Search Count: 3,853
All
Selected
 |
Organism: Scenedesmaceae sp.
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-22 Classification: LIPID BINDING PROTEIN Ligands: AXT |
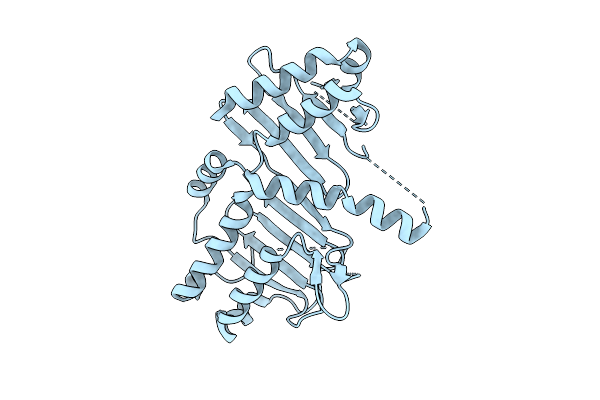 |
Organism: Streptomyces novoguineensis
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2026-04-08 Classification: LYASE Ligands: HOH |
 |
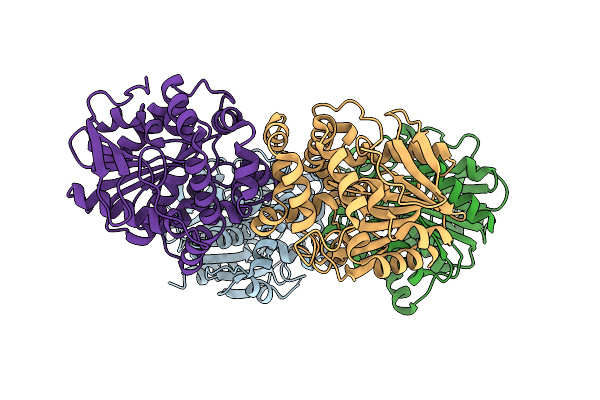
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Trp185Tyr/His280Ala With 2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-04-01 Classification: TRANSFERASE |
 |
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Met/Trp185Tyr/His280Ala With 2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-03-25 Classification: TRANSFERASE |
 |
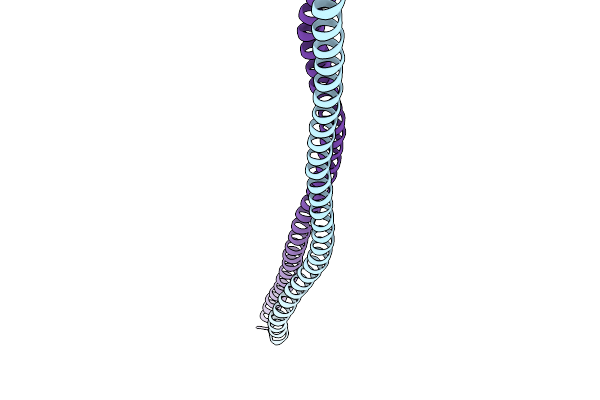
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Met/His280Ala With 2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-01-21 Classification: TRANSFERASE |
 |
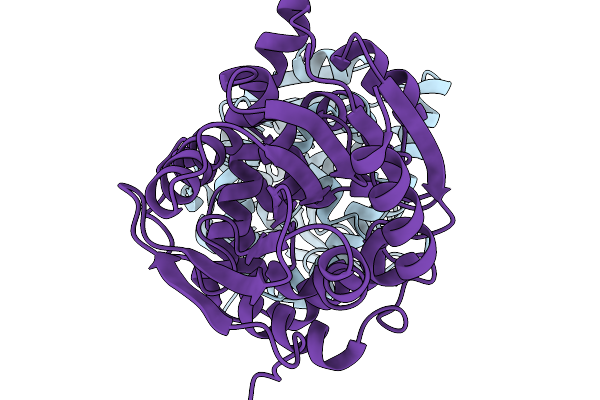
Crystal Structure Of The Coiled-Coil Based Fibril Core Of The Tardigrade Cahs-8 Protein
Organism: Hypsibius exemplaris
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-01-14 Classification: PROTEIN FIBRIL |
 |
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163-His280Ala With (S)-2-Fluoro-3-Phenylpropanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:2.01 Å Release Date: 2026-01-07 Classification: HYDROLASE |
 |
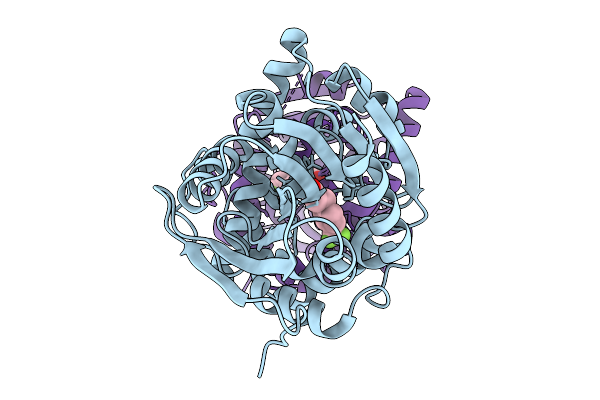
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Lys/Trp185Tyr/His280Ala With (S)-2-Fluoro-3-(4-(Trifluoromethyl)Phenyl)Propanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2025-12-31 Classification: TRANSFERASE Ligands: A1ES3 |
 |
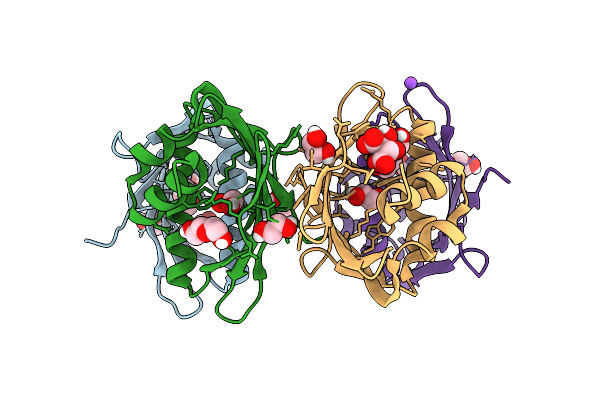
Crystal Structure Of The Fluoroacetate Dehalogenase Rpa1163 - Lys181Ser/Trp185Gly/His280Ala With (S)-2-Fluoro-3-(4-(Trifluoromethyl)Phenyl)Propanoic Acid
Organism: Rhodopseudomonas palustris cga009
