Search Count: 32,284
All
Selected
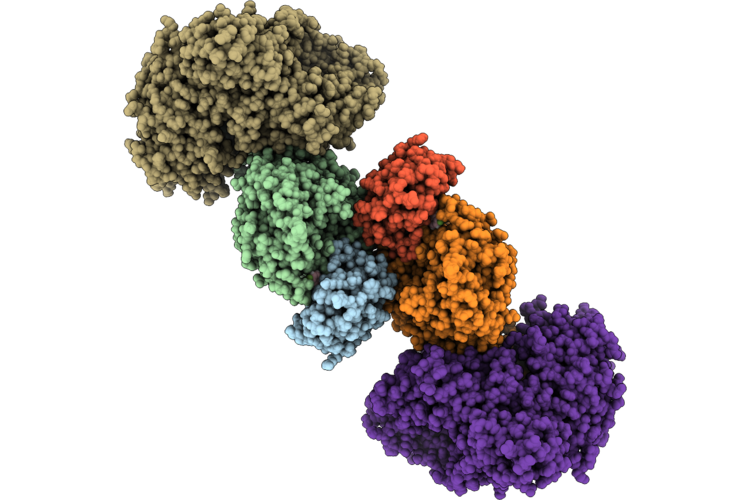 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: CYTOSOLIC PROTEIN Ligands: A1C4S, ZN |
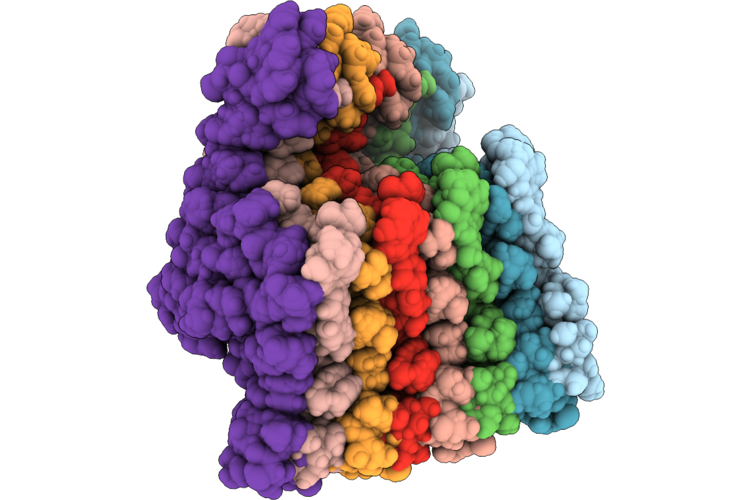 |
Ssnmr Structure Of Anti-Necroptosis Viral:Human Functional Hetero-Amyloid M45:Ripk3
Organism: Homo sapiens, Murid betaherpesvirus 1
Method: SOLID-STATE NMR Release Date: 2026-04-22 Classification: PROTEIN FIBRIL |
 |
Organism: Mycobacterium phage ds6a
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-04-22 Classification: VIRUS Ligands: ZN, PO4 |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PEG, DSN, CA |
 |
Organism: Mycobacterium phage ds6a
Method: X-RAY DIFFRACTION Resolution:2.75 Å Release Date: 2026-04-22 Classification: HYDROLASE |
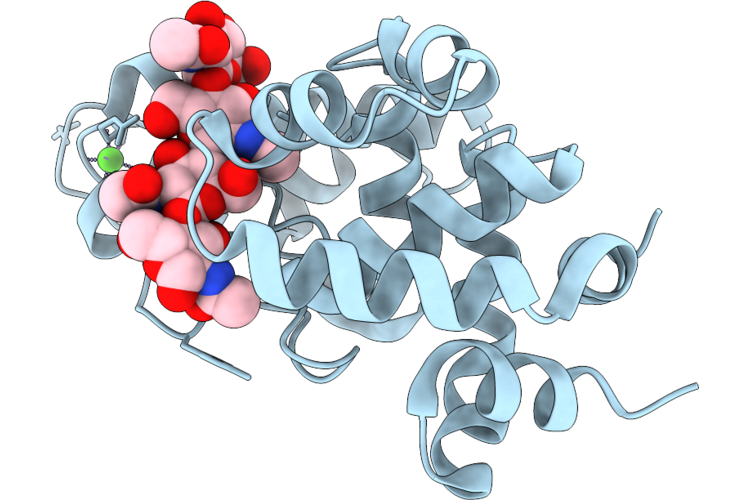 |
Crystal Structure Of Gh19 E228Q Domain Of D29-Lysa In Complex With (Glcnac)4
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: CA |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:1.63 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: MES, CA, MPD |
