Search Count: 6,347
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: MAN, FMT, BMA, NA |
 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:0.94 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: ACT, FMT, NA |
 |
Structural Characterization Of A N-Acetyl-D-Glucosamine-Binding Site In Human Rnase2
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: ACT, NAG, PG4, FMT, PEG, NA |
 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-22 Classification: PROTEIN BINDING Ligands: GLA, ACT, FMT, PEG, EDO, NA, CL |
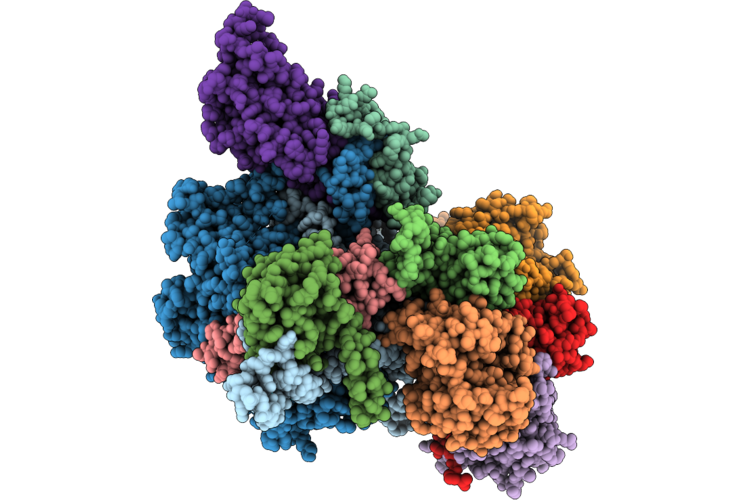 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.84 Å Release Date: 2026-04-22 Classification: RNA BINDING PROTEIN/RNA |
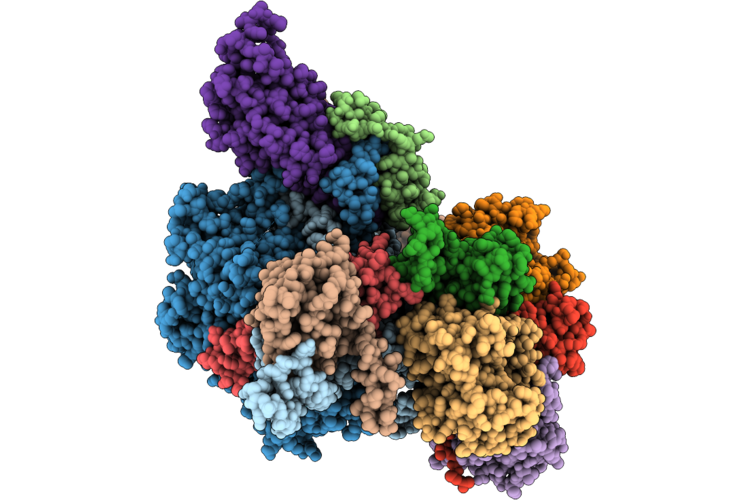 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.47 Å Release Date: 2026-04-22 Classification: RNA BINDING PROTEIN/RNA Ligands: ZN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.93 Å Release Date: 2026-04-22 Classification: RNA BINDING PROTEIN/RNA Ligands: ZN |
 |
Structural Characterization Of A N-Acetylmuramic Acid-Binding Site In Human Rnase2
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.01 Å Release Date: 2026-04-15 Classification: PROTEIN BINDING Ligands: MUB, AMU, ACT, FMT, NA, CL |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-04-08 Classification: RNA BINDING PROTEIN Ligands: ZN, MG |
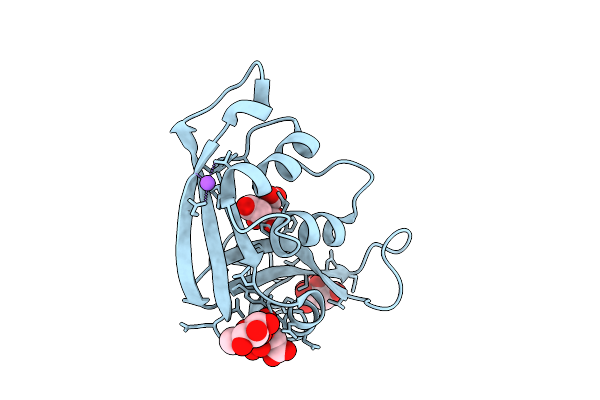 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.31 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: CIT, GLC, NA, CL |
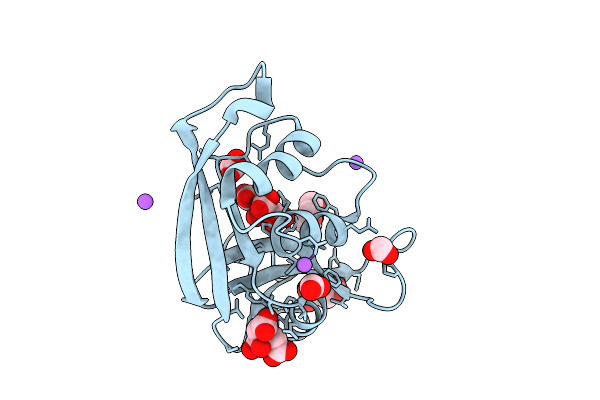 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.17 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: CIT, FMT, PEG, ACT, NA |
 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.04 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: B3P, FMT, PEG, NA |
 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.04 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: MLI, PEG, ACT, FMT |
 |
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.04 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: PEG, ACT, FMT, FUL, FUC |
 |
Structural Characterization Of Glucose- And Fucose-Binding Sites In Human Rnase2
Organism: Escherichia coli 'bl21-gold(de3)plyss ag'
Method: X-RAY DIFFRACTION Resolution:1.01 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: PEG, FUC, ACT, GLC, NA |
 |
Structural Characterization Of A N-Acetyl-Beta-Neuraminic Acid-Binding Site In Human Rnase2
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.01 Å Release Date: 2026-03-18 Classification: PROTEIN BINDING Ligands: SLB, B3P, ACT |
 |
Joint X-Ray/Neutron Structure Of Wild-Type Bacillus Halodurans Rnase H1 In The Apo-Form
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:2.48 Å, 1.90 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: SO4, DOD |
 |
Joint X-Ray/Neutron Structure Of D132N Bacillus Halodurans Rnase H1 In The Apo-Form
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION, NEUTRON DIFFRACTION Resolution:2.4 Å, 2.20 Å Release Date: 2026-03-18 Classification: HYDROLASE Ligands: SO4, DOD |
 |
Room-Temperature X-Ray Structure Of D132N Bacillus Halodurans Rnase H1 In Complex With Rna/Dna Duplex
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-03-18 Classification: HYDROLASE/RNA/DNA Ligands: MG, PO3 |
 |
Room-Temperature X-Ray Structure Of D132N Bacillus Halodurans Rnase H1 In Complex With Complementary Rna/Dna Duplex
Organism: Halalkalibacterium halodurans
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-03-18 Classification: HYDROLASE/RNA/DNA Ligands: MG |

