Search Count: 5,394
All
Selected
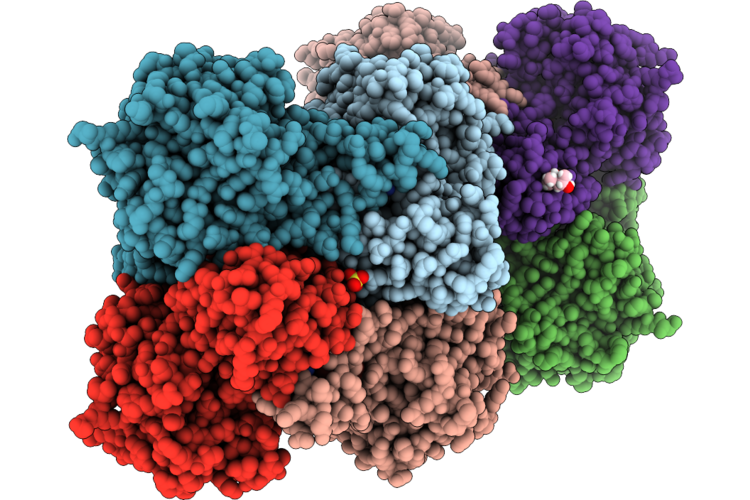 |
Crystal Structure Of Human S-Adenosyl-L-Homocysteine Hydrolase Complex With Adenosine
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.59 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: NAD, ADN, K, SO4, MPD |
 |
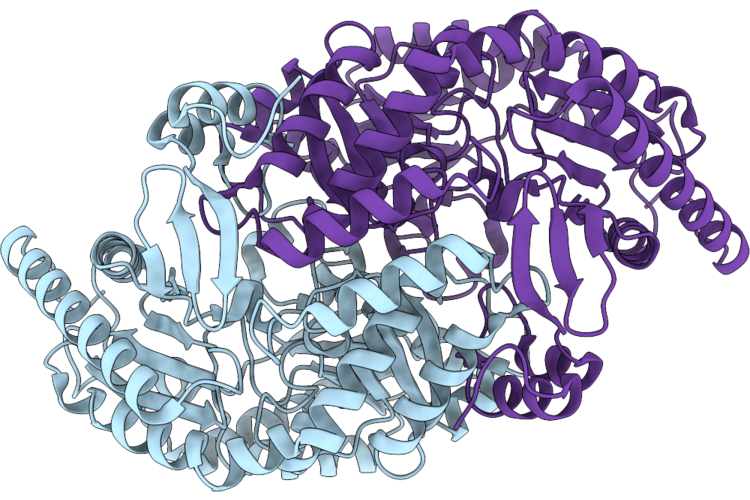
Crystal Structure Of Human S-Adenosyl-L-Homocysteine Hydrolase In Complex With Adenosine And Cadmium Ions.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.93 Å Release Date: 2026-04-29 Classification: HYDROLASE Ligands: NAD, ADN, CD, K |
 |
Organism: Phaeobacter porticola
Method: X-RAY DIFFRACTION Resolution:1.73 Å Release Date: 2026-04-29 Classification: TRANSFERASE |
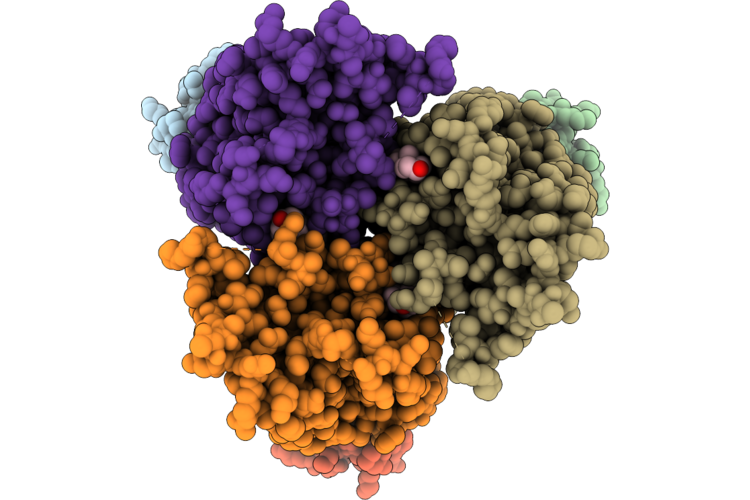 |
Structure Of Cxcr4 In Complex With A De-Novo Designed Mini-Protein Antagonist
Organism: Synthetic construct, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.28 Å Release Date: 2026-04-22 Classification: SIGNALING PROTEIN Ligands: CLR, D21 |
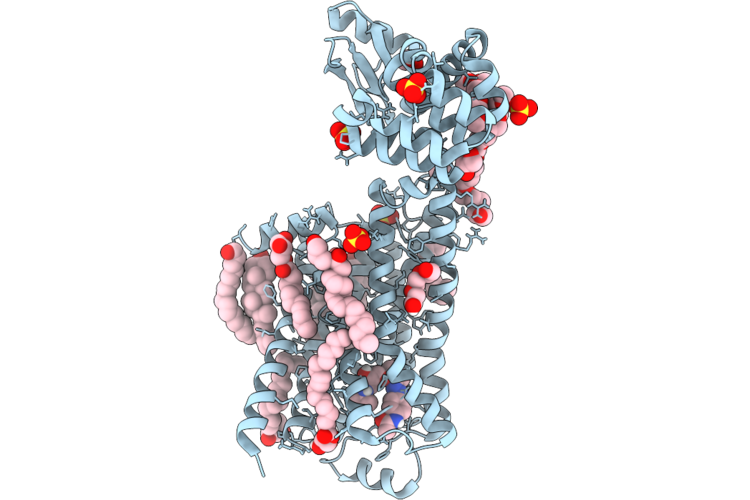 |
Dark Structure Of Beta-2 Adrenergic Receptor With Photoazolol In Dark State Recorded At Swissfel
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: SO4, CLR, 12P, OLC, 1PE, EDO, GOL, A1JHU, PLM |
 |
Mixed Model Refinement Of Beta-2 Adrenergic Receptor With Photoazolol In Dark State And Light State, 10 Seconds After Light Activation, Recorded At Swissfel
Organism: Homo sapiens, Tequatrovirus t4
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: SO4, ACM, CLR, PLM, 12P, OLC, 1PE, EDO, GOL, A1JHU |
