Search Count: 22,344
All
Selected
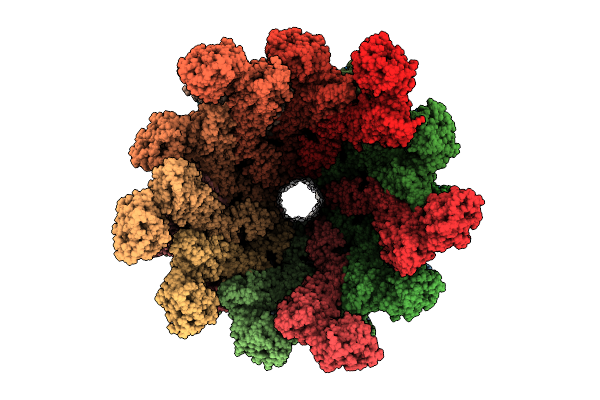 |
Organism: Vibrio alginolyticus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: STRUCTURAL PROTEIN |
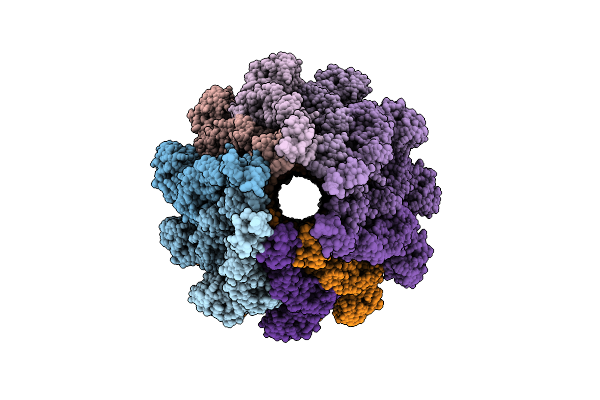 |
Organism: Vibrio alginolyticus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: STRUCTURAL PROTEIN |
 |
Atomic Model Of Cx43 Gap Junction Channel Rigid-Body Fitted To The In Situ Structure Of The Human Cx43 Gap Junction
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: MEMBRANE PROTEIN |
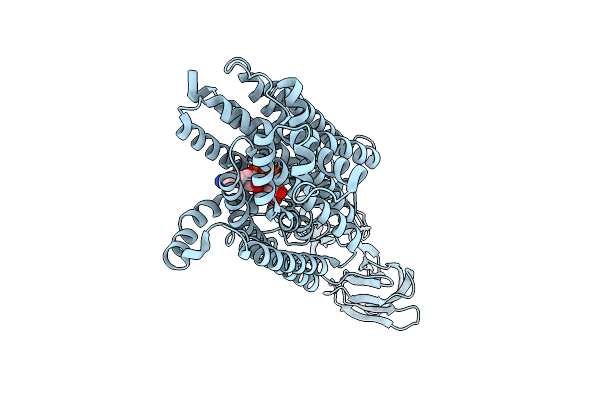 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.38 Å Release Date: 2026-04-15 Classification: MEMBRANE PROTEIN Ligands: NAD |
 |
Organism: Zika virus
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN Ligands: MG, EDO |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: ZN |
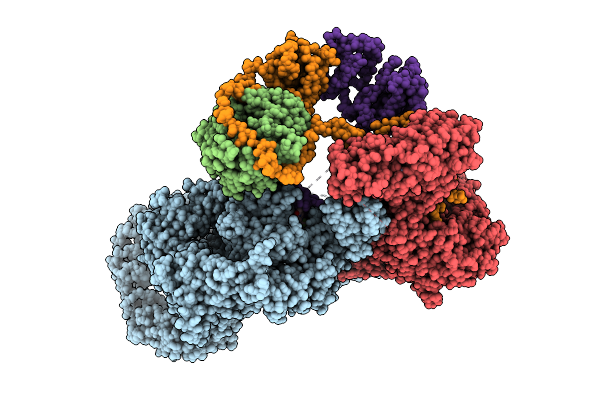 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: ZN, A1AID |
