Search Count: 1,923
All
Selected
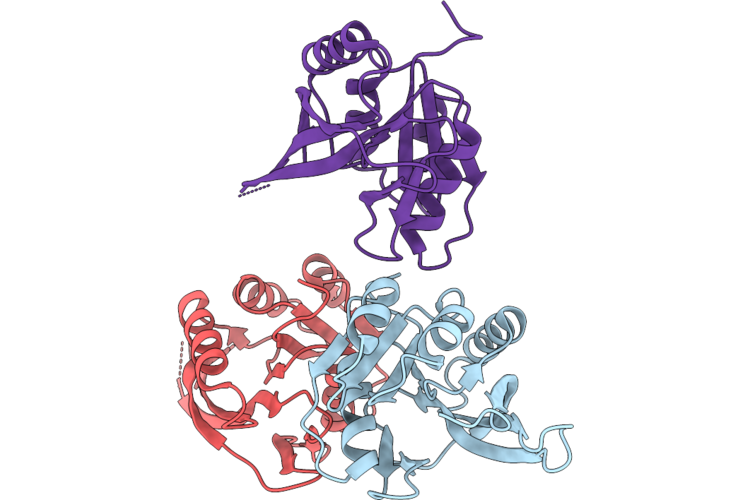 |
Organism: Bacillus cereus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-22 Classification: TRANSFERASE |
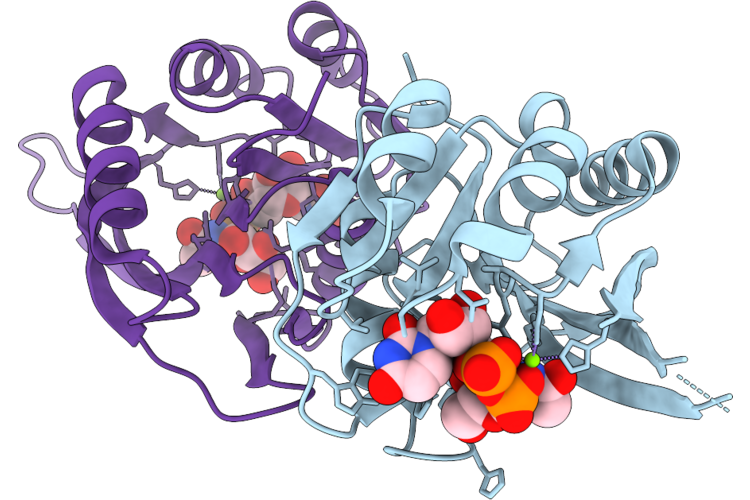 |
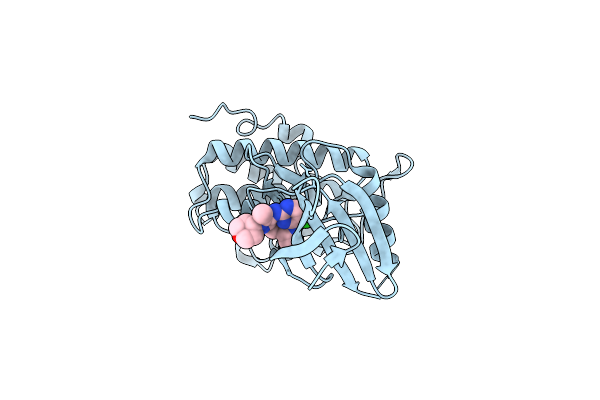
Crystal Structure Of Bacillus Cereus Gmar In Complex With Udp-Glcnac And Mg2+
Organism: Bacillus cereus
Method: X-RAY DIFFRACTION Resolution:2.42 Å Release Date: 2026-04-22 Classification: TRANSFERASE Ligands: UD1, MG |
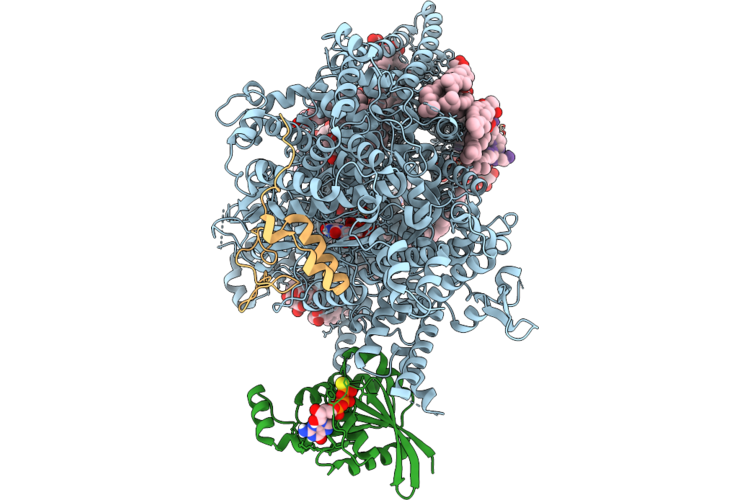 |
Structure Of Beta-1,3-Glucan Synthase In Complex With Caspofungin, Rho1 And Long Glucan
Organism: Saccharomyces cerevisiae, Synthetic construct
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP, GSP, A1CHR |
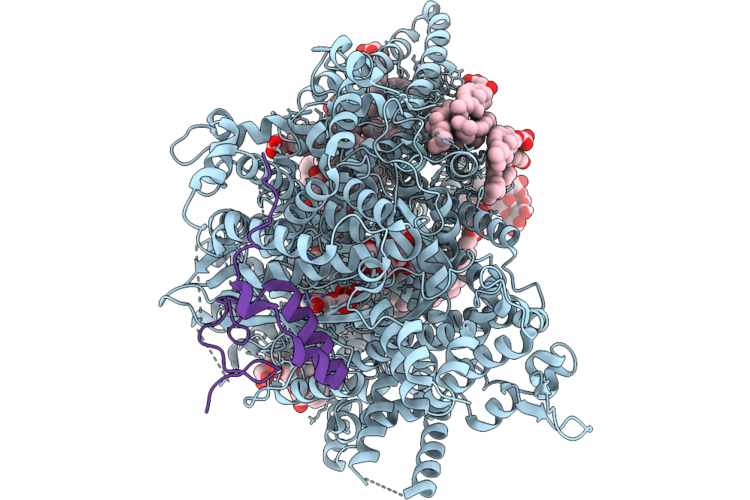 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) In Complex With Short Glucan
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP |
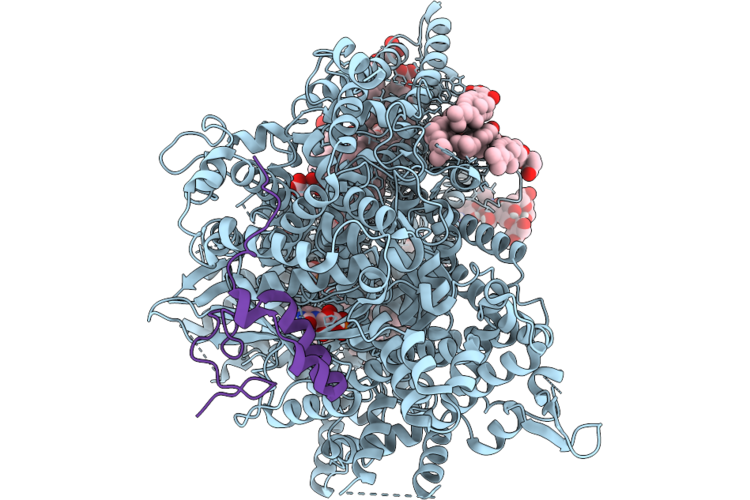 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Relevant Ground State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: NAG, Y01, 3PE, UDP |
