Search Count: 22,130
All
Selected
 |
Crystal Structure Of Putative L-Amino Acid N-Acyltransferase Mnat From Pseudomonas Aeruginosa
Organism: Pseudomonas aeruginosa pao1
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN Ligands: EDO |
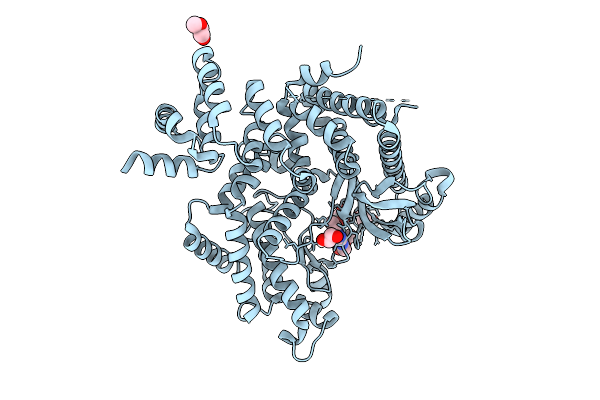 |
The Structure Of Human Vacuolar Protein Sorting 34 Catalytic Domain Bound To Rd-I-137
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: A1CD6, GOL, PEG, CL |
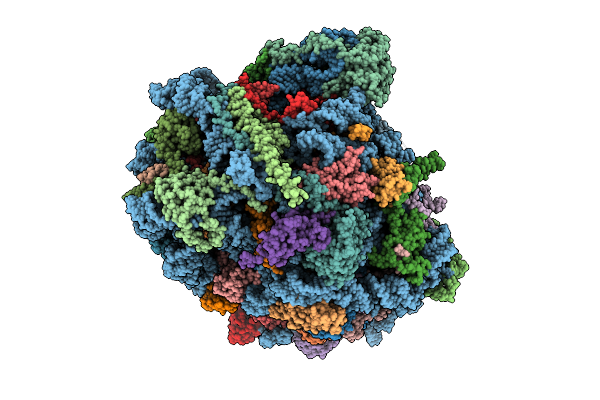 |
In Vitro Reconstituted Complex Of Purified S. Pombe Large Ribosomal Subunit And Snor
Organism: Schizosaccharomyces pombe 972h-
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: RIBOSOME Ligands: ZN |
 |
Crystal Structure Of An Engineered Tpr Domain In Complex With The Hsp90 Peptide Meevd
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.13 Å Release Date: 2026-04-15 Classification: CHAPERONE Ligands: SO4 |
 |
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-15 Classification: OXIDOREDUCTASE Ligands: HEM, MG, O |
 |
Protein Structure Of The First Glycoside Hydrolase Family 30, Subfamily 12 Endoxylanase
Organism: Anaerobacterium chartisolvens
