Search Count: 22,889
All
Selected
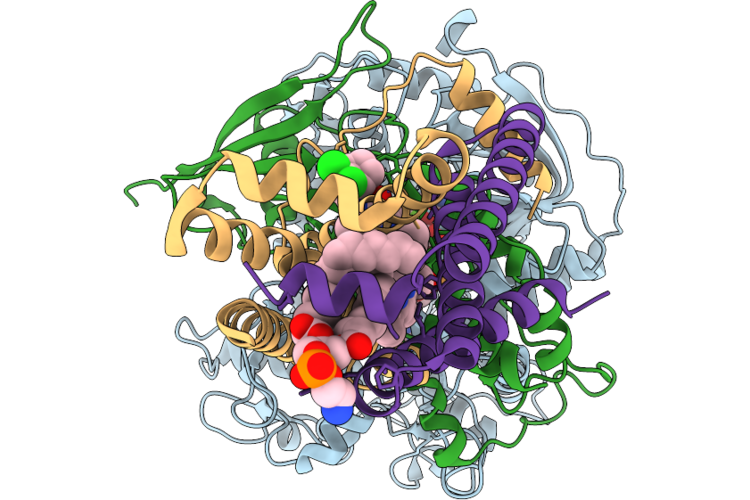 |
The Cryo-Em Structure Of Human Succinate Dehydrogenase In Complex With Benzovindiflupyr
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FAD, FES, SF4, F3S, A1EGM, HEM, PEV |
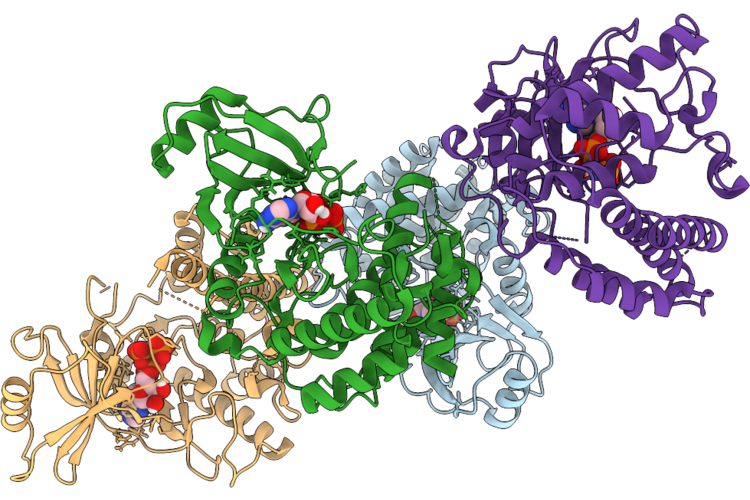 |
Caldalkalibacillus Thermarum Acyl-Coa Dehydrogenase Member 10 Bound To Anp And Magnesium
Organism: Caldalkalibacillus thermarum
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-05-06 Classification: TRANSFERASE Ligands: ANP |
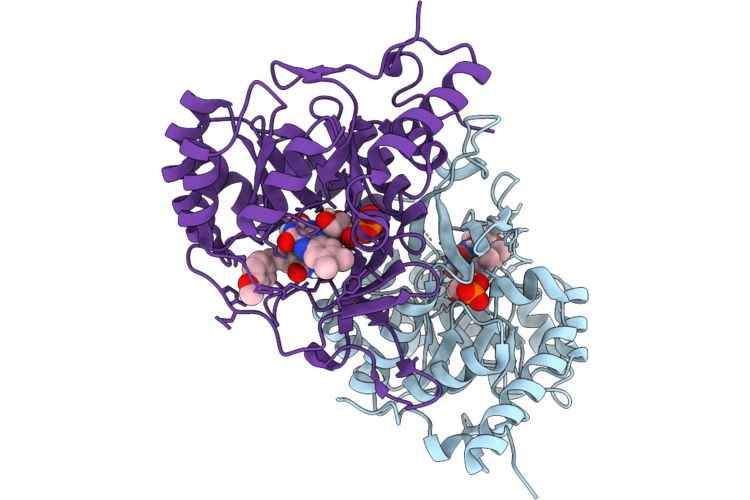 |
Crystal Structure Of Dihydroorotate Dehydrogenase From Leishmania Brasiliensis In Complex With 5-[(E)-3-(P-Methoxyphenyl)-2-Propenylidene]-2,4,6(1H,3H,5H)-Pyrimidinetrione
Organism: Leishmania braziliensis
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FMN, 5TI |