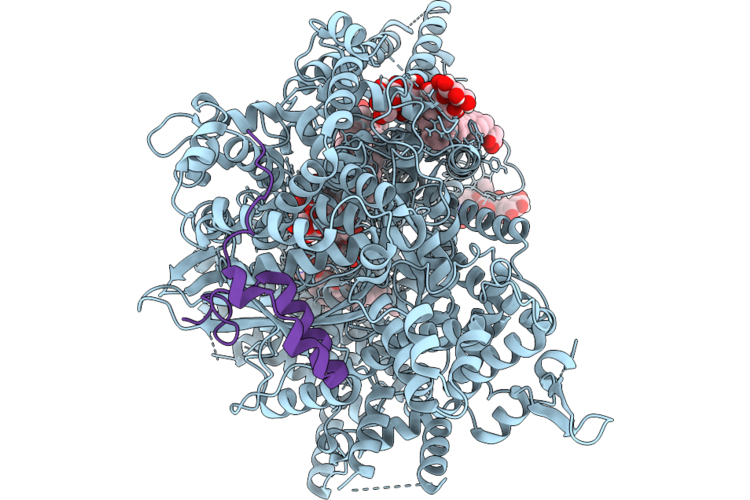 |
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Less Relevant L2 State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: Y01, 3PE, LMN |
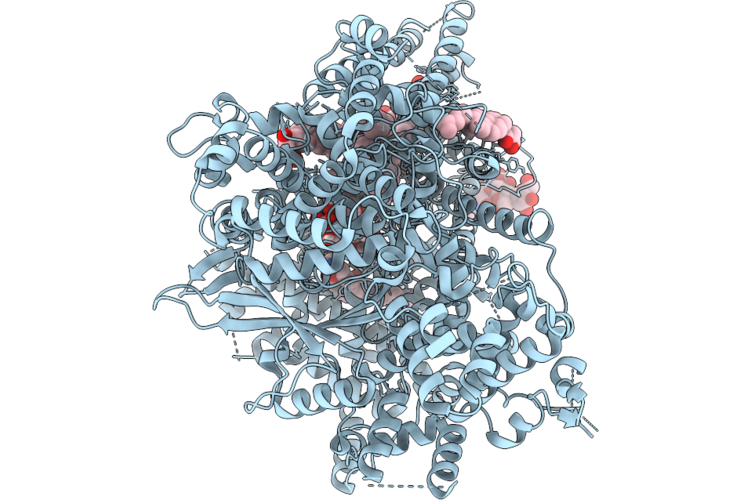 |
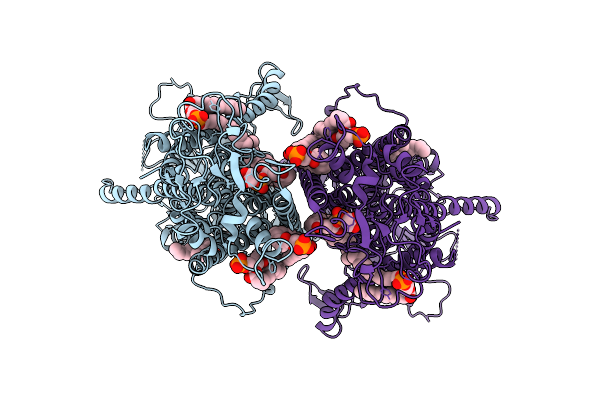
Structure Of Beta-1,3-Glucan Synthase From Saccharomyces Cerevisiae (Scfks1) At The Catalytically Less Relevant L1 State
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: Y01, 3PE, LMN |
 |
Organism: Saccharomyces cerevisiae
Method: ELECTRON MICROSCOPY Resolution:3.50 Å Release Date: 2026-04-22 Classification: MEMBRANE PROTEIN Ligands: Y01 |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: LIPID TRANSPORT Ligands: PGW |
 |
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 1 ((11R)-8-Chloro-3,11-Dimethyl-2-(Oxan-4-Yl)-2,4,10,11,12,13-Hexahydro-9,5-(Azeno)Pyrazolo[3,4-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.03 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5F |
