Search Count: 1,17,624
All
Selected
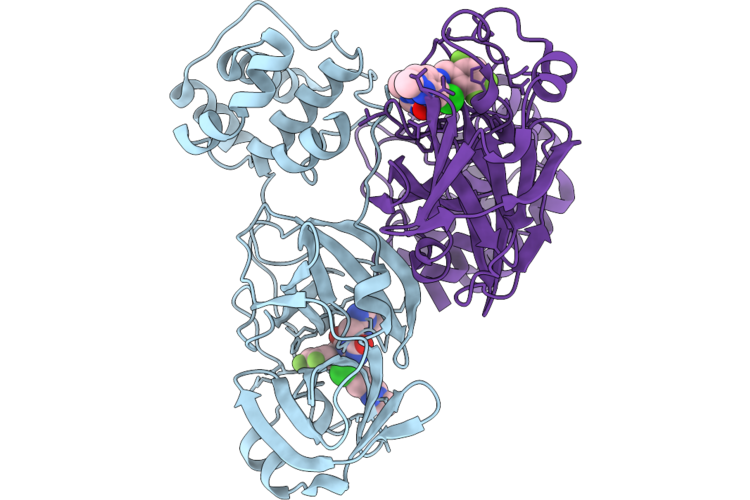 |
Room Temperature X-Ray Structure Of Sars Cov-2 Main Protease Intermediate Precursor With Ensitrelvir (Esv)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.05 Å Release Date: 2026-05-06 Classification: HYDROLASE/HYDROLASE INHIBITOR Ligands: 7YY |
 |
Organism: Bacillus thuringiensis
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: UDP, NOV |
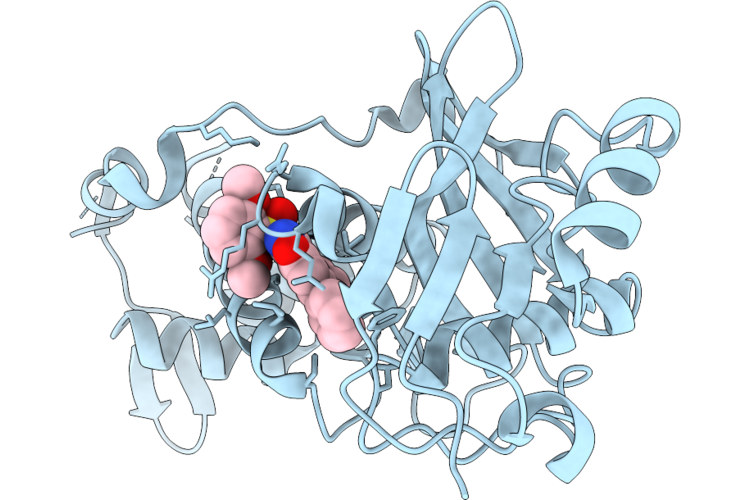 |
Crystal Structure Of Myst Histone Acetyltransferase Kat6A In Complex With Inhibitor Compound 9
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.29 Å Release Date: 2026-05-06 Classification: TRANSCRIPTION Ligands: A1MGC |
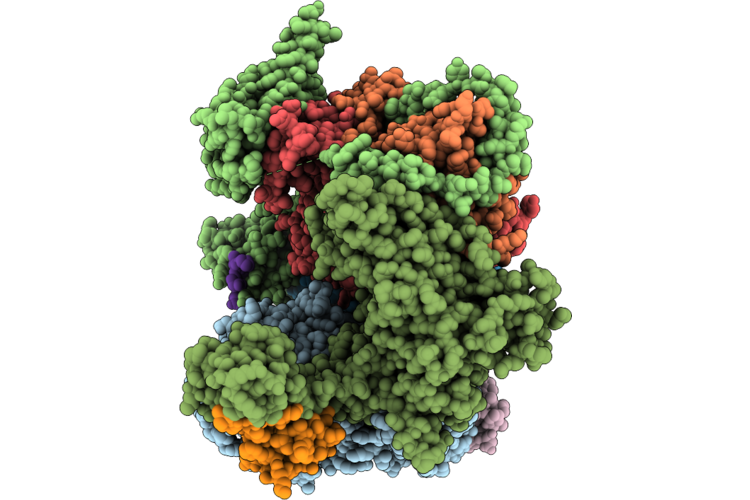 |
Cryo-Em Structure Of The Human Holo-Tfiih And Xpc Initial Encounter Complex