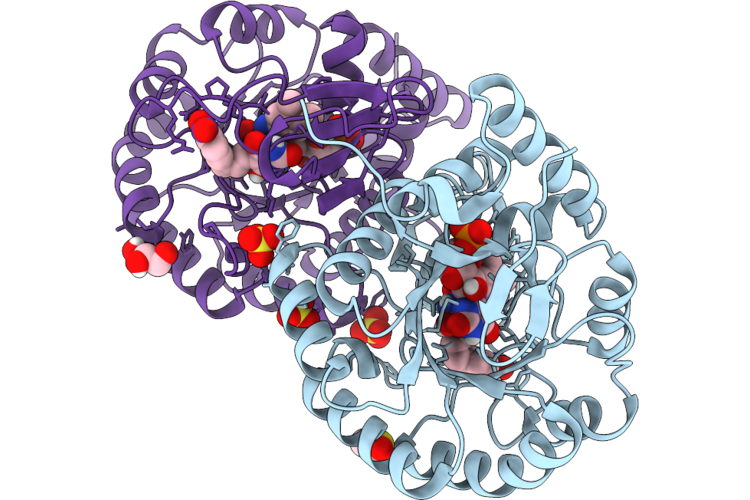 |
Crystal Structure Of Dihydroorotate Dehydrogenase From Leishmania Brasiliensis In Complex With (E)-5-(3-(4-Nitrophenyl)Allylidene)Pyrimidine-2,4,6(1H,3H,5H)-Trione
Organism: Leishmania braziliensis
Method: X-RAY DIFFRACTION Resolution:1.97 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: FMN, DMS, A1CAU, SO4, GOL |
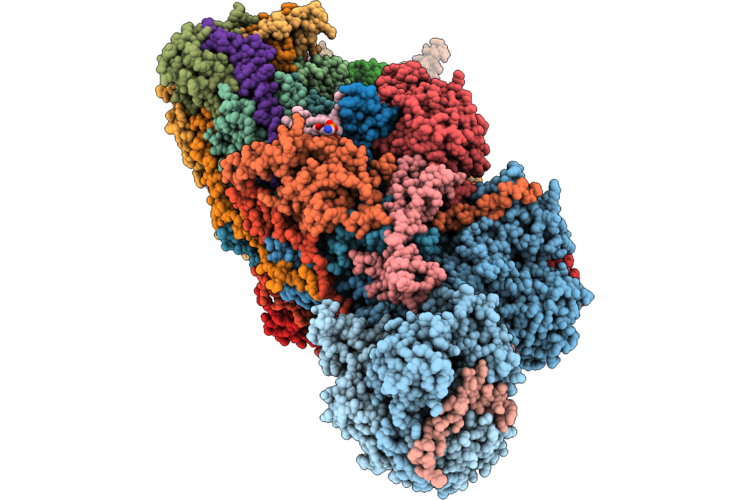 |
Organism: Ovis aries
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: 3PE, PC1, DGT, MG, MYR, ZMP, SF4, FMN, NAI, FES, K, A1JBT, ZN, NDP |
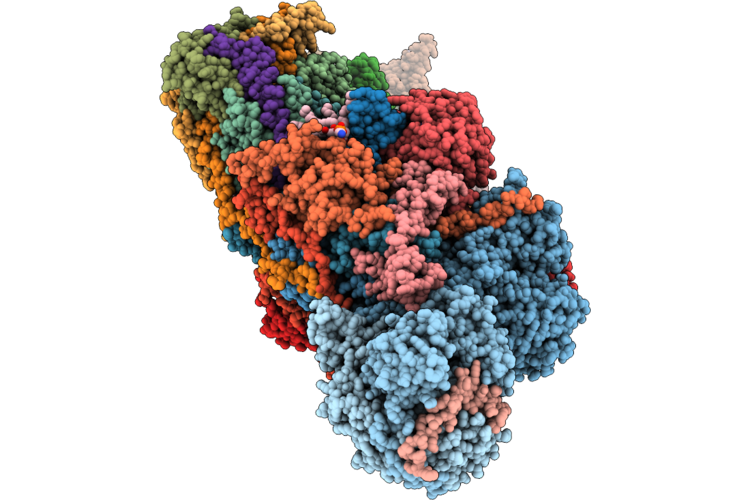 |
Organism: Ovis aries
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: 3PE, PC1, A1JBT, SF4, DGT, MG, MYR, ZMP, FMN, NAI, FES, K, ZN, NDP |
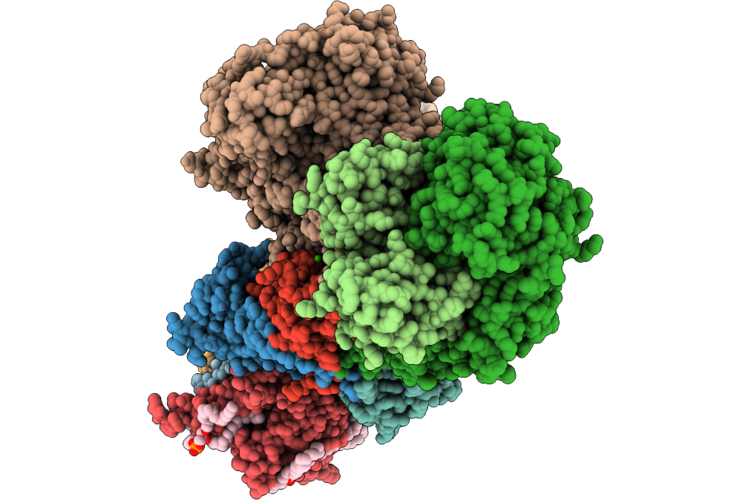 |
Organism: Escherichia coli bw25113, Escherichia coli bl21(de3)