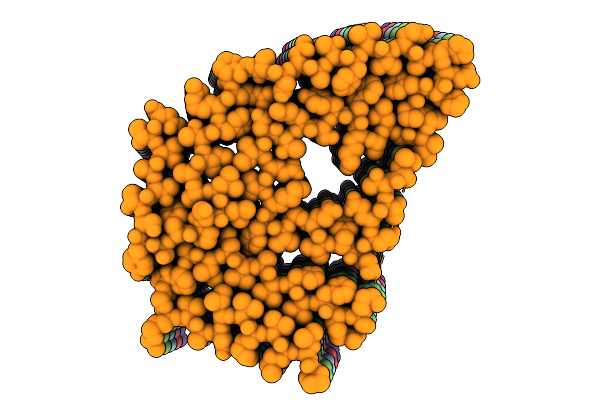 |
Cryo-Em Structure Of Hepatic Amyloid Fibril From A Variant Attrv122Delta, Single Filament Morphology
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-15 Classification: PROTEIN FIBRIL |
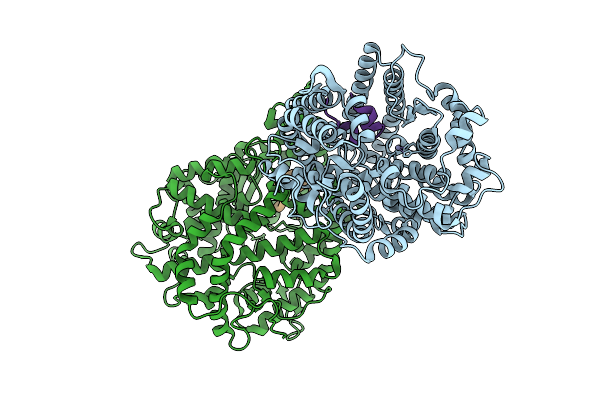 |
Organism: Homo sapiens, Synthetic plasmid
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: ZN |
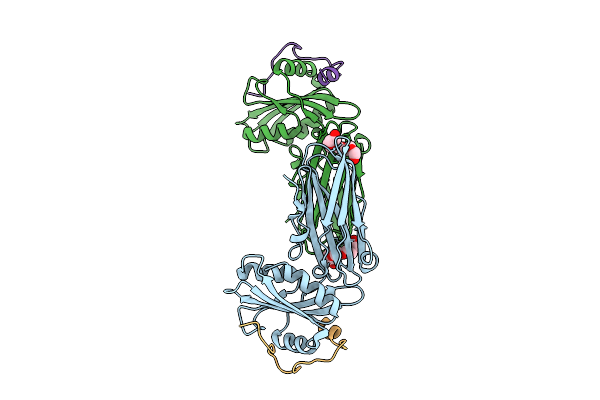 |
Structure Of Alpha Appendage Of Ap2 Bound To The Extended Fxdxf Motif Derived Of Ccdc32
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-08 Classification: ENDOCYTOSIS Ligands: 1PE |
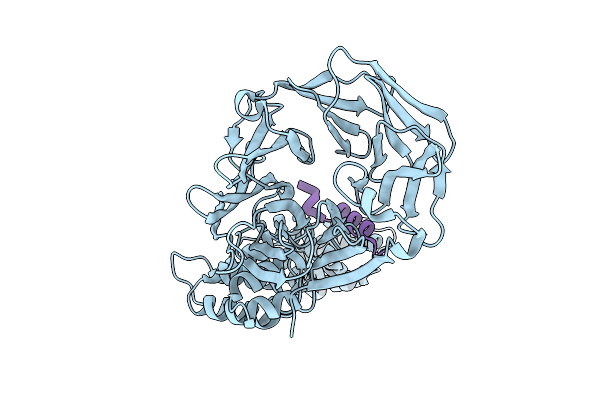 |
Crystal Structure Of Clathrin Heavy Chain In Complex With A Peptidomimetic Inhibitor Of The Tacc3 Interaction
Organism: Rattus norvegicus, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-08 Classification: CELL CYCLE |