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN |
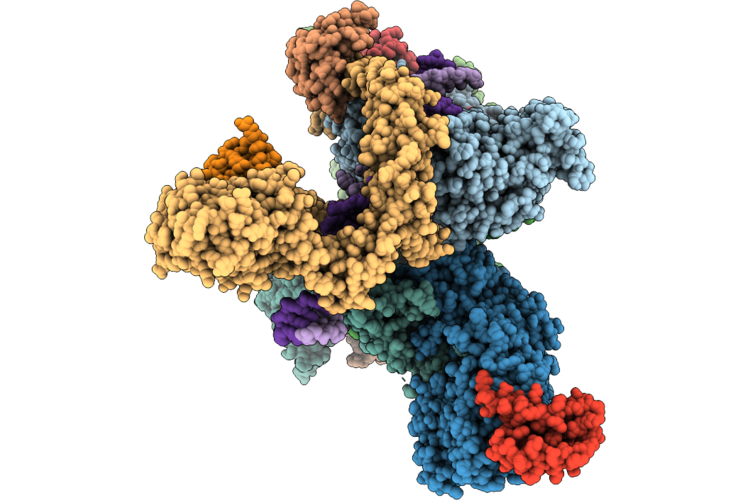 |
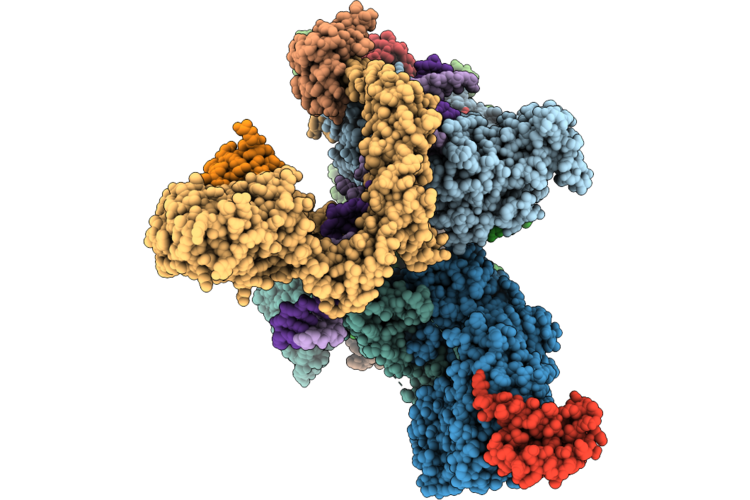
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Composite Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.32 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
 |
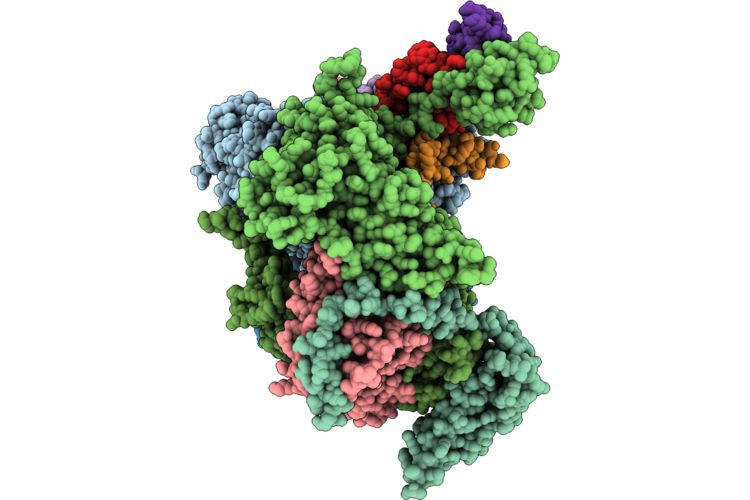
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc Complex Bound To Bulky Lesion-Mimic Dna (Consensus Map)
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.91 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN, CA |
 |
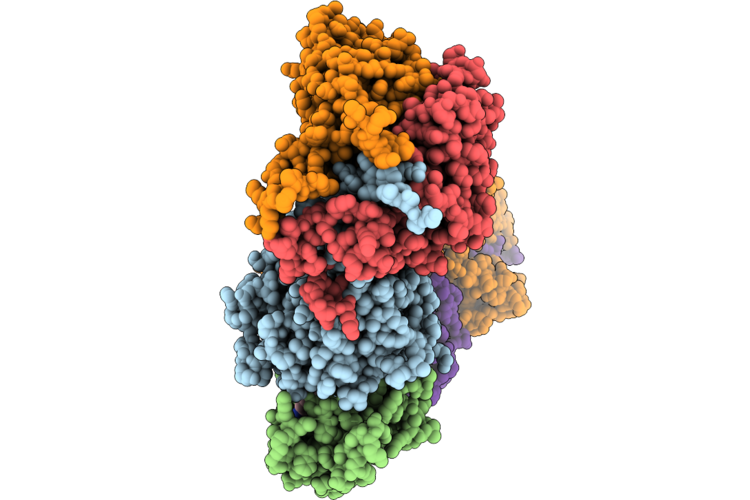
Cryo-Em Structure Of The Human Holo-Tfiih-Xpc-Xpa Complex Bound To Bulky Lesion-Mimic Dna
Organism: Homo sapiens, Synthetic construct
