Search Count: 54,514
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2026-04-22 Classification: STRUCTURAL PROTEIN Ligands: SO4, CL |
 |
Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans.
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Reprocessing And Re-Refinement Of Damage Free Ferric State Of Dye Type Peroxidase Aa From Streptomyces Lividans
Organism: Streptomyces lividans 1326
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM |
 |
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN Ligands: EDO |
 |
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN |
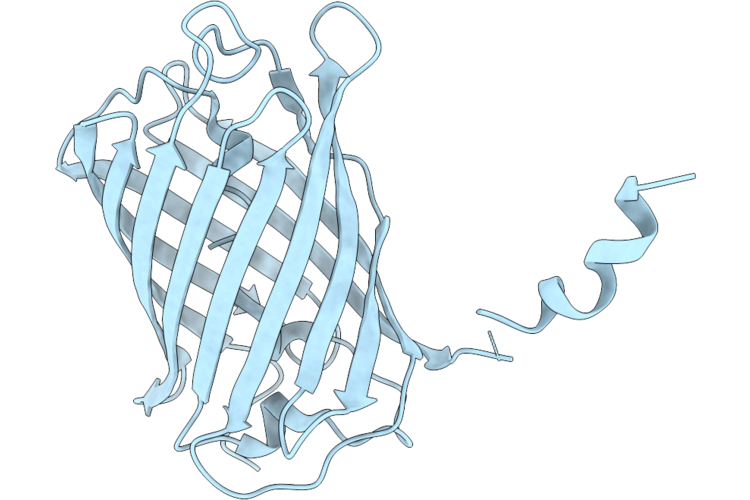 |
Crystal Structure Of Lssmorange Under Room Temperature At Ph 8.0, 250Ps After Pump Laser, Refined Against 8.33 Times Extrapolated Structure Factor
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN |
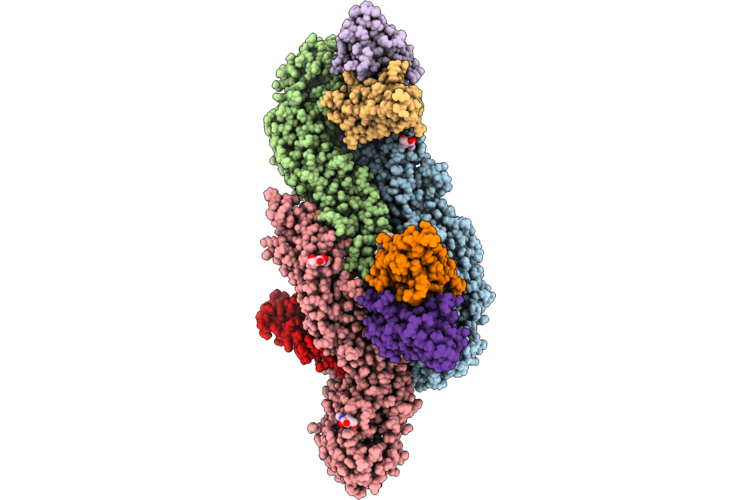 |