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2025-12-31 Classification: TRANSFERASE Ligands: A1ES3 |
 |
Organism: Micromonospora sp. hm134
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2025-07-16 Classification: BIOSYNTHETIC PROTEIN Ligands: EDO, PEG, CIT, NA |
 |
Organism: Pseudorhizobium banfieldiae
Method: X-RAY DIFFRACTION Resolution:1.89 Å Release Date: 2025-05-21 Classification: OXIDOREDUCTASE Ligands: SER, 4MO, F3S, EDO, PEG, MGD, SO4, P33, GOL, O, FES, 1PE |
 |
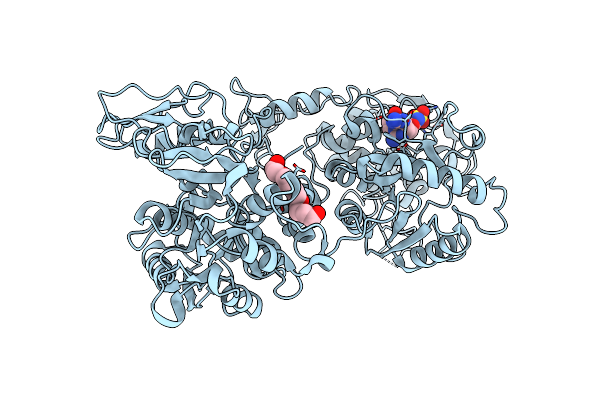
Organism: Nanorana parkeri
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2025-05-07 Classification: ANTITOXIN Ligands: 1PE, YGU |
 |
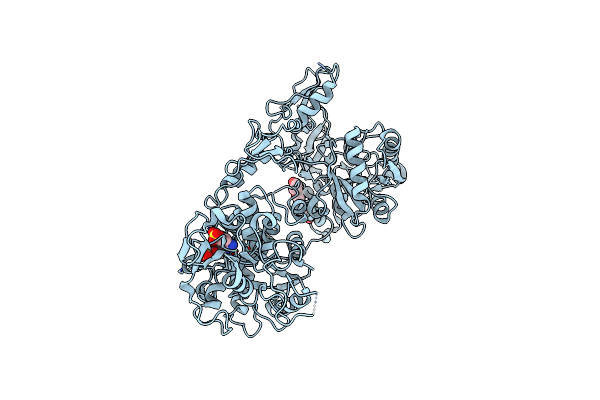
Organism: Nanorana parkeri
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-05-07 Classification: ANTITOXIN Ligands: 1PE, YGZ |
 |
Organism: Nanorana parkeri
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-05-07 Classification: ANTITOXIN Ligands: 1PE, YH9 |
 |
Organism: Nanorana parkeri
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2025-05-07 Classification: ANTITOXIN Ligands: 1PE, YGF |
 |
Organism: Nanorana parkeri
Method: X-RAY DIFFRACTION Resolution:1.95 Å Release Date: 2025-05-07 Classification: ANTITOXIN Ligands: 1PE, YGQ |
 |
Organism: Orthotospovirus tomatomaculae
Method: ELECTRON MICROSCOPY Release Date: 2025-04-16 Classification: VIRAL PROTEIN Ligands: RTP |
 |
Organism: Orthotospovirus tomatomaculae
Method: ELECTRON MICROSCOPY Release Date: 2025-01-29 Classification: VIRAL PROTEIN |
 |
Organism: Nocardioides sp. s-1144
Method: X-RAY DIFFRACTION Resolution:1.55 Å Release Date: 2024-12-04 Classification: HYDROLASE Ligands: SO4 |
 |
Structure Of A Solute Binding Protein From Desulfonauticus Sp. Bound To L-Tryptophan
Organism: Desulfonauticus sp. 38_4375
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2024-11-06 Classification: PEPTIDE BINDING PROTEIN Ligands: TRP, EDO, PEG, ACT, PG4 |