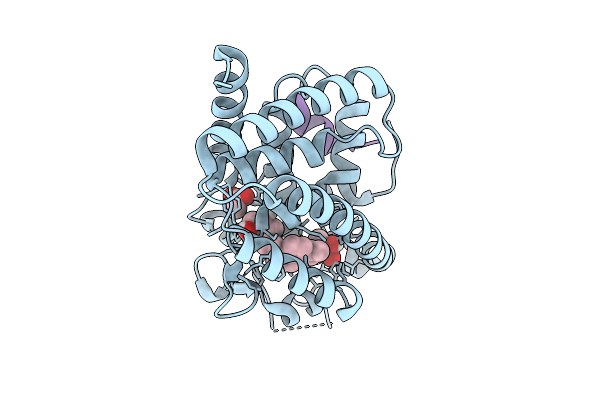 |
Organism: Danio rerio, Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-04-08 Classification: TRANSCRIPTION Ligands: ACT, A1JD0 |
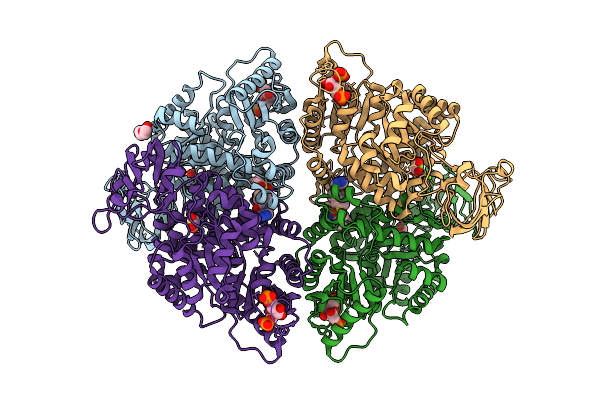 |
Liver Pyruvate Kinase In Complex With A Phthalazine-Based Fluorescent Probe Iv
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.62 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: MG, A1JEB, OXL, FBP, PEG |
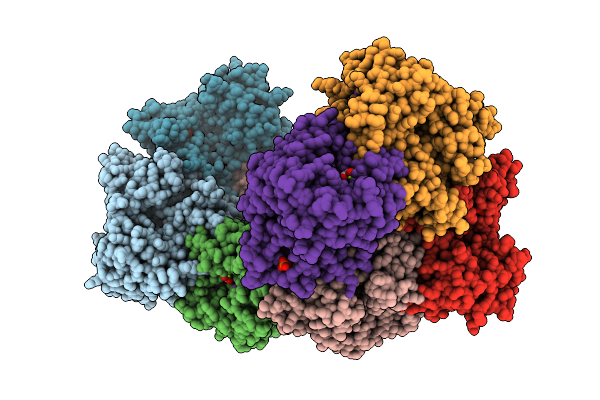 |
Structure Of Liver Pyruvate Kinase In Complex With Liver Pyruvate Kinase In Complex With Fluorescent Probe I
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: FBP, OXL, MG, K, A1JFN |
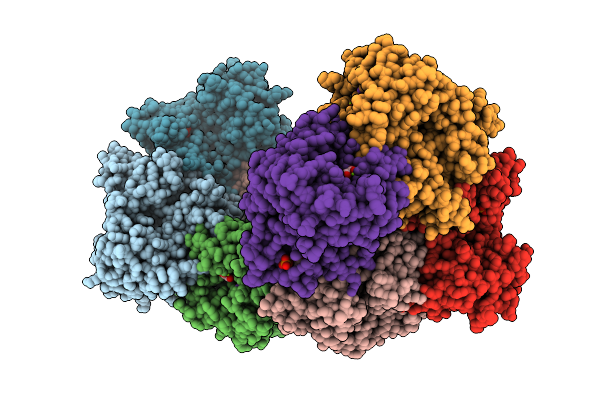 |
Structure Of Liver Pyruvate Kinase In Complex With Liver Pyruvate Kinase In Complex With Fluorescent Probe Ii
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.27 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: FBP, OXL, MG, K, A1JFT |