 |
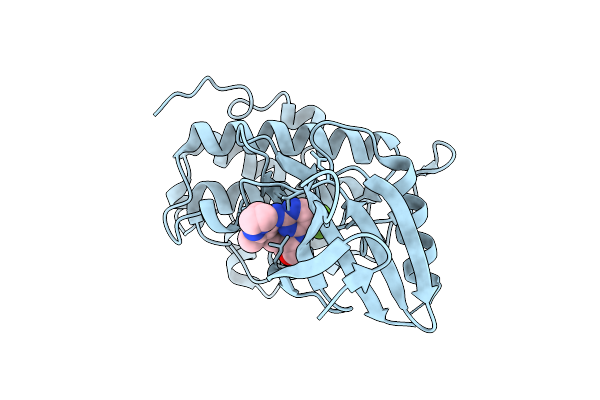
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 7 ((10As,13As)-3-Cyclobutyl-1-Methyl-8-(Trifluoromethyl)-3,4,10A,11,13A,14-Hexahydro-10H,13H-9,5-(Azeno)Furo[3,4-K]Pyrazolo[4,3-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5G |
 |
Structure Of Chk1 10-Pt. Mutant Complex With Macrocyclic Lrrk2 Inhibitor Compound 12 ((10As,13As)-3-Cyclopropyl-1-Methyl-8-(Trifluoromethyl)-3,4,10A,11,13A,14-Hexahydro-10H,13H-9,5-(Azeno)Furo[3,4-K]Pyrazolo[4,3-B][1,4,6,10]Oxatriazacyclotridecine)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-04-08 Classification: TRANSFERASE/INHIBITOR Ligands: A1C5H |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.70 Å Release Date: 2026-04-01 Classification: CELL CYCLE Ligands: A1MBI |
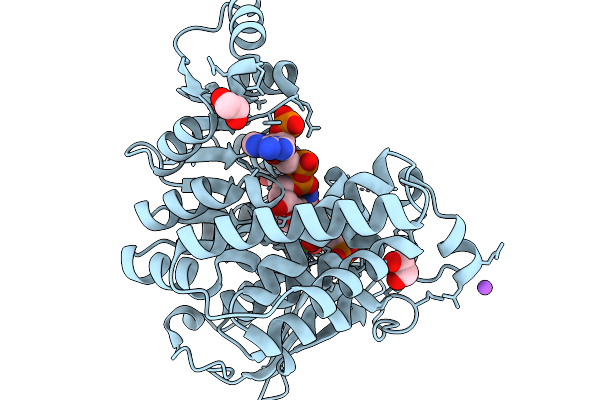 |
1-Deoxy-D-Xylulose 5-Phosphate Reductoisomerase (Ispc) From Acinetobacter Baumannii In Complex With A Fosmidomycin Analog, Nadph, And A Magnesium Ion
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: EDO, NDP, A1CFA, MG, CL, NA |