Organism: Homo sapiens, Dengue virus type 2
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: VIRUS Ligands: NAG |
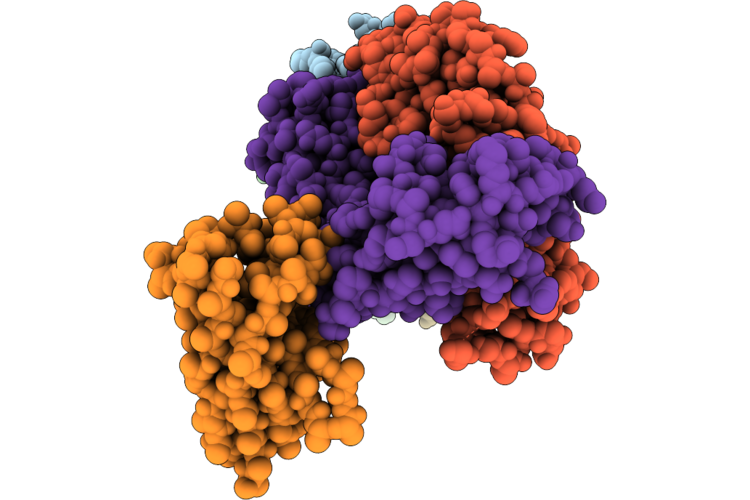 |
Organism: Homo sapiens, Lama glama
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: IMMUNE SYSTEM/Hydrolase |
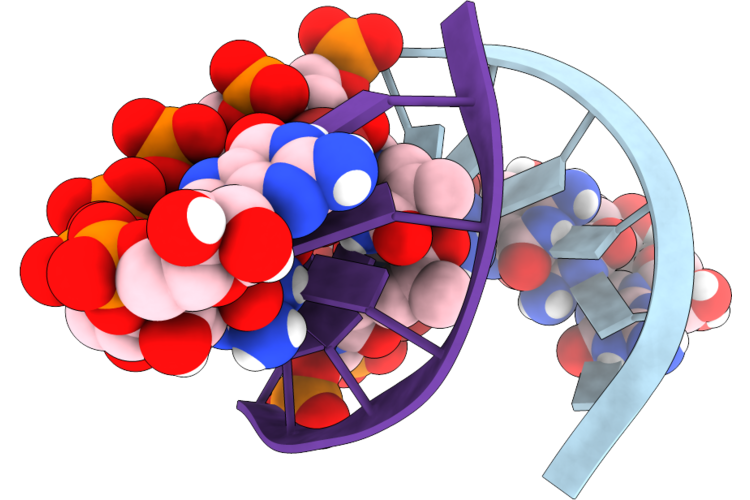 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-04-22 Classification: RNA Ligands: A1B9T, 5BU, 5GP, C5P, C |
 |
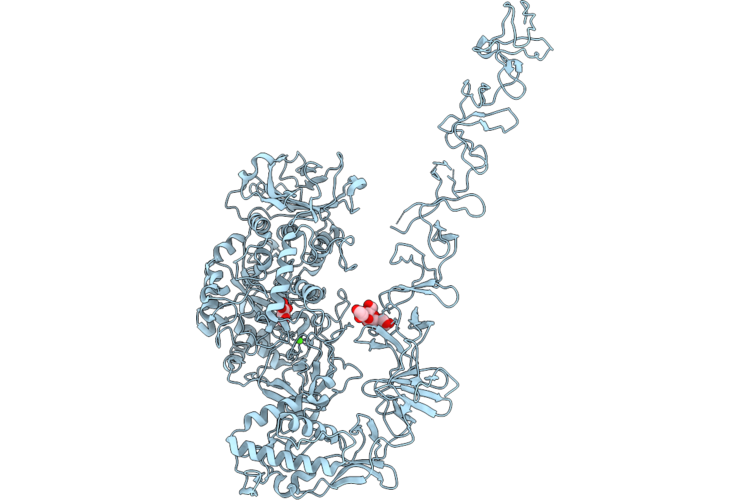
Structure Of Full-Length Streptococcus Mutans Gtfd Active Site Mutant (D465A, D584A) In Complex With Sucrose
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: EDO |
 |
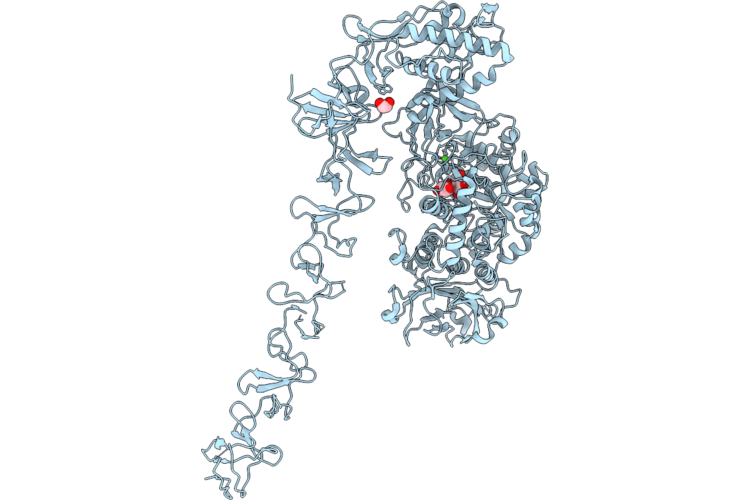
Structure Of Full-Length Streptococcus Mutans Gtfd Active Site Mutant (D465A, D584A) With Active Site Glucose
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: CA, GLC, EDO |
 |