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: FES, CA, FMN, SF4, 3PE, LFA, CDL, TRD, UQ8 |
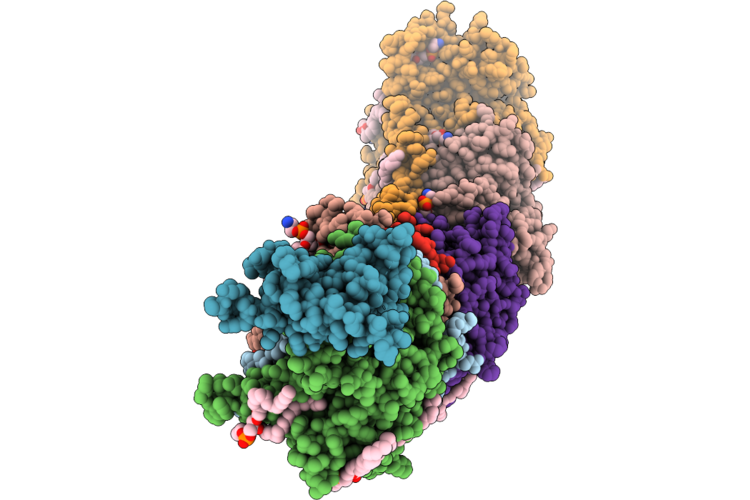 |
Organism: Escherichia coli bw25113
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: 3PE, LFA, CDL, TRD, UQ8 |
 |
Organism: Escherichia coli bw25113
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: FES, CA, FMN, SF4 |
 |
Organism: Escherichia coli bl21(de3), Escherichia coli bw25113
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: FES, SF4, FMN, CA, 3PE, 7PH, UQ8 |
 |
Local Refinement Of E. Coli Complex I D79N Nuoa Mutant Hydrophilic Domain In Lmng
Organism: Escherichia coli bw25113
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: SF4, FES, CA, FMN |
 |
Local Refinement Of E. Coli Complex I D79N Nuoa Mutant Membrane Domain In Lmng