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-05-06 Classification: DNA BINDING PROTEIN Ligands: SF4, ZN |
 |
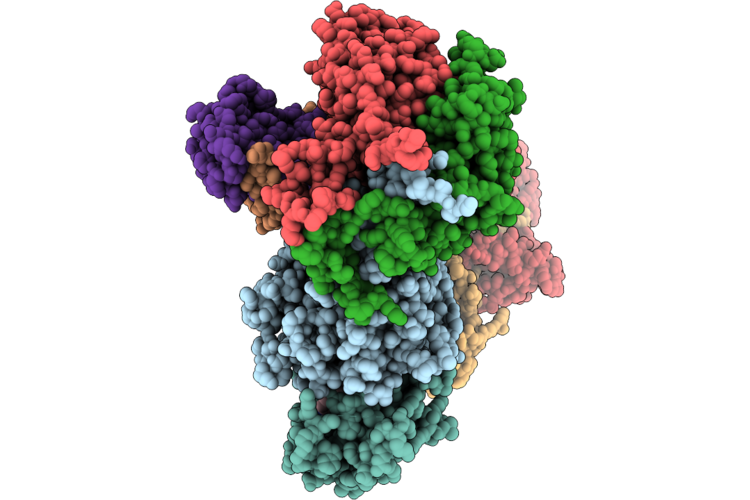
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2 Via Unc10088
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J20 |
 |
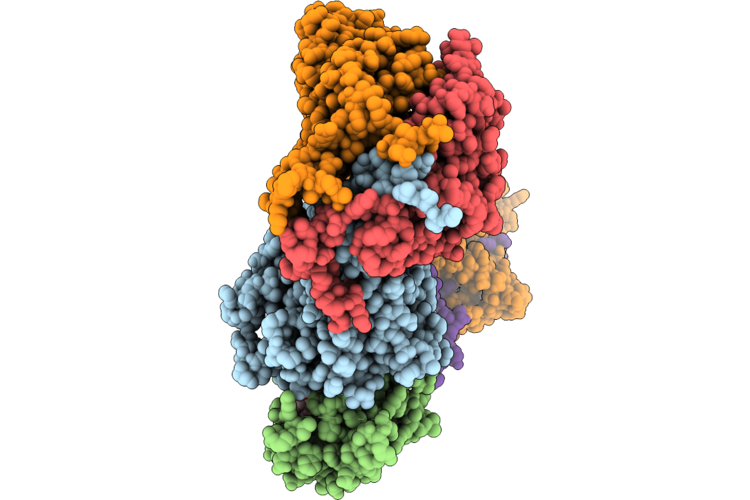
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2, Unc10088 And Bach1
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J20 |
 |
Cryo-Em Structure Of The Cul1-Rbx1-Skp1-Fbxo22 Scf Ubiquition Ligase In Complex With Nsd2 Via Unc10415667
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: LIGASE Ligands: A1J21 |
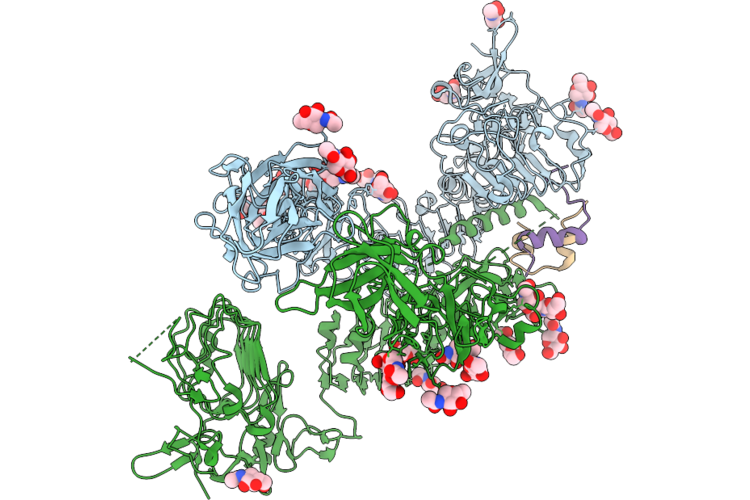 |
Drosophila Melanogaster Insulin Receptor Ectodomain In Complex With One Dilp2 Molecule
