Search Count: 55,470
All
Selected
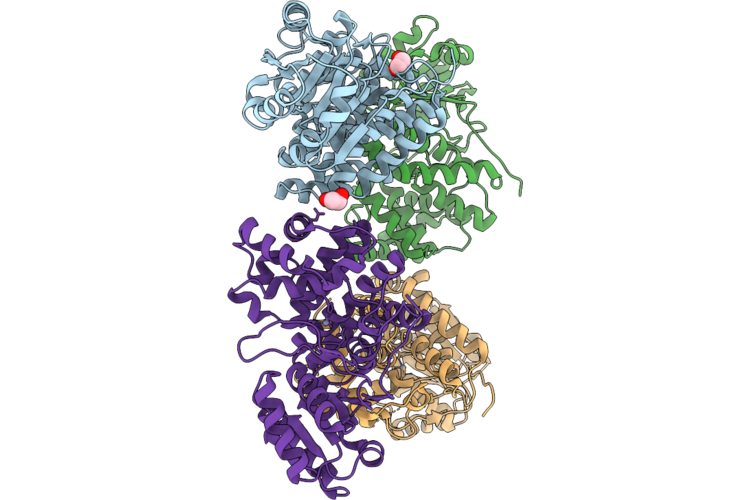 |
Organism: Streptococcus agalactiae
Method: X-RAY DIFFRACTION Resolution:3.10 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: EDO, ZN |
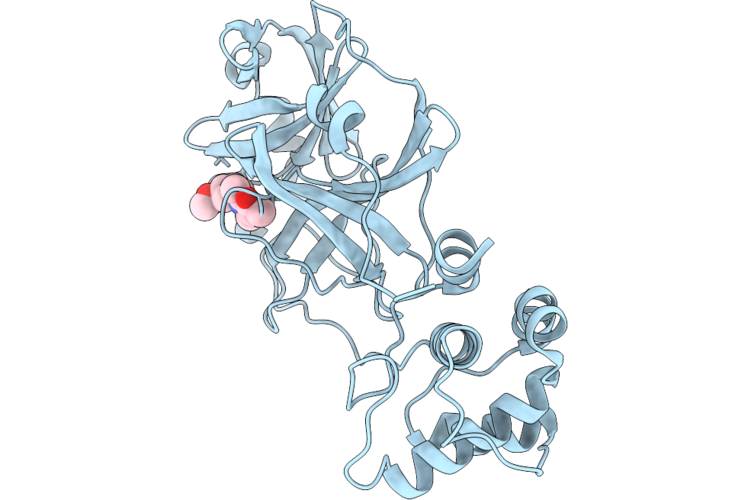 |
Crystal Structure Of Sars-Cov-2 Main Protease (Mpro) In Complex With Covalent Inhibitor A02
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2026-05-13 Classification: HYDROLASE Ligands: A1BGG |
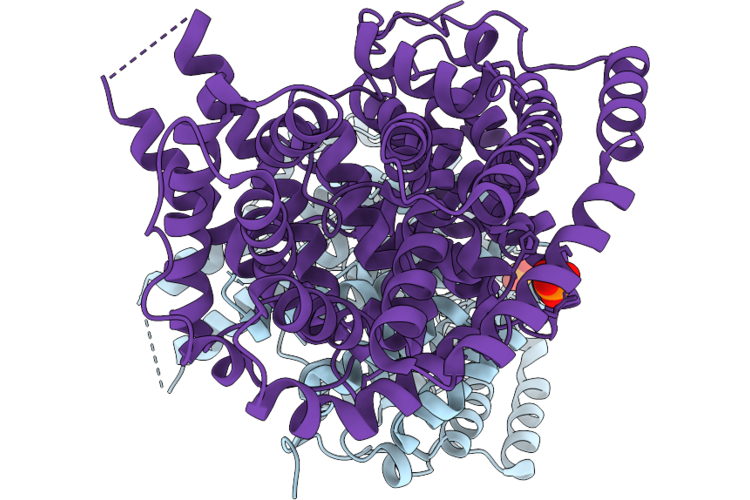 |
Organism: Phyla dulcis
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-05-13 Classification: LYASE Ligands: PO4 |
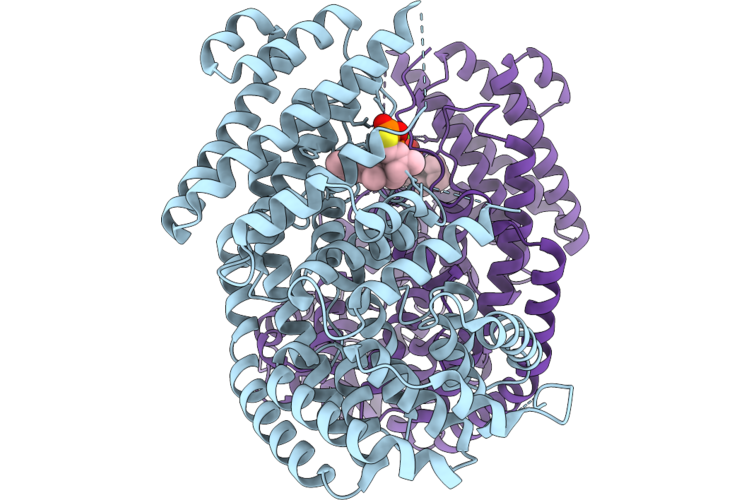 |
Organism: Phyla dulcis
Method: X-RAY DIFFRACTION Resolution:2.89 Å Release Date: 2026-05-13 Classification: LYASE Ligands: MG, FPS |
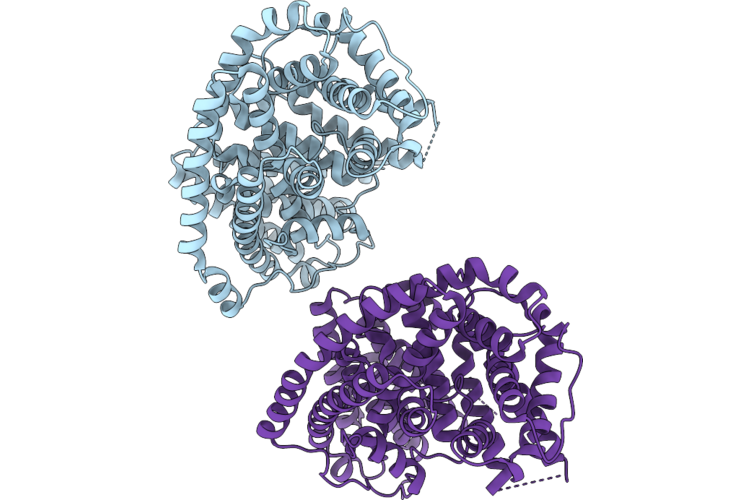 |
Crystal Structure Of Bicyclogermacrene Synthase Mutant I290V/I385C/V434C/L454C/V476W/L558I