Structure Of Full-Length Streptococcus Mutans Gtfd With Inhibitor Maltose In Active Site
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: CA, EDO |
 |
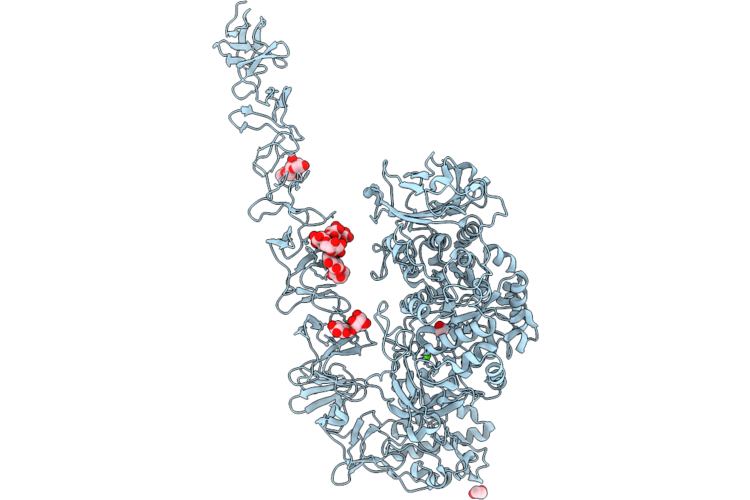
Structure Of Streptococcus Mutans Gtfd With C-Terminally Truncated Domain V In Complex With Acarbose
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:3.15 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: CA |
 |
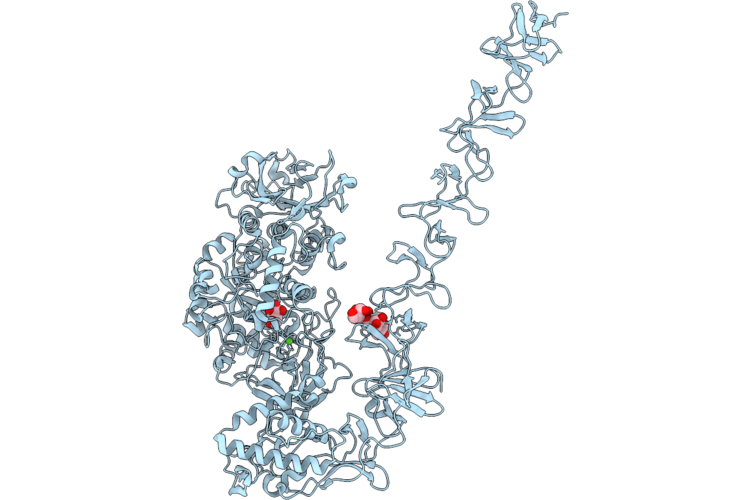
Structure Of Full-Length Streptococcus Mutans Gtfd In Complex With Dextran 1000 In Domain V
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: CA, EDO |
 |
Structure Of Full-Length Streptococcus Mutans Gtfd In Complex With Dextran 5000 In Domain V
Organism: Streptococcus mutans
Method: X-RAY DIFFRACTION Resolution:2.35 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: CA, GLC |
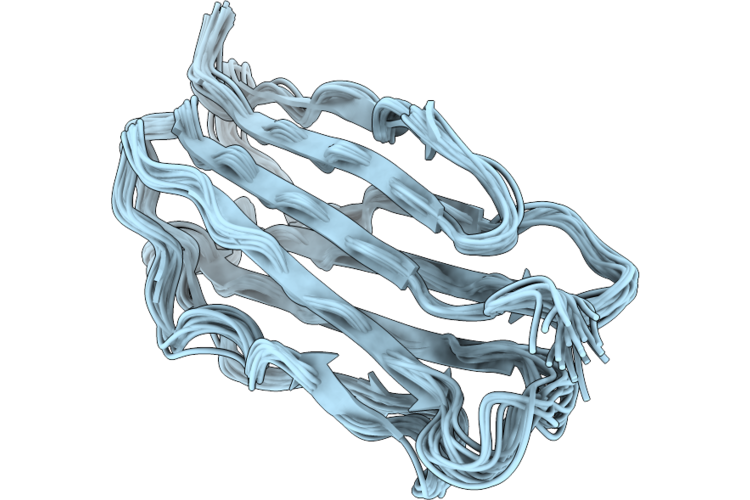 |