 |
Dark Structure Of Beta-2 Adrenergic Receptor With Photoazolol In Dark State Recorded At Lcls
Organism: Homo sapiens, Enterobacteria phage t4
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: SO4, CLR, PLM, 12P, A1JHU, OLC, EDO, GOL |
 |
Mixed Model Refinement Of Beta-2 Adrenergic Receptor With Photoazolol In Dark State And Light State, 17 Nanoseconds After Light Activation, Recorded At Lcls
Organism: Homo sapiens, Tequatrovirus t4
Method: X-RAY DIFFRACTION Resolution:2.60 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: SO4, ACM, CLR, PLM, 12P, OLC, EDO, GOL, 1PE, A1JHU |
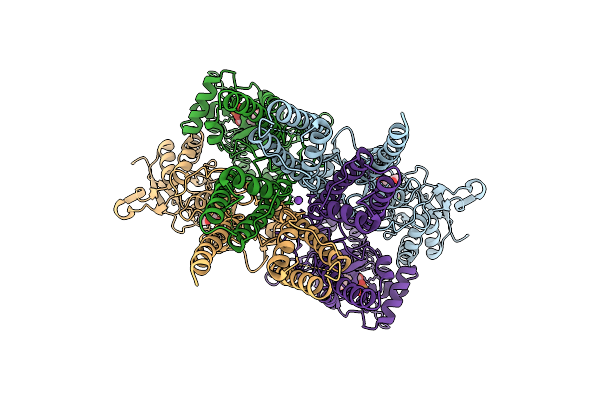 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: ZK1, NA |
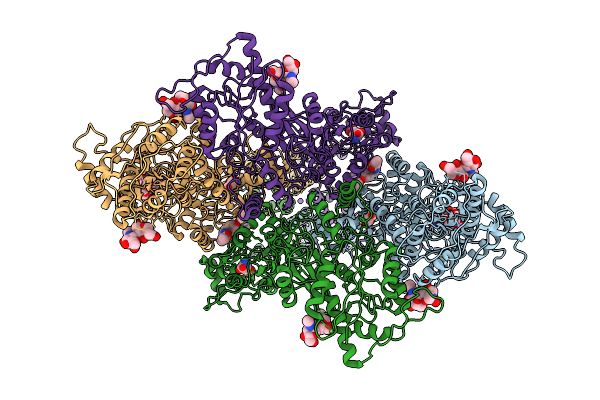 |
Composite Map Of Glua1/A2 In The Activated State, In Complex With Positive Allosteric Modulator (R,R)-2B And Agonist Glutamate (Atd-Lbd-Tmd)
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: GLU, NA, FWF |
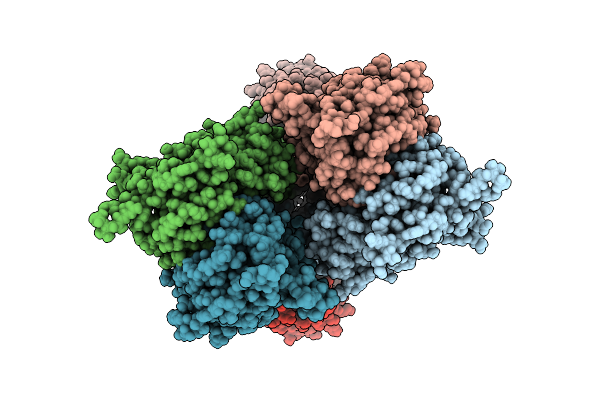 |
Heteromeric Glua1/A2-Cnih1 In The Activated State, Composite Map Of Lbd-Tmd
Organism: Rattus norvegicus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: GLU, POV, FWF, NA |
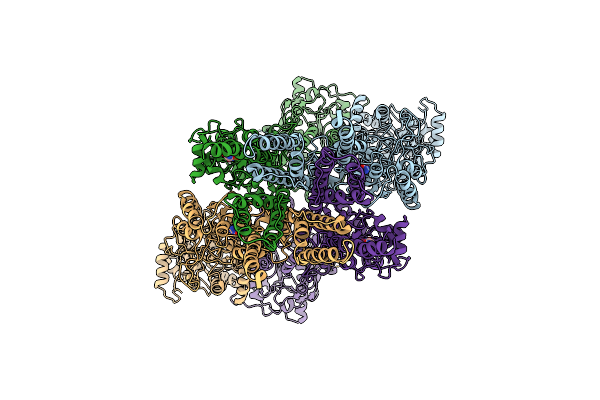 |
Heteromeric Glua1/A2 In The Desensitized State, Composite Map Of Atd-Lbd-Tmd
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: QUS |
 |
Crystal Structure Of The S-Adenosyl-L-Homocysteine Hydrolase From P. Aeruginosa, Q65N Mutant Probed With Rubidium To Confirm Disruption Of A Potassium Binding Site.
