Search Count: 8,297
All
Selected
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: XDC, ZN, MG, CA, GOL, NA, NAG, MAN, PO4, TRS, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PO4, ZN, MG, CA, NA, NAG, GOL, TRS, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
Tissue Non-Specific Alkaline Phosphatase -S110A Bound To Ppi With Ethylene Glycol
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: POP, ZN, MG, CA, NA, NAG, MAN, PO4, FLC |
 |
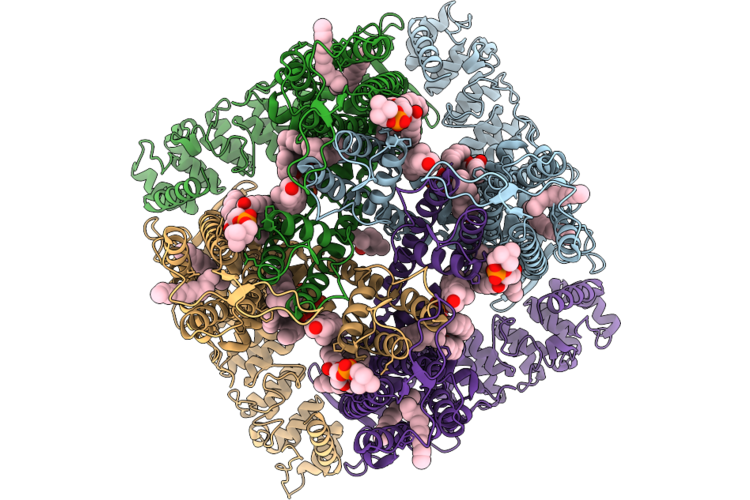
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ATP, ZN, MG, CA, GOL, NAG, FLC, PO4 |
 |
Organism: Oryctolagus cuniculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: ERG, CPL, PIO, IT9 |
 |
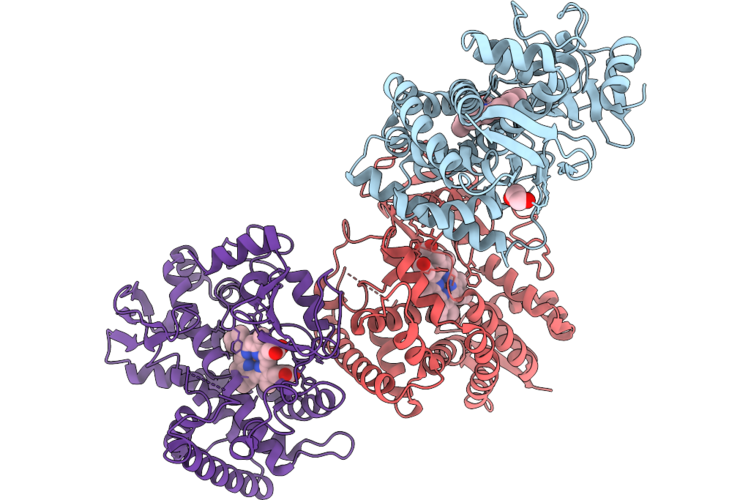
Organism: Micromonospora rosaria
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
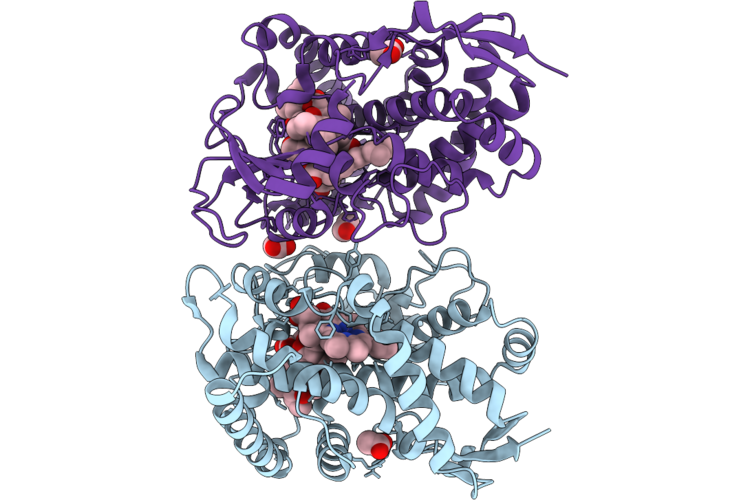
Organism: Micromonospora rosaria
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, EDO |
 |
Organism: Micromonospora rosaria