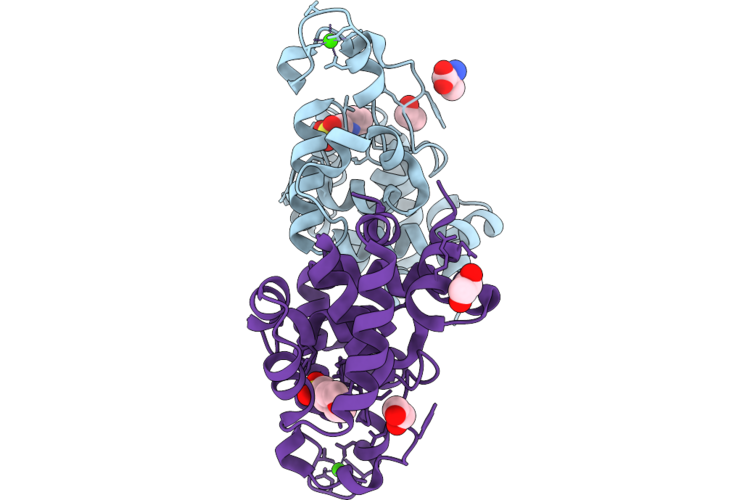 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: MES, CA, EDO, ALA |
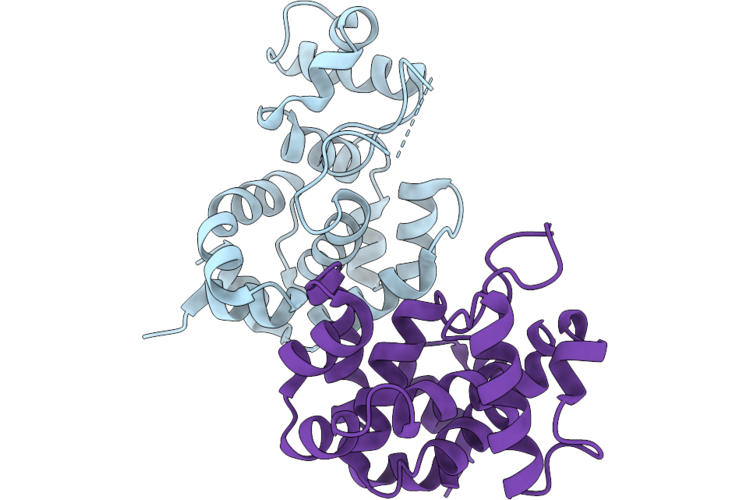 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.04 Å Release Date: 2026-04-22 Classification: HYDROLASE |
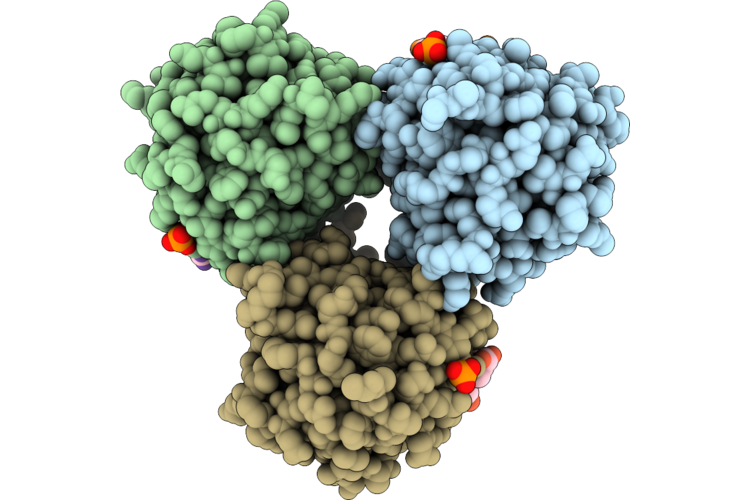 |
Crystal Structure Of Ami2B Domain Of Ds6A-Lysa In Complex With L-Ala-D-Iso-Gln-L-Lys- D-Ala-D-Ala
Organism: Mycobacterium phage ds6a, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PO4, ZN |
 |
Crystal Structure Of Gh19 Domain Of D29-Lysa In Complex With Product (Glcnac)
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: NAG |
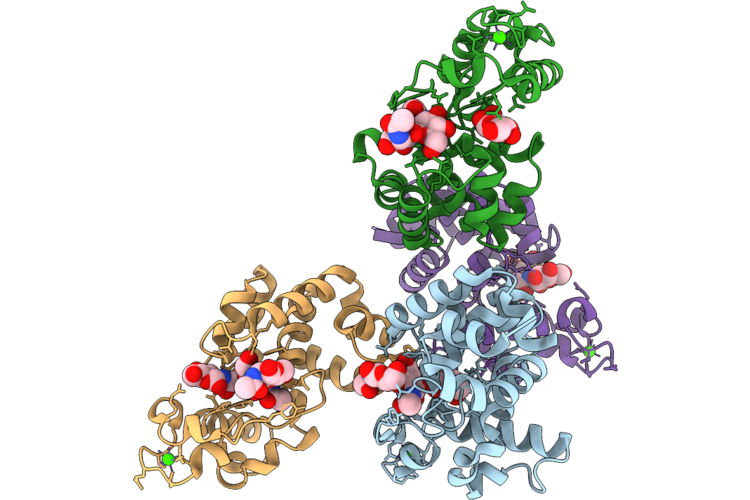 |
Crystal Structure Of Gh19 Domain Of D29-Lysa In Complex With Products (Glcnac Y Glcnac2)
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: NAG, CA |
 |
Organism: Mycobacterium phage d29
Method: X-RAY DIFFRACTION Resolution:3.20 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
