Search Count: 1,19,578
All
Selected
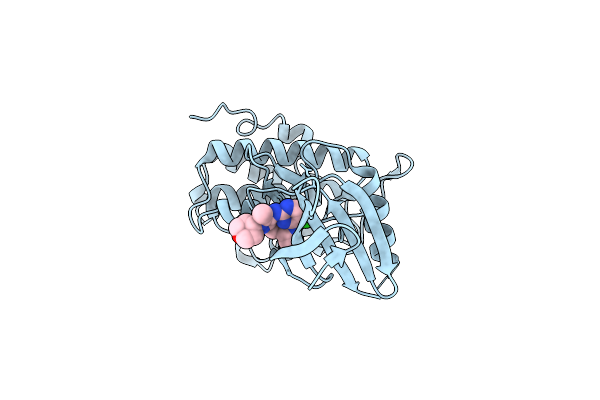 |
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 1 ((11R)-8-Chloro-3,11-Dimethyl-2-(Oxan-4-Yl)-2,4,10,11,12,13-Hexahydro-9,5-(Azeno)Pyrazolo[3,4-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5F |
 |
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 7 ((10As,13As)-3-Cyclobutyl-1-Methyl-8-(Trifluoromethyl)-3,4,10A,11,13A,14-Hexahydro-10H,13H-9,5-(Azeno)Furo[3,4-K]Pyrazolo[4,3-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5G |
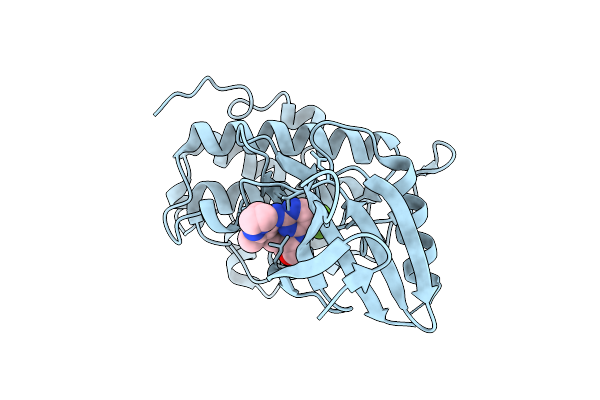 |
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 12 ((10As,13As)-3-Cyclopropyl-1-Methyl-8-(Trifluoromethyl)-3,4,10A,11,13A,14-Hexahydro-10H,13H-9,5-(Azeno)Furo[3,4-K]Pyrazolo[4,3-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5H |
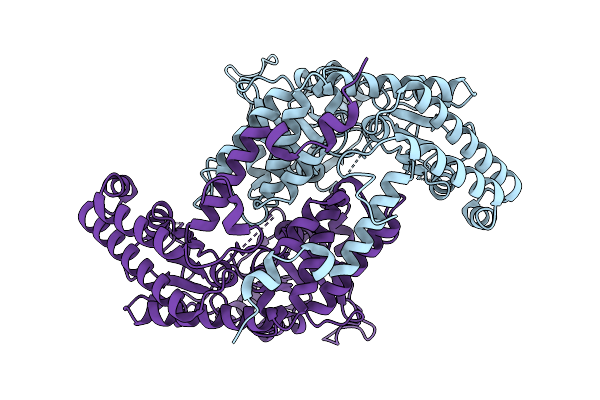 |
Cryo-Em Structure Of The A149T Dimer Variant Of Serine Hydroxymethyltransferase 8 From Soybean Cultivar Essex In Complex With Plp
Organism: Glycine max
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: GG5, A1C99, ZN |
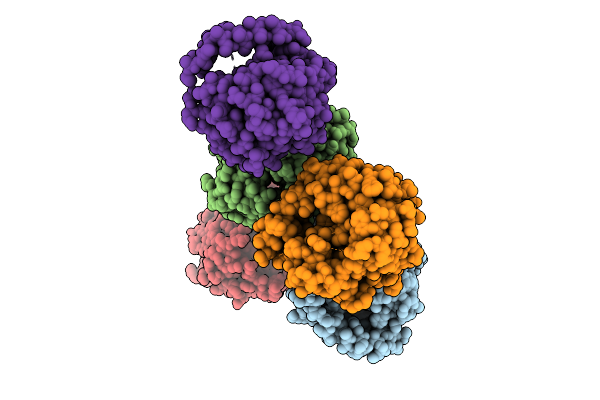 |
Crystal Structure Of Apo Alpha/Beta-Hydrolase Macrolide Esterase Estt From Sphingobacterium Faecium (S126A Mutant)
Organism: Sphingobacterium faecium
Method: X-RAY DIFFRACTION Resolution:3.19 Å Release Date: 2026-04-08 Classification: HYDROLASE |
 |