Organism: Drosophila melanogaster
Method: ELECTRON MICROSCOPY Resolution:3.70 Å Release Date: 2026-05-06 Classification: SIGNALING PROTEIN Ligands: NAG |
 |
Drosophila Melanogaster Insulin Receptor Ectodomain In Complex With Two Dilp2 Molecules
Organism: Drosophila melanogaster
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: SIGNALING PROTEIN Ligands: NAG |
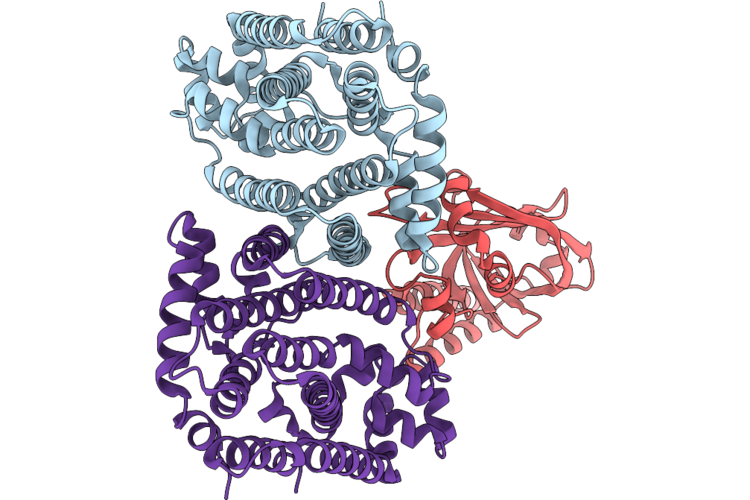 |
Organism: Oryza sativa subsp. japonica
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSFERASE |
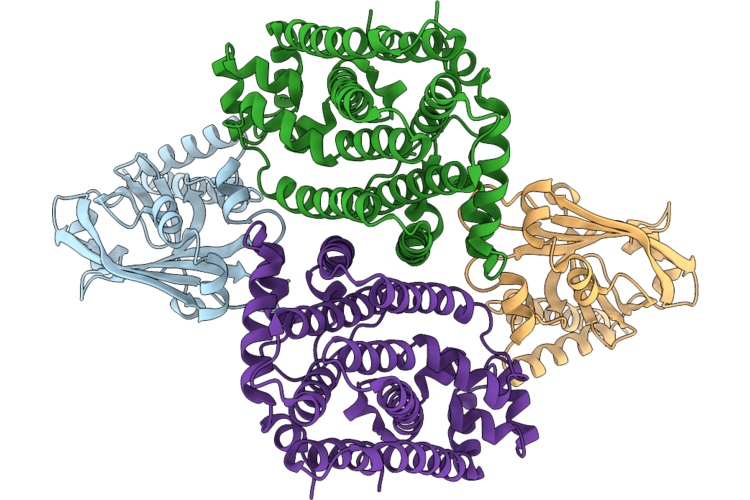 |
Organism: Oryza sativa japonica group
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSFERASE |
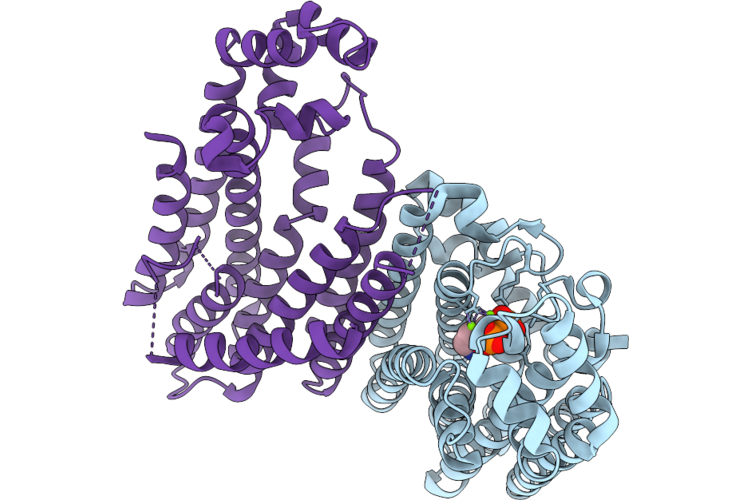 |
Organism: Sorghum bicolor
