Search Count: 1,356
All
Selected
 |
Organism: Staphylococcus aureus
Method: SOLUTION NMR Release Date: 2026-04-15 Classification: PROTEIN BINDING |
 |
Organism: Staphylococcaceae
Method: SOLUTION NMR Release Date: 2026-04-15 Classification: PROTEIN BINDING |
 |
Organism: Staphylococcaceae
Method: SOLUTION NMR Release Date: 2026-04-15 Classification: PROTEIN BINDING |
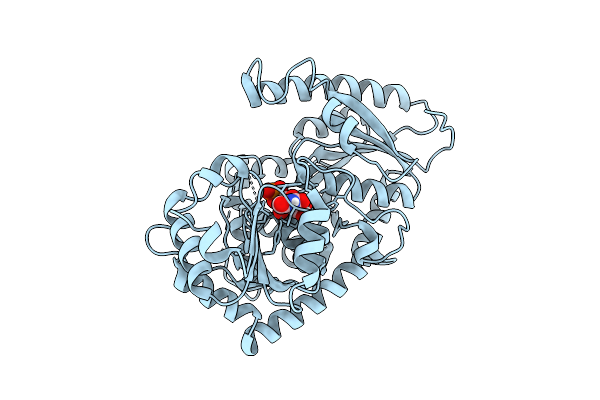 |
Crystal Structure Of The Glycosyltransferase Qsfuct From Quillaja Saponaria
Organism: Quillaja saponaria
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: UDP |
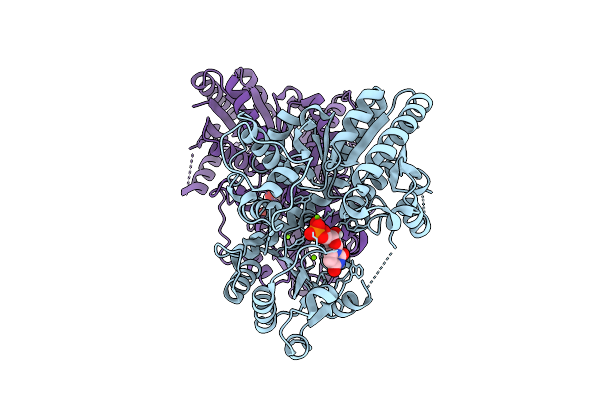 |
Crystal Structure Of The Glycosyltransferase Svfuct From Saponaria Vaccaria
Organism: Gypsophila vaccaria
Method: X-RAY DIFFRACTION Resolution:1.74 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: UDP, MG |
 |
Organism: Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.67 Å Release Date: 2026-03-11 Classification: DE NOVO PROTEIN Ligands: LFI |
 |
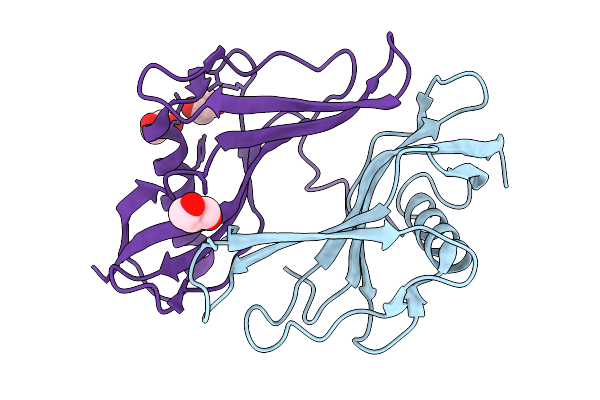
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2026-03-04 Classification: PROTEIN BINDING Ligands: PGE, FLC |
 |