Method: X-RAY DIFFRACTION Resolution:1.92 Å Release Date: 2026-04-15 Classification: HYDROLASE Ligands: GOL, FMT, ACY, EDO, NA, PEG |
 |
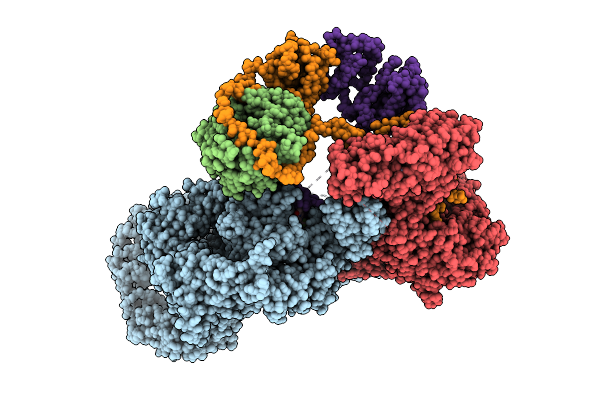
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: ZN |
 |
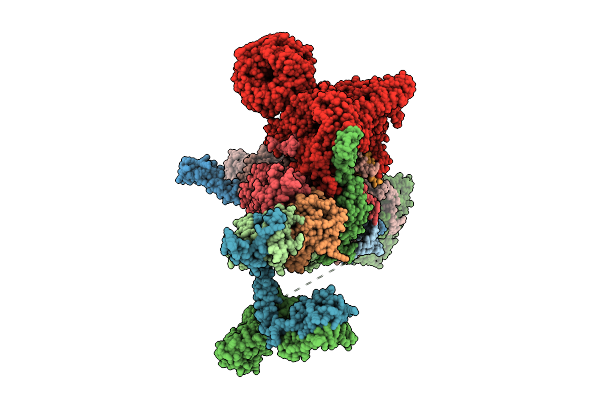
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-04-15 Classification: TRANSFERASE Ligands: ZN, A1AID |
 |
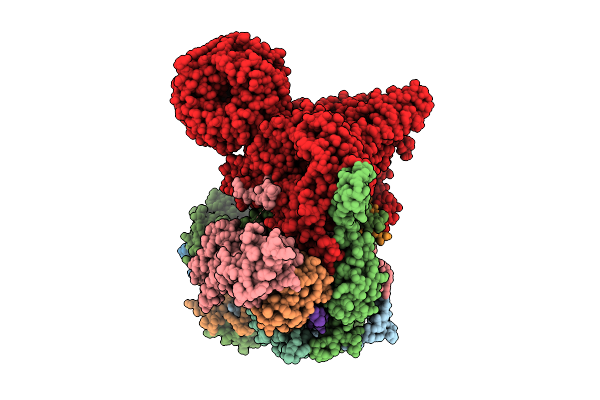
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Proximal Dhx15 State)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
 |
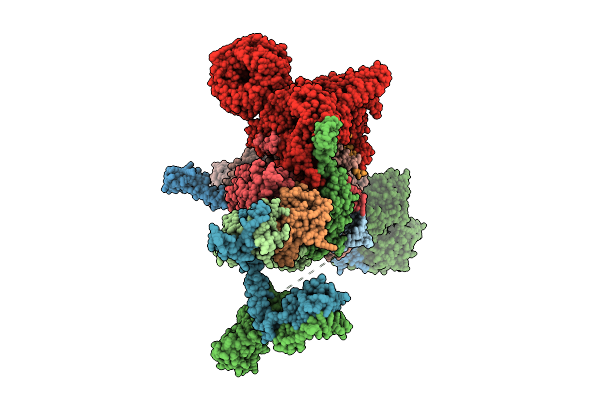
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Core)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
 |
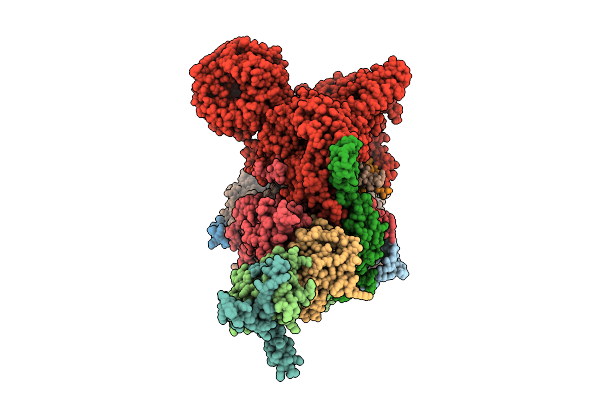
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Distal Dhx15 State)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
 |
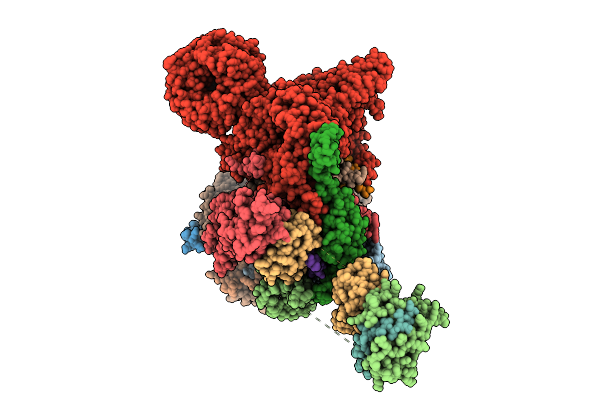
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Sf3A State 1)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
 |
Cryo-Em Structure Of The Endogenous U2/Branchpoint Spliceosomal Complex (Sf3A State 2)
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: SPLICING Ligands: ZN |
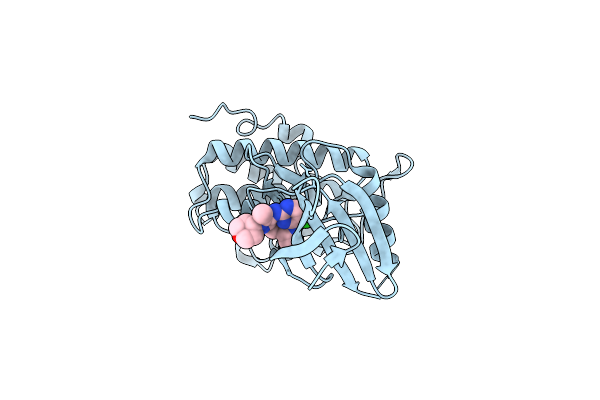 |