Organism: Phyla dulcis
Method: X-RAY DIFFRACTION Resolution:2.30 Å Release Date: 2026-05-13 Classification: LYASE |
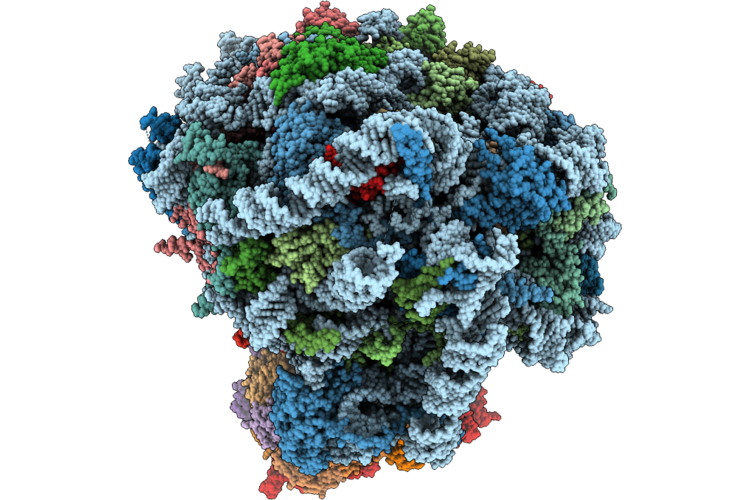 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
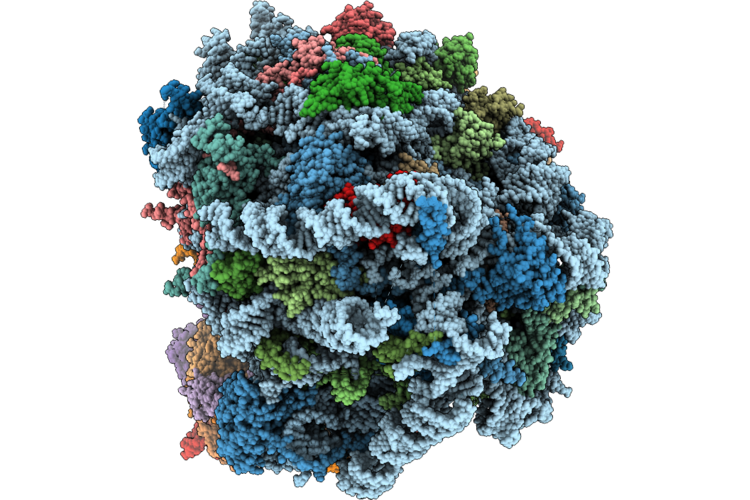 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
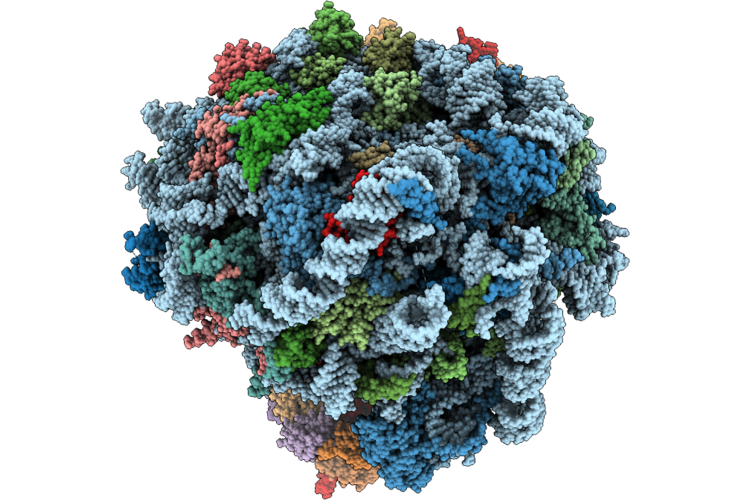 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: RIBOSOME Ligands: MG, ZN |
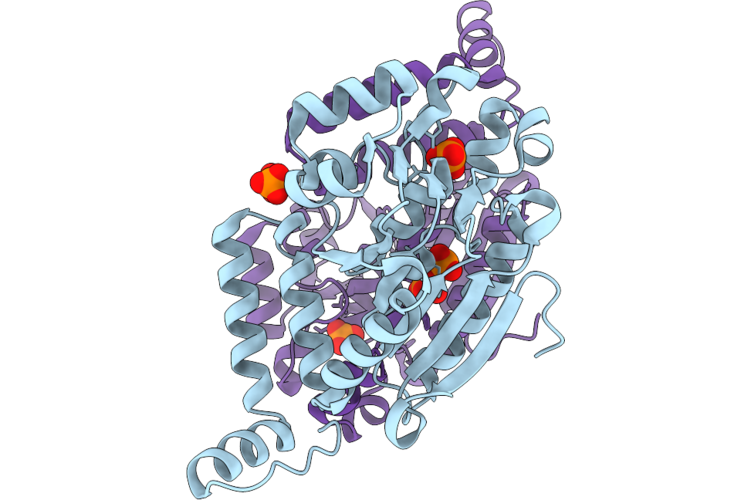 |
Crystal Structure Of Rattus Norvegicus Enoyl-Coa Hydratase In Unliganded Form
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-05-13 Classification: LYASE Ligands: PO4 |
 |
Crystal Structure Of Rattus Norvegicus Enoyl-Coa Hydratase In Complex With 3S-Hydroxybutanoyl-Coa
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:2.69 Å Release Date: 2026-05-13 Classification: LYASE Ligands: 3HC |
 |