 |
1-Deoxy-D-Xylulose 5-Phosphate Reductoisomerase (Ispc) From Acinetobacter Baumannii In Complex With A Fosmidomycin Analog, Nadph, And A Magnesium Ion
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: NDP, A1CFB, EDO, MG, CL |
 |
1-Deoxy-D-Xylulose 5-Phosphate Reductoisomerase (Ispc) From Acinetobacter Baumannii In Complex With A Fosmidomycin Analog, Nadph, And A Magnesium Ion
Organism: Acinetobacter baumannii
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-03-25 Classification: OXIDOREDUCTASE Ligands: NDP, A1CE6, EDO, MG |
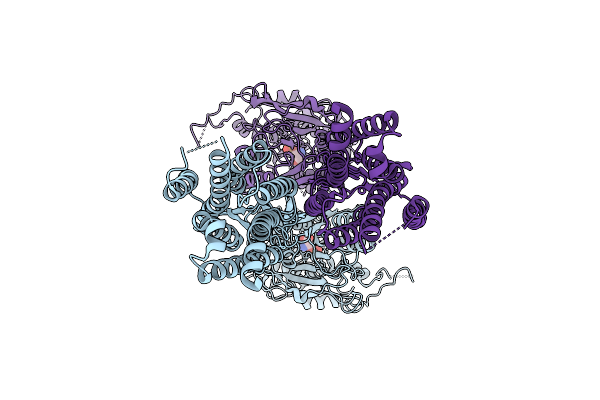 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.20 Å Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: SEP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.82 Å Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.39 Å Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: SEP |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:3.51 Å Release Date: 2026-03-18 Classification: MEMBRANE PROTEIN Ligands: SEP |