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: A1MAI, HEM, ACT, MLT, EDO |
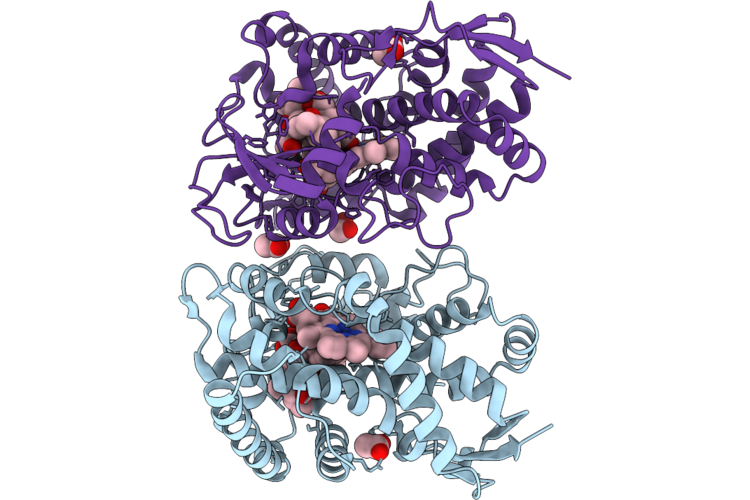 |
Crystal Structure Of The Cytochrome P450 Rosc In Complexed With 20-Dihydrorosamicin
Organism: Micromonospora rosaria
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: L0O, HEM, ACT, MLT, EDO |
 |
Crystal Structure Of The Cytochrome P450 Rosc In Complexed With 20-Deoxo-20-Dihydrorosamicin
Organism: Micromonospora rosaria
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: A1MAJ, HEM, ACT, MLT, EDO |
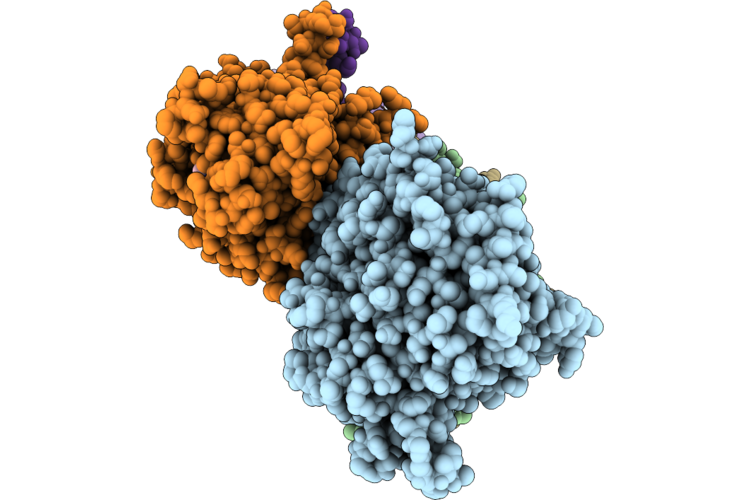 |
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MOTOR PROTEIN Ligands: GTP, MG, GDP, TA1, ANP |
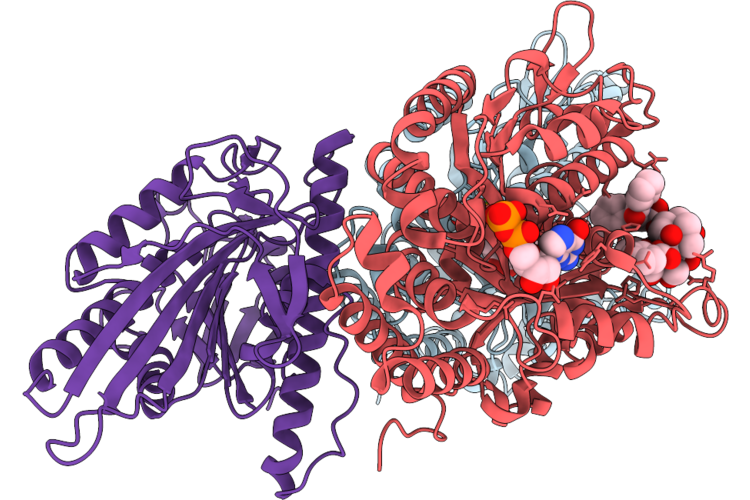 |
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-04-22 Classification: MOTOR PROTEIN Ligands: GTP, MG, GDP, TA1 |
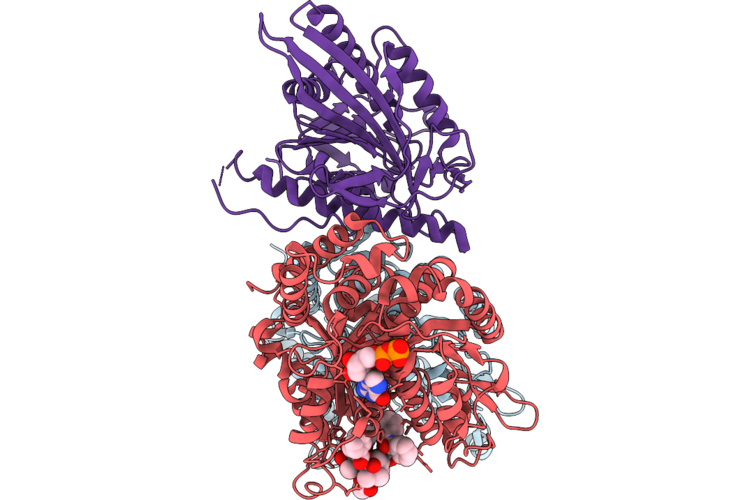 |
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MOTOR PROTEIN Ligands: GTP, MG, GDP, TA1 |
 |
Organism: Homo sapiens, Sus scrofa