Crystal Structure Of Rattus Norvegicus Enoyl-Coa Hydratase In Complex With 3S-Hydroxyhexanoyl-Coa
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-05-13 Classification: LYASE Ligands: 3H9 |
 |
Crystal Structure Of Rattus Norvegicus Enoyl-Coa Hydratase In Complex With 3S-Hydroxydecanoyl-Coa
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-05-13 Classification: LYASE Ligands: HSC |
 |
Crystal Structure Of Rattus Norvegicus Enoyl-Coa Hydratase In Complex With 3S Hydroxyhexanoyl-Pan And 3',5', Diphosphate Adenosine
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-05-13 Classification: LYASE Ligands: COA, A1JF6, A3P |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.00 Å Release Date: 2026-05-13 Classification: RIBOSOME Ligands: K, MG, C, NA, ZN, HYG |
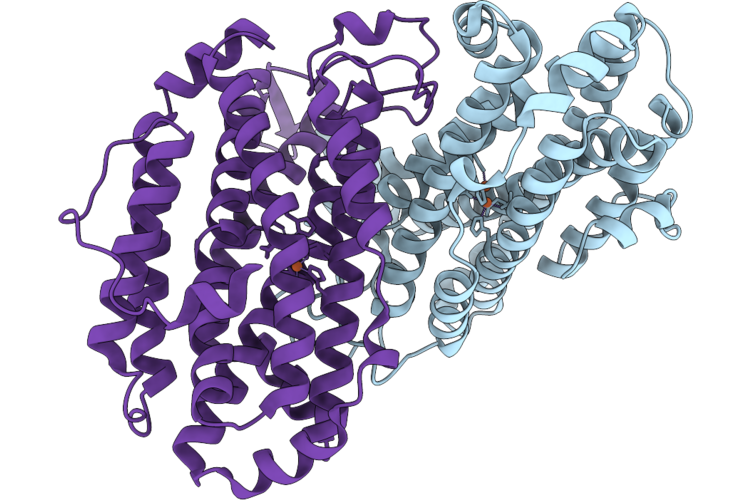 |
Xfel Structure Of Oxidised Ribonucleotide Reductase R2A Y122F Mutant From E. Coli
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE |