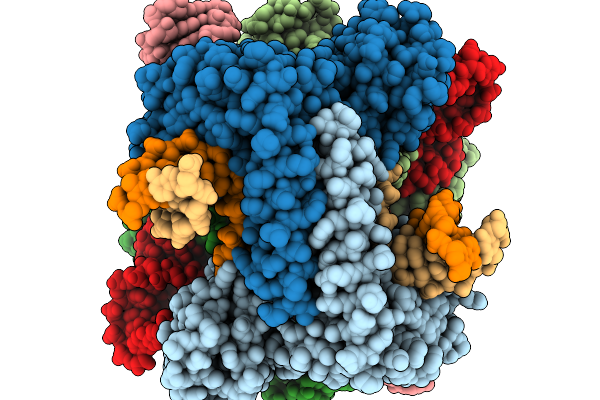
Organism: Staphylococcus aureus
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-03-04 Classification: PROTEIN BINDING Ligands: GOL, EDO |
 |
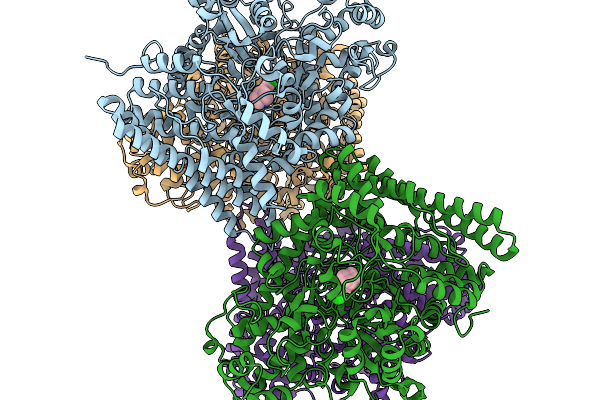
Organism: Citrobacter rodentium icc168
Method: ELECTRON MICROSCOPY Resolution:2.81 Å Release Date: 2026-02-18 Classification: RECOMBINATION Ligands: MG |
 |
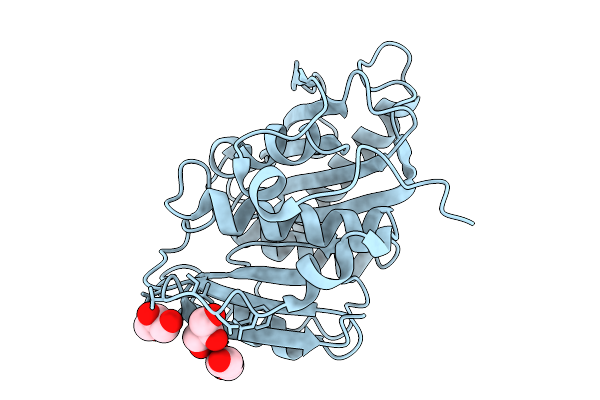
Organism: Oleidesulfovibrio alaskensis
Method: X-RAY DIFFRACTION Resolution:1.34 Å Release Date: 2026-02-11 Classification: HYDROLASE Ligands: A1JCI |
 |
Organism: Piscinibacter sakaiensis
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: GOL |
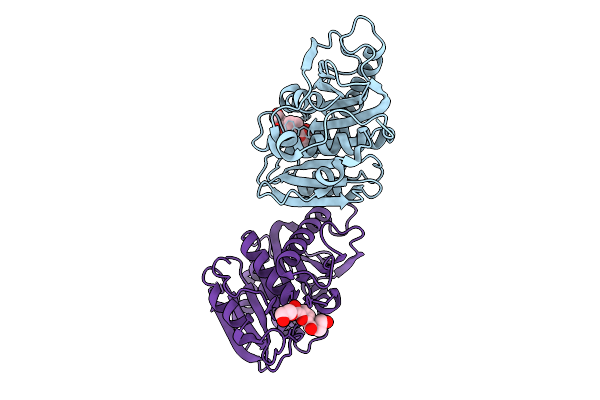 |
Organism: Piscinibacter sakaiensis
Method: X-RAY DIFFRACTION Resolution:2.49 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: A1L8P |
 |
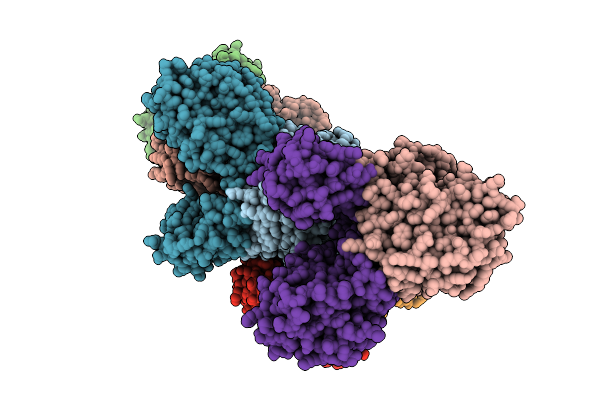
Organism: Borreliella burgdorferi
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: MN, FE, AMP |
 |
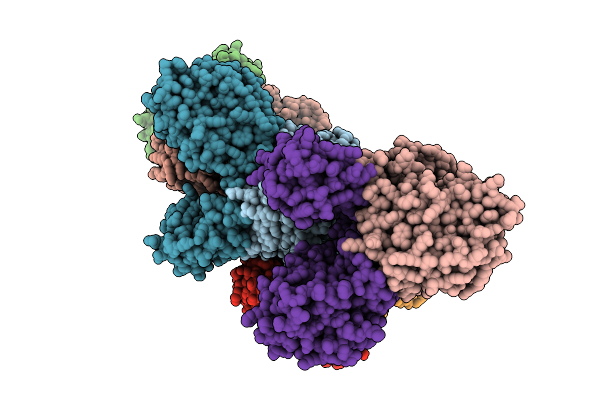
Organism: Borreliella burgdorferi
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: MN, FE, AMP |
 |