Organism: Escherichia coli bw25113
Method: ELECTRON MICROSCOPY Release Date: 2026-05-06 Classification: PROTON TRANSPORT Ligands: 3PE, 7PH, CDL, UQ8 |
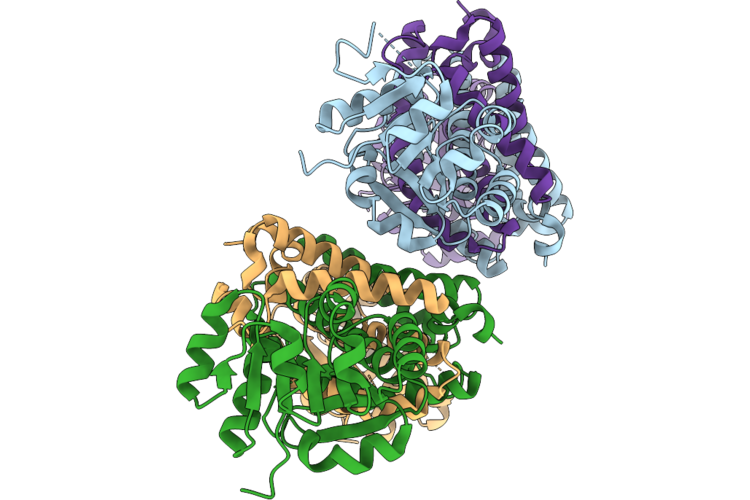 |
Crystal Structure Of Imine Reductase Mutant(Ahtanredam) From Actinoalloteichus Hymeniacidonis In Complex With Nadph
Organism: Actinoalloteichus hymeniacidonis
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE |
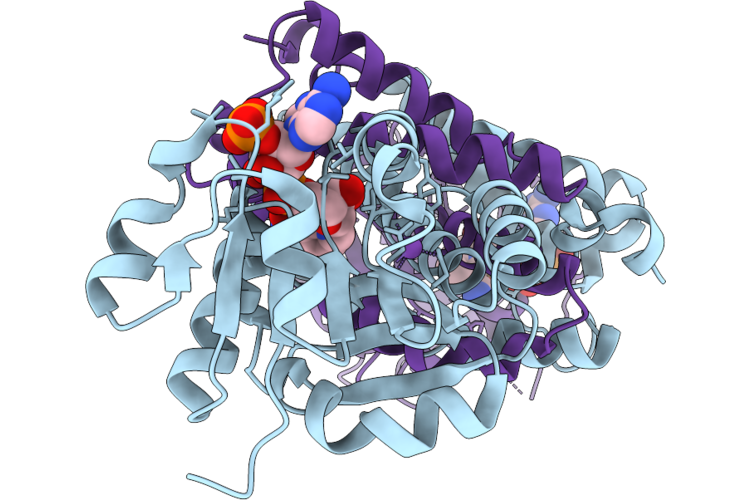 |
Crystal Structure Of Imine Reductase Mutant(Ahtanredam) From Actinoalloteichus Hymeniacidonis In Complex With Nadph
Organism: Actinoalloteichus hymeniacidonis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-05-06 Classification: OXIDOREDUCTASE Ligands: NAP, NA |
 |
Crystal Structure Of Human L-Lactate Dehydrogenase B Protein In Complex With Nadh, Oxamate And Sertraline