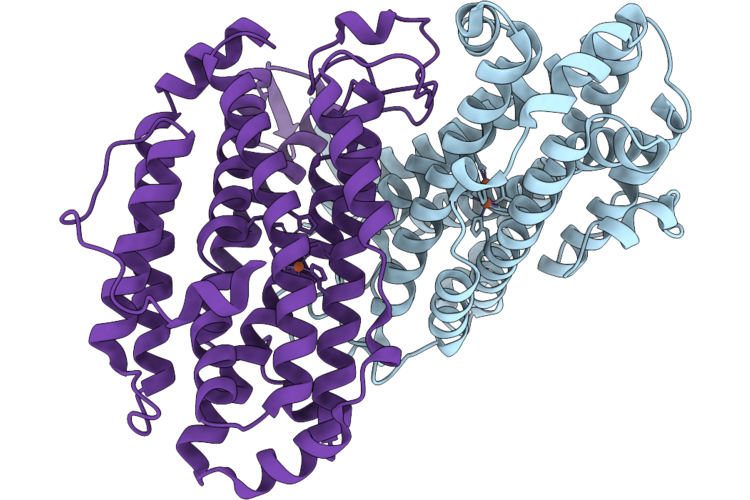 |
Xfel Structure Of Ribonucleotide Reductase R2A Y122F Mutant From E. Coli,Reduced Form
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE2 |
 |
Coproheme Decarboxylase H117A Mutant From Listeria Monocytogenes In Complex With Iron Coproporphyrin Iii
Organism: Listeria monocytogenes egd-e
Method: X-RAY DIFFRACTION Resolution:2.19 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FEC |
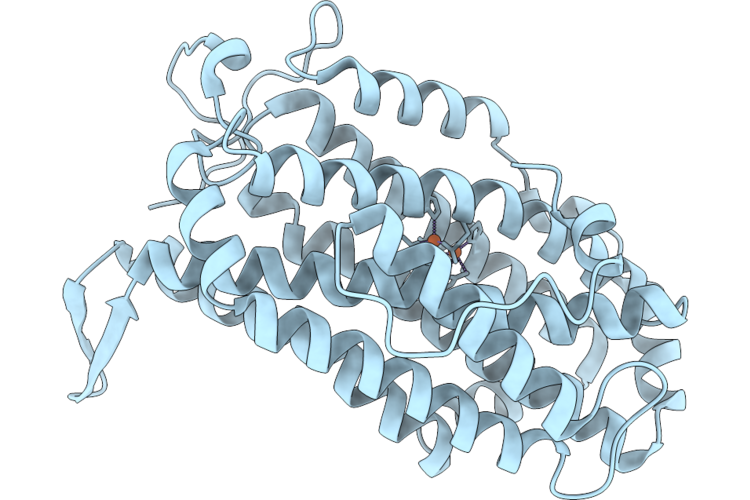 |
Xfel Structure Of Oxidised Ribonucleotide Reductase R2A Y122F Mutant From E. Coli, Hexagonal P6122 Form
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE |
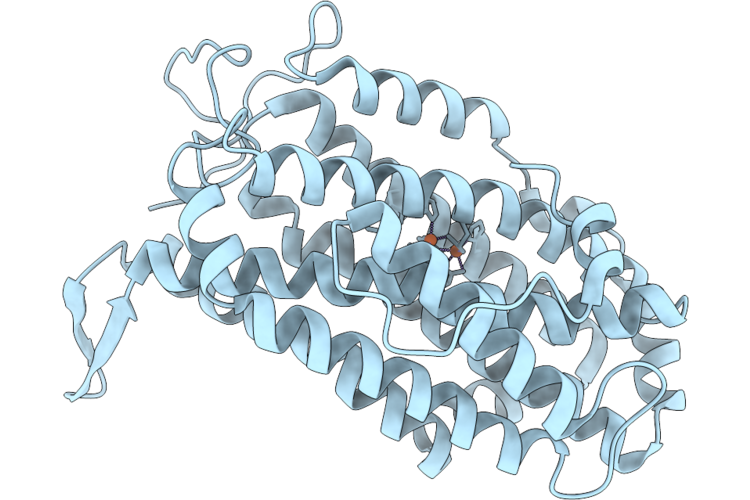 |
Xfel Structure Of Ribonucleotide Reductase R2A Y122F Mutant From E. Coli,Reduced Form, Hexagonal P6122
Organism: Escherichia coli
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-05-13 Classification: OXIDOREDUCTASE Ligands: FE2 |