 |
Organism: Homo sapiens, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.39 Å Release Date: 2026-04-08 Classification: HYDROLASE |
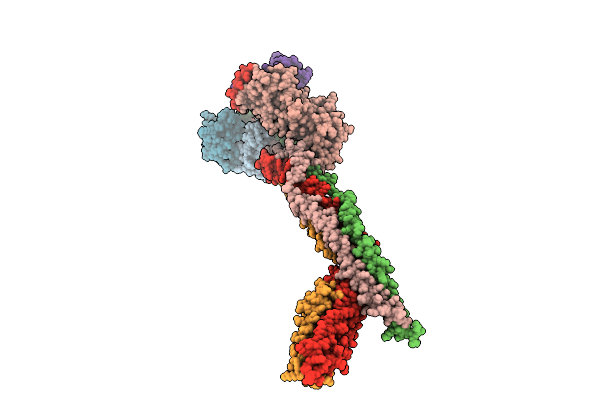 |
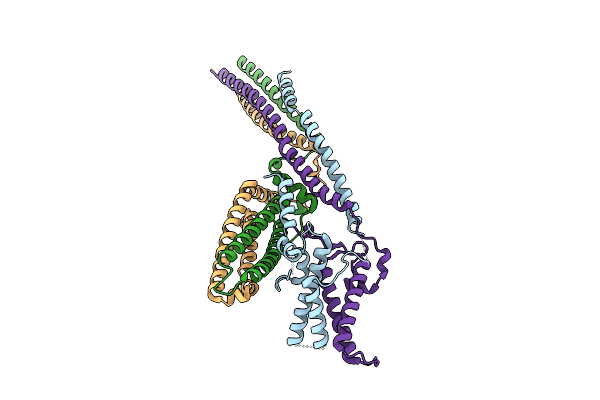
Cryo-Em Structure Of The Saccharomyces Cerevisiae Kmn Junction Complex Lacking The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
 |
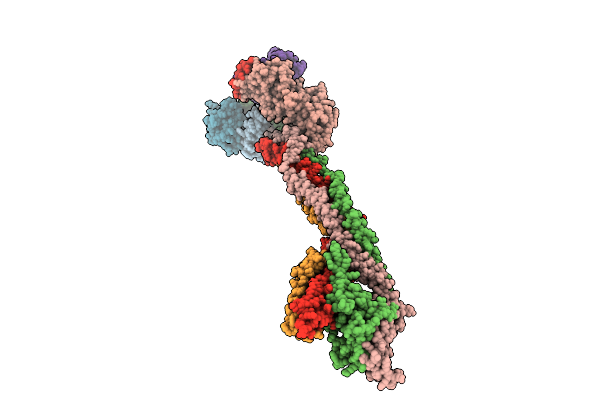
Cryo-Em Structure Of The Base Of The Saccharomyces Cerevisiae Kmn Junction Complex Containing The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
 |
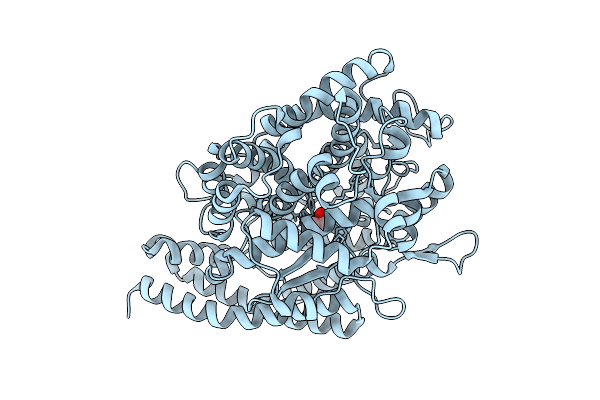
Cryo-Em Structure Of The Saccharomyces Cerevisiae Kmn Junction Complex Containing The Mis12C(Mtw1C) Head 2 Domain
Organism: Saccharomyces cerevisiae, Saccharomyces cerevisiae s288c
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: CELL CYCLE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: MLA, ZN |