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.06 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: OXM, NAI, SRE |
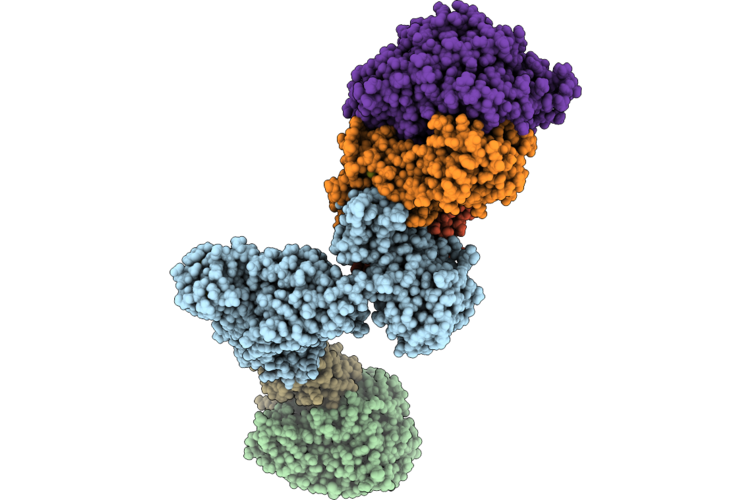 |
Structure Of The Mvhagd-Hdrabc Dimer Of M. Marburgensis Under State 2 Substate A (Composite Structure)
Organism: Methanothermobacter marburgensis str. marburg
Method: ELECTRON MICROSCOPY Resolution:2.19 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, 9S8, FES, NFU |
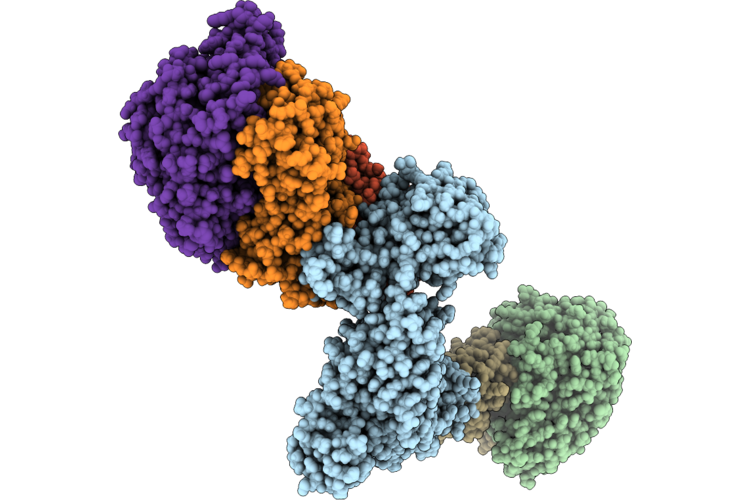 |
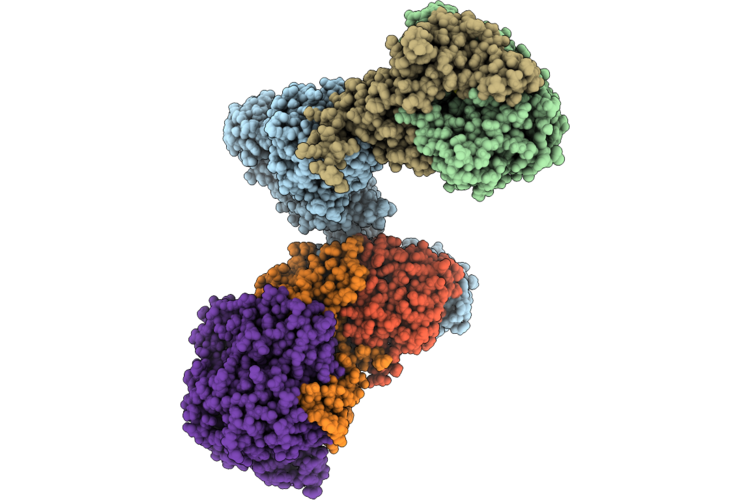
Structure Of The Mvhagd-Hdrabc Dimer Of M. Marburgensis Under State 2 Substate B (Composite Structure)
Organism: Methanothermobacter marburgensis
Method: ELECTRON MICROSCOPY Resolution:3.25 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, 9S8, FES, NFU |
 |
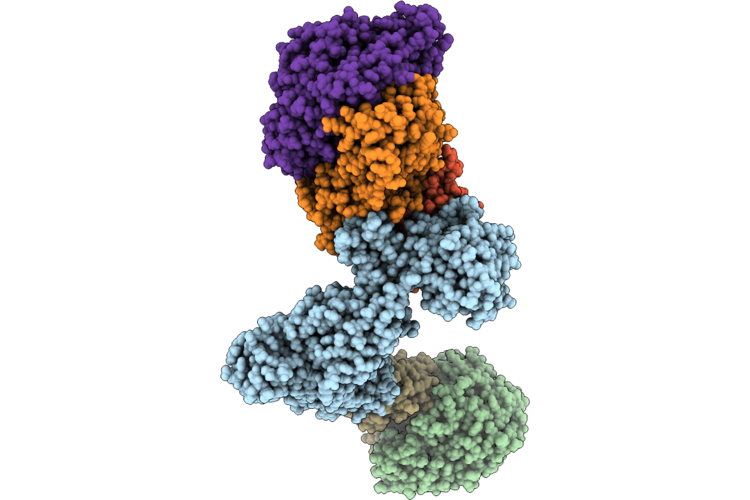