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MOTOR PROTEIN Ligands: MG, ANP, GTP, GDP, TA1 |
 |
Crystal Structure Of Putative L-Amino Acid N-Acyltransferase Mnat From Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN Ligands: EDO |
 |
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica Bound To C-Terminal Region Of Collagen Model Peptide (Pro-Hyp-Gly)10
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |
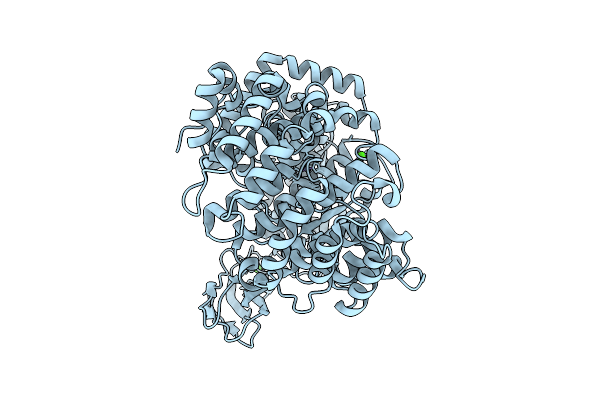 |
Organism: Hathewaya histolytica
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: ZN, CA |
 |
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica In Complex With Collagen Model Peptide (Pro-Hyp-Gly)10
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |
 |
Cryo-Em Structure Of Collagenase H (E416Q Mutant) From Hathewaya Histolytica In Complex With Collagen Model Peptide (Pro-Hyp-Gly)12
Organism: Hathewaya histolytica, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: METAL BINDING PROTEIN Ligands: CA, ZN |