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.91 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAD, GOL, PO4 |
 |
Crystal Structure Of S-Adenosyl-L-Homocysteine Hydrolase From P. Aeruginosa: Q65A Mutant Soaked With Adenosine And Probed With Rubidium To Confirm Disruption Of A Potassium Binding Site.
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAD, ADN, PO4 |
 |
Crystal Structure Of S-Adenosyl-L-Homocysteine Hydrolase From P. Aeruginosa, Q65N Mutant Soaked With Adenosine And Probed With Rubidium To Confirm Disruption Of A Potassium Binding Site.
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:2.34 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAD, ADN, PO4 |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-04-08 Classification: RNA BINDING PROTEIN Ligands: ZN, MG |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.70 Å Release Date: 2026-04-08 Classification: RNA BINDING PROTEIN Ligands: MG, ZN |
 |
Organism: Chlamydomonas reinhardtii
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-04-01 Classification: RIBOSOME Ligands: MG, K, CLM |
 |
Organism: Chlamydomonas reinhardtii
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-01 Classification: RIBOSOME Ligands: K, MG |
 |
Crystal Structure Of Feruloyl Esterase From Fusarium Oxysporum G122S Variant In Complex With Benzoic Acid
Organism: Fusarium oxysporum
Method: X-RAY DIFFRACTION Resolution:1.71 Å Release Date: 2026-04-01 Classification: HYDROLASE Ligands: NAG, PEG, EDO, BEZ, CA |