Structure Of The Mvhagd-Hdrabc Dimer Of M. Marburgensis Under State 1 Substate A (Composite Structure)
Organism: Methanothermobacter marburgensis
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, FES, 9S8, NFU |
 |
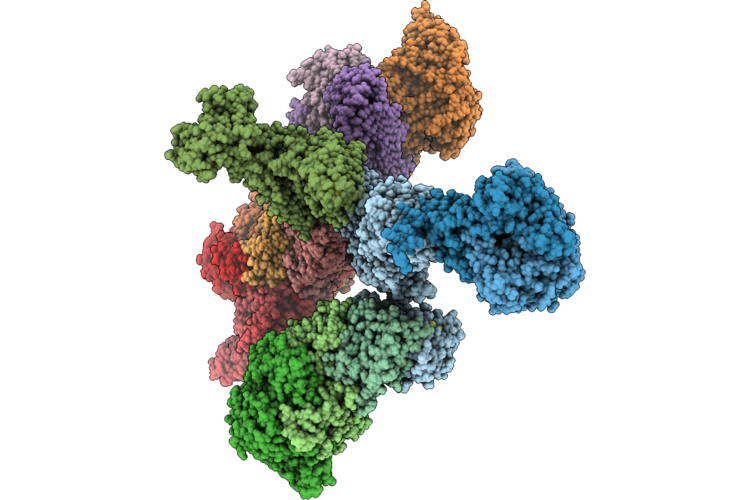
Structure Of The Mvhagd-Hdrabc Dimer Of M. Marburgensis Under State 1 Substate B (Composite Structure)
Organism: Methanothermobacter marburgensis
Method: ELECTRON MICROSCOPY Resolution:3.29 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, 9S8, NFU, FES |
 |
Structure Of The Mvh-Hdr-Fmd Complex Of Methanothermobacter Marburgensis (Composite Structure)
Organism: Methanothermobacter marburgensis
Method: ELECTRON MICROSCOPY Resolution:2.70 Å Release Date: 2026-04-29 Classification: OXIDOREDUCTASE Ligands: SF4, FAD, 9S8, FES, ZN, MGD, MO, H2S, F3S, NFU |