Crystal Structure Of Apo Alpha/Beta-Hydrolase Macrolide Esterase Estt From Sphingobacterium Thalpophilum (S102A Mutant)
Organism: Sphingobacterium thalpophilum
Method: X-RAY DIFFRACTION Resolution:2.18 Å Release Date: 2026-04-08 Classification: HYDROLASE |
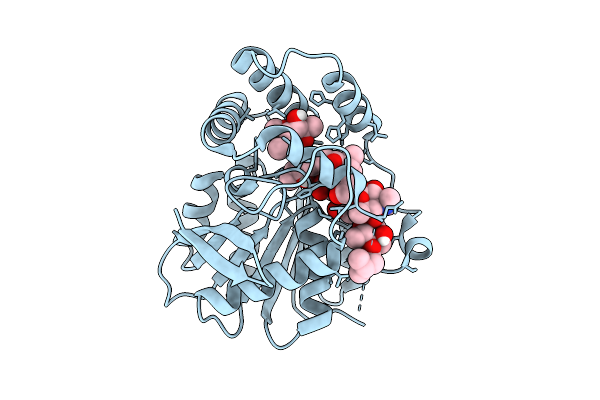 |
Crystal Structure Of Alpha/Beta-Hydrolase Macrolide Esterase Estt From Bacillus Cereus (S102A Mutant) In Complex With Linearized Tylvalosin
Organism: Bacillus cereus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: A1DAR |
 |
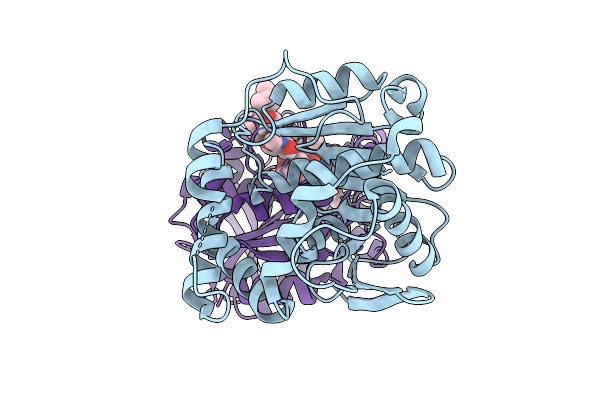
Crystal Structure Of Alpha/Beta-Hydrolase Macrolide Esterase Estx From Escherichia Coli (S102A Mutant) In Complex With Linearized Tylosin
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: A1E2O |
 |
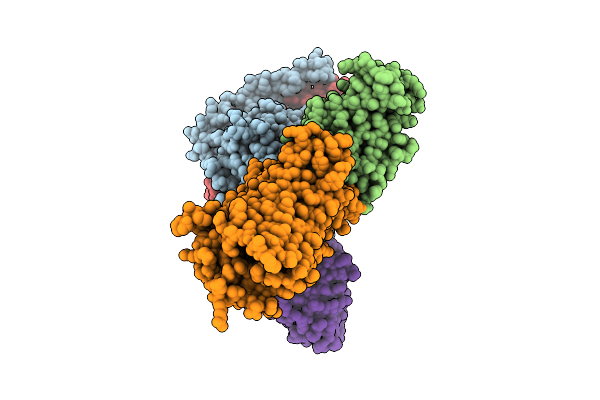
Crystal Structure Of Alpha/Beta-Hydrolase Macrolide Esterase Estx From Escherichia Coli (S102A Mutant) In Complex With Linearized Tylvalosin
Organism: Escherichia coli #1/h766
Method: X-RAY DIFFRACTION Resolution:1.99 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: A1DAR |
 |
Organism: Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM Ligands: A1E2G |
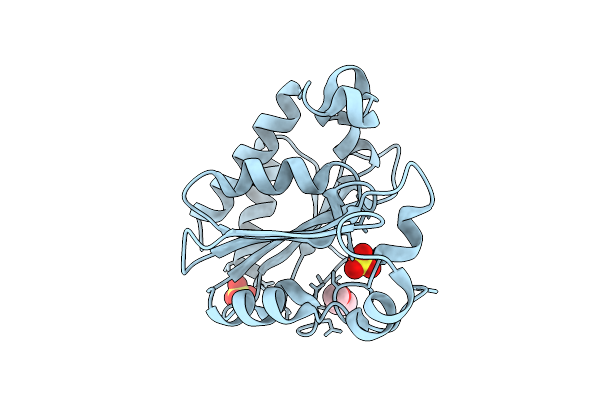 |
Crystal Structure Of Poly-Gamma-Glutamate Hydrolase From Bacillus Phage Pm1
Organism: Bacillus phage pm1
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: PEG, SO4 |
 |