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 1 ((11R)-8-Chloro-3,11-Dimethyl-2-(Oxan-4-Yl)-2,4,10,11,12,13-Hexahydro-9,5-(Azeno)Pyrazolo[3,4-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5F |
 |
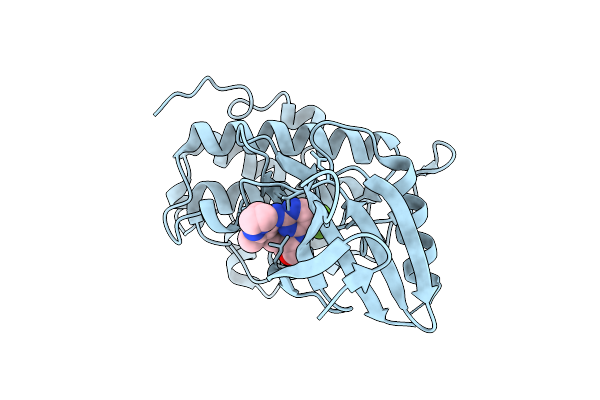
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 7 ((10As,13As)-3-Cyclobutyl-1-Methyl-8-(Trifluoromethyl)-3,4,10A,11,13A,14-Hexahydro-10H,13H-9,5-(Azeno)Furo[3,4-K]Pyrazolo[4,3-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5G |
 |
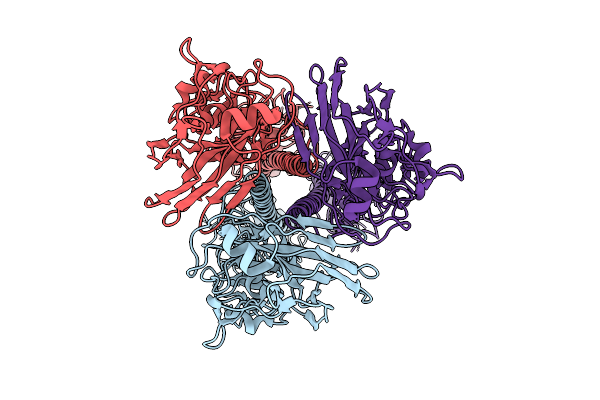
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 12 ((10As,13As)-3-Cyclopropyl-1-Methyl-8-(Trifluoromethyl)-3,4,10A,11,13A,14-Hexahydro-10H,13H-9,5-(Azeno)Furo[3,4-K]Pyrazolo[4,3-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5H |
 |
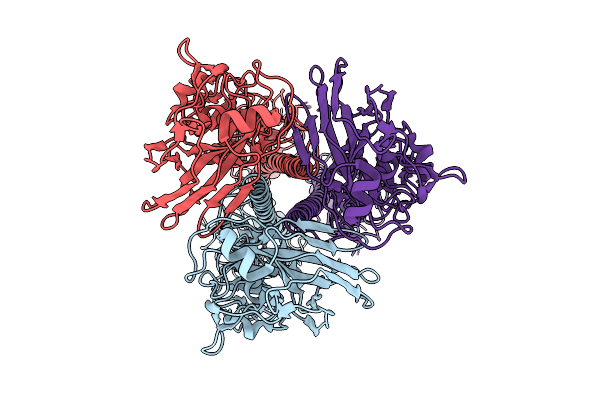
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.39 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.16 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.67 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
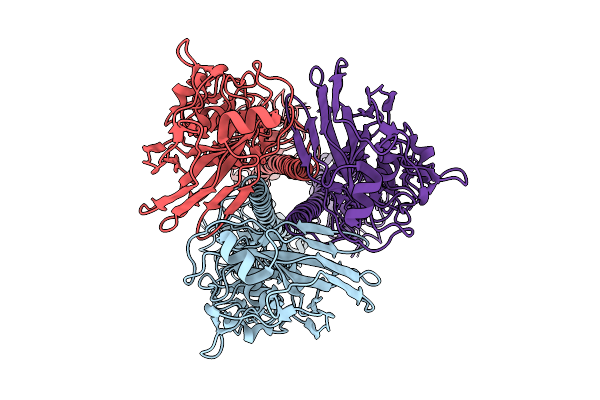 |
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.47 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |