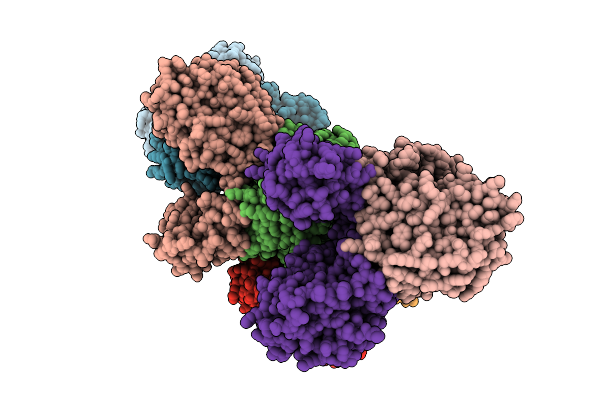
Organism: Borreliella burgdorferi
Method: X-RAY DIFFRACTION Resolution:3.30 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: MN, FE, AMP |
 |
Organism: Borreliella burgdorferi
Method: X-RAY DIFFRACTION Resolution:3.00 Å Release Date: 2026-01-21 Classification: HYDROLASE Ligands: MN, FE, A1JDM |
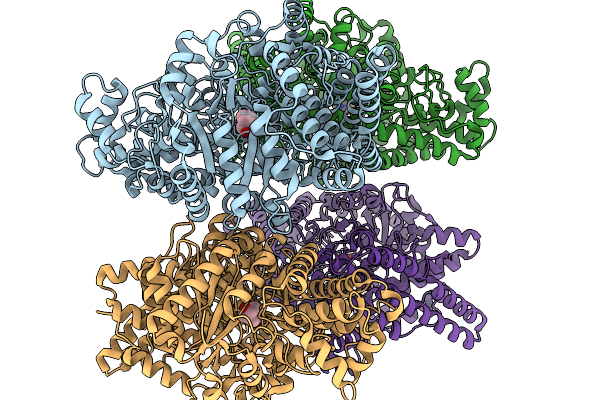 |
Organism: Oleidesulfovibrio alaskensis
Method: X-RAY DIFFRACTION Resolution:1.84 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1IXC |
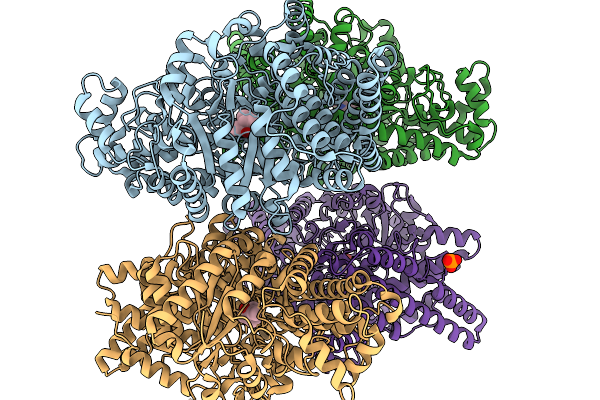 |
Organism: Oleidesulfovibrio alaskensis
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-01-14 Classification: OXIDOREDUCTASE Ligands: A1IXB, PO4 |
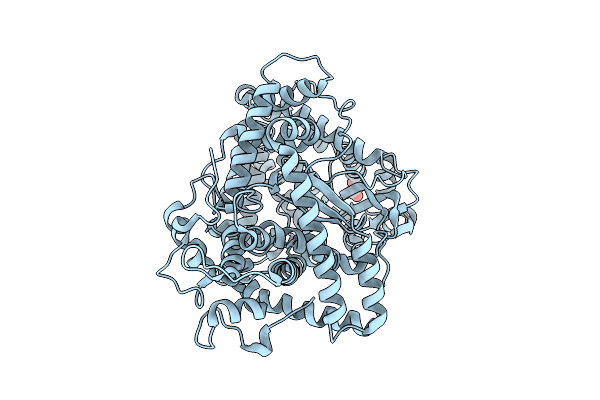 |
Organism: Porphyromonas gingivalis
Method: X-RAY DIFFRACTION Resolution:1.76 Å Release Date: 2025-12-17 Classification: HYDROLASE Ligands: EDO, ZN |
 |
Cryo Em Structure Of Rc-Dlh Complex Model Ii From Gemmatimonas Groenlandica
Organism: Gemmatimonas groenlandica
Method: ELECTRON MICROSCOPY Release Date: 2025-12-03 Classification: PHOTOSYNTHESIS Ligands: BCL, BPH, LMT, MQ8, FE, CD4, CRT, PEX, HEC, V7N |