Crystal Structure Of Poly-Gamma-Glutamate Hydrolase From Bacillus Phage Pm1 In Complex With A Zinc Ion
Organism: Bacillus phage pm1
Method: X-RAY DIFFRACTION Resolution:1.38 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: PEG, ZN |
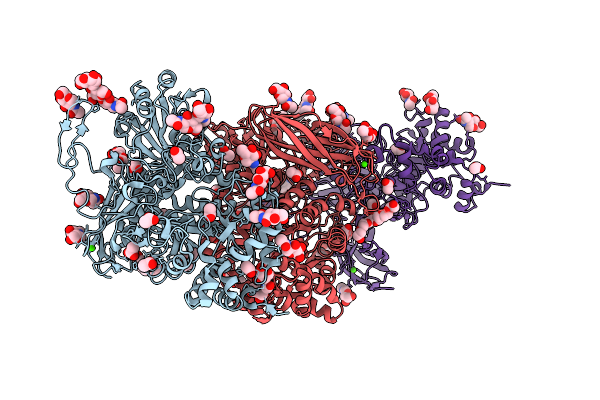 |
Crystal Structure Of Isoprimeverose-Producing Enzyme From Phaeoacremonium Minimum
Organism: Rattus rattus, Phaeoacremonium minimum
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAG, ACT, PG4, CA, PGE, P6G |
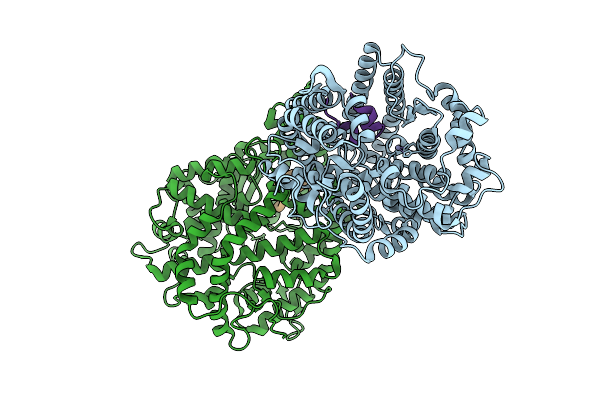 |
Organism: Homo sapiens, Synthetic plasmid
Method: X-RAY DIFFRACTION Resolution:2.02 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: ZN |
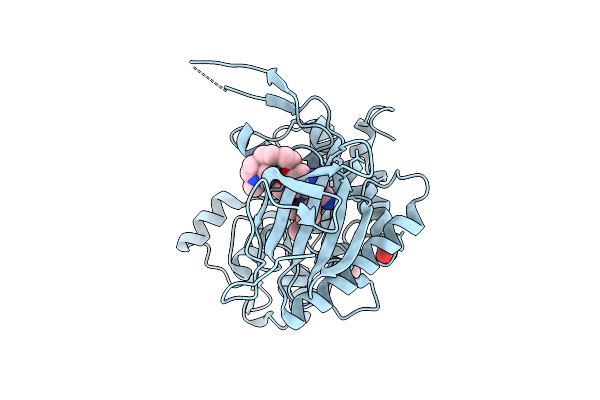 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.10 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: EDO, A1J3A, IOD |
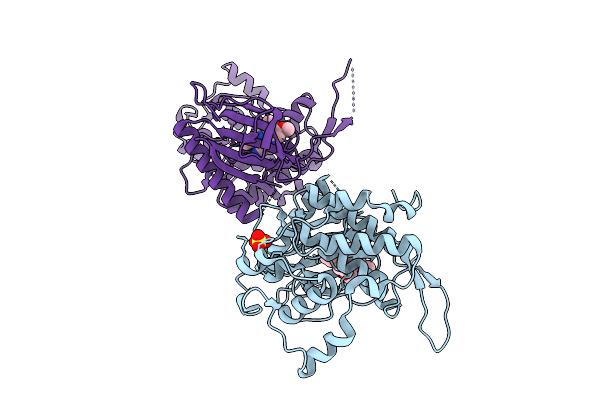 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: SO4, A1J3B |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.03 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN,HYDROLASE/INHIBITOR Ligands: A1A5Z, CL |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN,HYDROLASE/INHIBITOR Ligands: A1A54, CL |
 |
Sdacs: A Synergistic Strategy Using Both Crystallographic And Alphafold Structures To Improve Thermostability And Activity Of N-Acetylhexosaminidase
Organism: Cellvibrio mixtus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: TRS, GOL |

