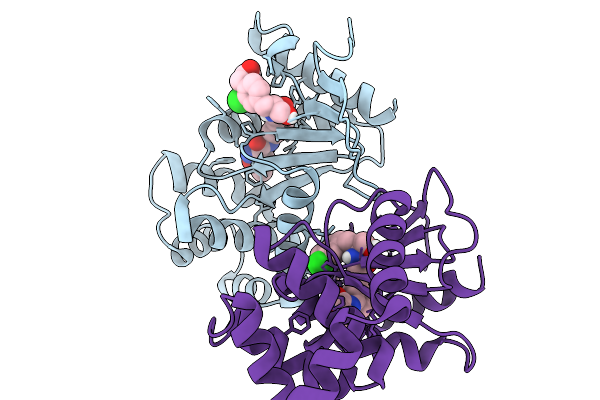 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJP |
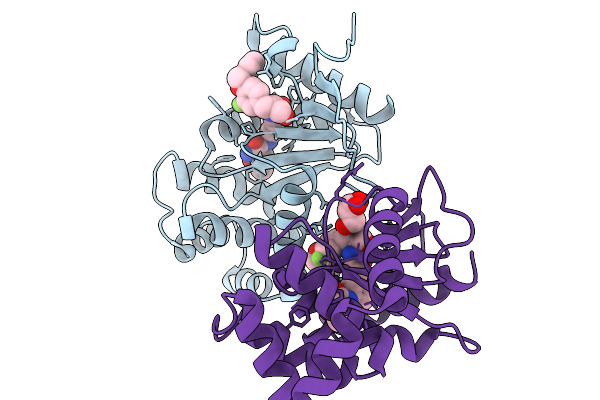 |
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: A1CJQ, GOL |
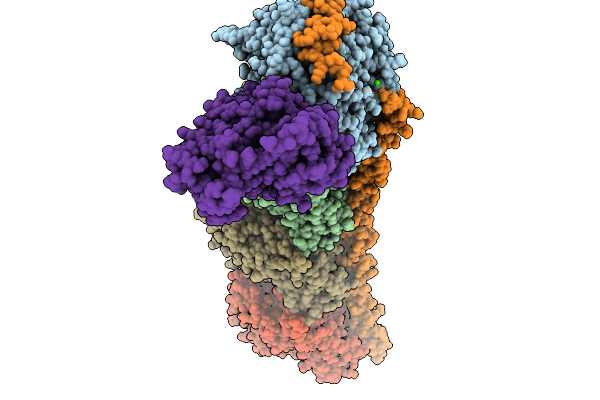 |
Organism: Rattus norvegicus, Gallus gallus, Sus scrofa
Method: X-RAY DIFFRACTION Resolution:2.54 Å Release Date: 2026-03-25 Classification: CELL CYCLE Ligands: GTP, MG, CA, MES, GDP, A1L8Z |