Method: X-RAY DIFFRACTION Resolution:2.43 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: A1EE9, MG |
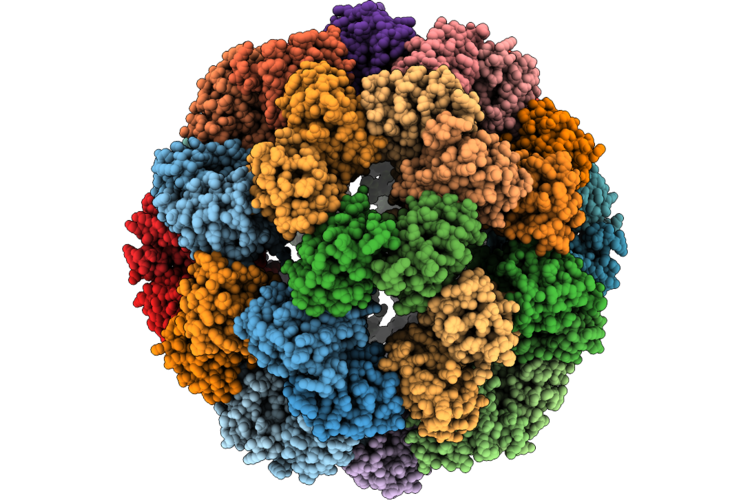 |
Organism: Listeria monocytogenes egd-e
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSCRIPTION |
 |
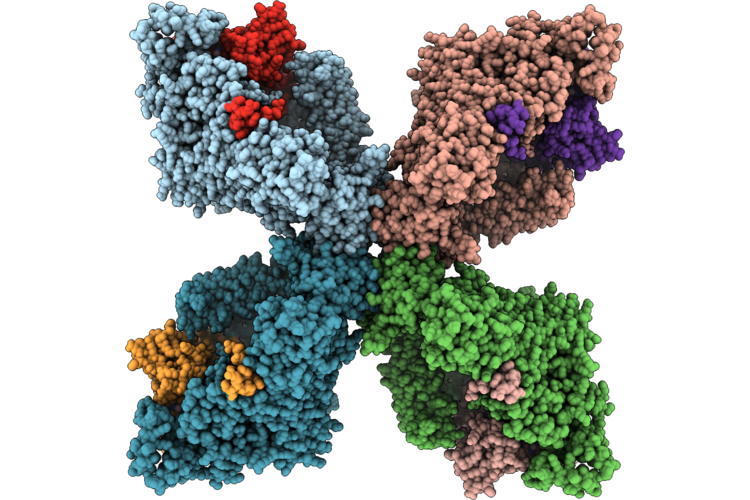
Organism: Listeria monocytogenes
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSCRIPTION |
 |
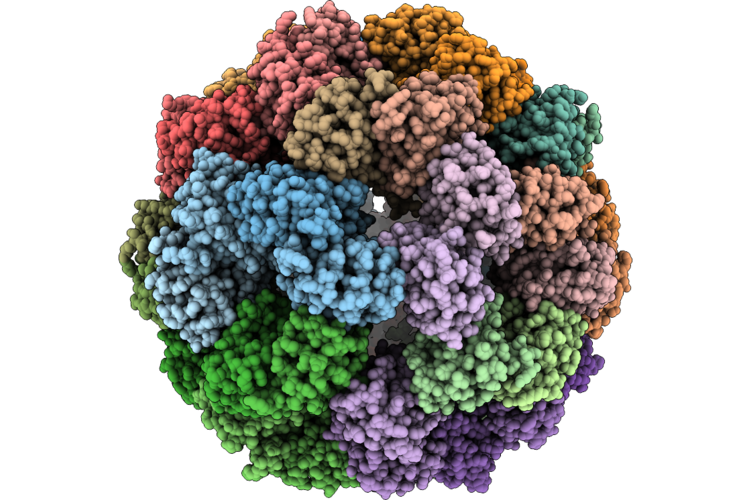
Organism: Escherichia coli, Escherichia phage phiv-1
Method: ELECTRON MICROSCOPY Resolution:2.33 Å Release Date: 2026-05-06 Classification: ANTIVIRAL PROTEIN Ligands: ATP, MG |
 |
Organism: Listeria monocytogenes
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSCRIPTION |
 |
Organism: Listeria monocytogenes
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: TRANSCRIPTION |