Crystal Structure Of A P. Aeruginosa Gyrase Chimera In Complex With 20Mer Dna And Ciprofloxacin
Organism: Pseudomonas aeruginosa, Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-04-22 Classification: DNA BINDING PROTEIN Ligands: CIT, CPF, SO4, MN, CL |
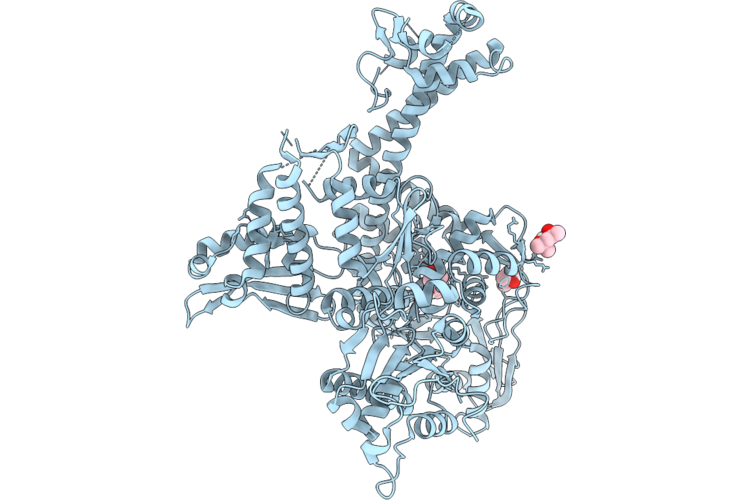 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-22 Classification: DNA BINDING PROTEIN |
 |
Organism: Periplaneta americana
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: GDS, EDO, MG, MPD, NA |
 |
Organism: Periplaneta americana
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: GSH, EDO, MG, CL |
 |
Organism: Blattella germanica
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: GDS, GOL, GSH |
 |
Organism: Blattella germanica
Method: X-RAY DIFFRACTION Resolution:1.33 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: GSH, EDO, CA |
 |
Organism: Mattirolomyces terfezioides
Method: SOLUTION NMR Release Date: 2026-04-22 Classification: UNKNOWN FUNCTION |