Organism: Mattirolomyces terfezioides
Method: SOLUTION NMR Release Date: 2026-04-22 Classification: UNKNOWN FUNCTION |
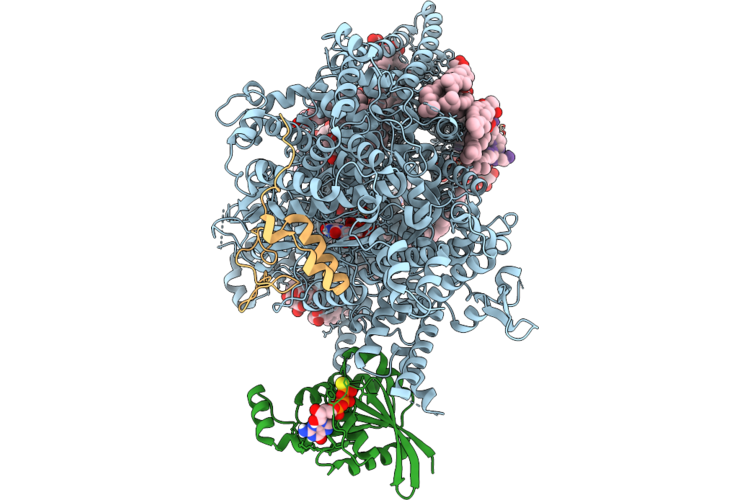 |
Structure Of Beta-1,3-Glucan Synthase In Complex With Caspofungin, Rho1 And Long Glucan
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP, GSP, A1CHR |
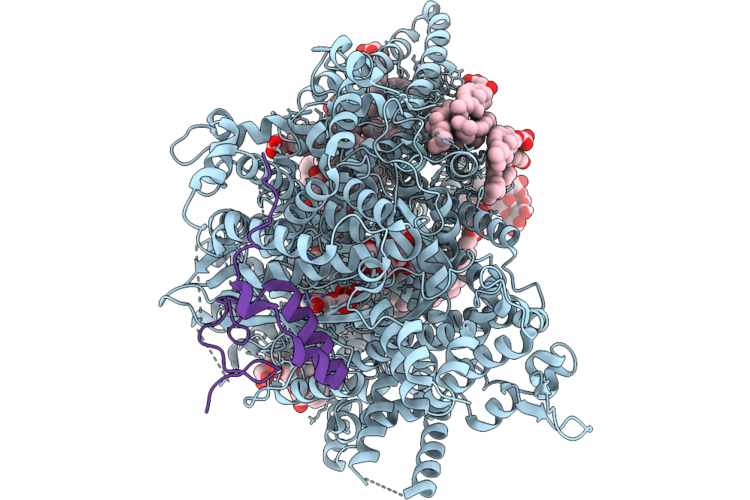 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) In Complex With Short Glucan
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP |
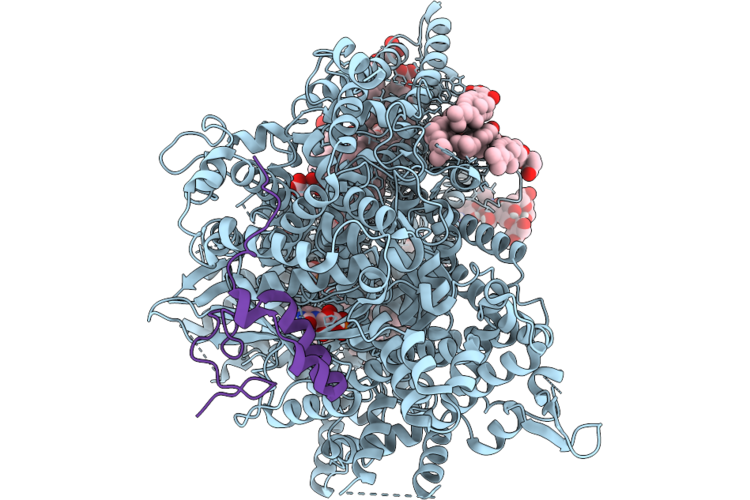 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Relevant Ground State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP |
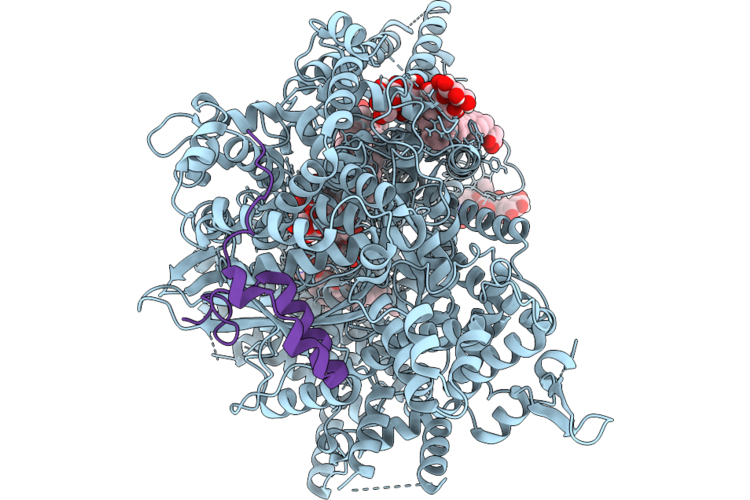 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Less Relevant L2 State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: Y01, 3PE, LMN |

