Search Count: 424
All
Selected
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-29 Classification: RNA Ligands: SPM |
 |
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-29 Classification: RNA |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.26 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: XDC, ZN, MG, CA, GOL, NA, NAG, MAN, PO4, TRS, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.12 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: PO4, ZN, MG, CA, NA, NAG, GOL, TRS, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.25 Å Release Date: 2026-04-22 Classification: HYDROLASE |
 |
Tissue Non-Specific Alkaline Phosphatase -S110A Bound To Ppi With Ethylene Glycol
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: POP, ZN, MG, CA, NA, NAG, MAN, PO4, FLC |
 |
Organism: Mus musculus
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-04-22 Classification: HYDROLASE Ligands: ATP, ZN, MG, CA, GOL, NAG, FLC, PO4 |
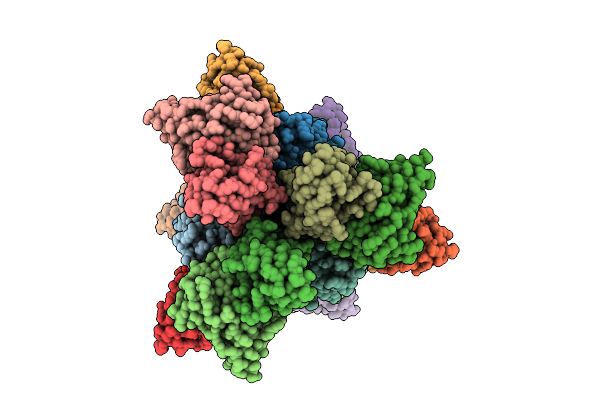 |
Cryo-Em Structure Of Sudan Ebolavirus Gp Bound By Three Neutralizing Antibodies 545S, 523S And 294S
Organism: Sudan ebolavirus, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
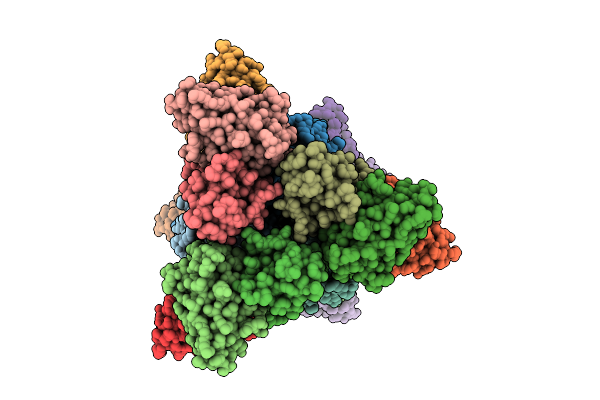 |
Cryo-Em Structure Of Sudan Ebolavirus Gp Bound By Three Neutralizing Antibodies 316L, 523S And 294S
Organism: Sudan ebolavirus, Macaca mulatta, Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: VIRAL PROTEIN/IMMUNE SYSTEM |
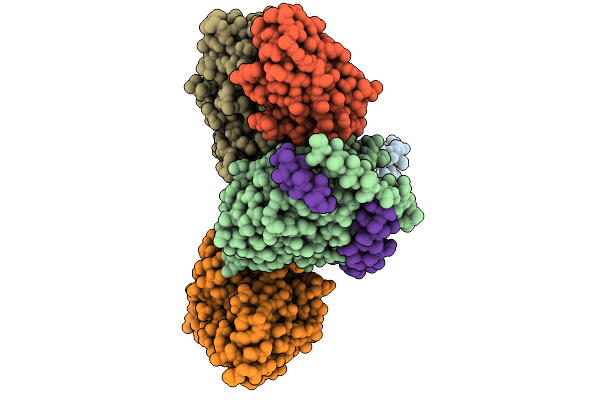 |
Organism: Synthetic construct, Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.60 Å Release Date: 2026-01-28 Classification: MEMBRANE PROTEIN/IMMUNE SYSTEM |
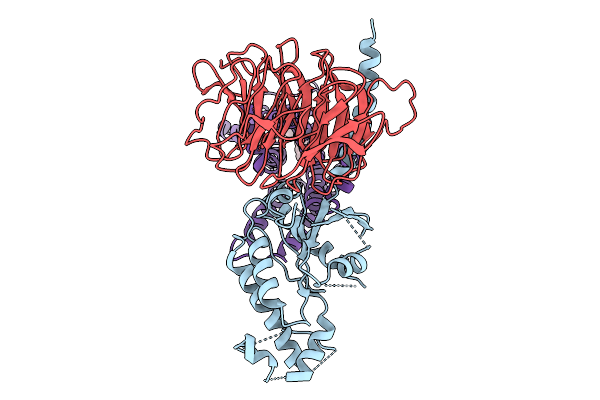 |
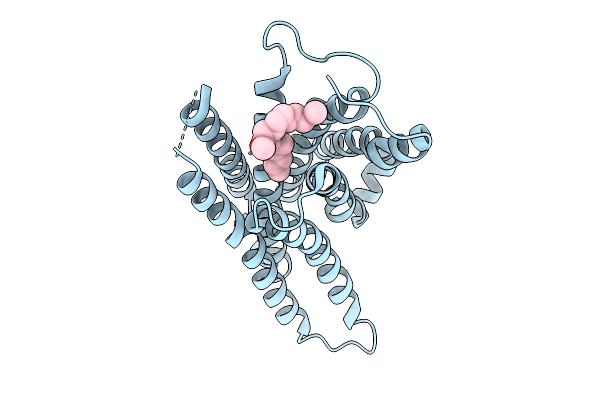
Organism: Synthetic construct, Homo sapiens, Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.70 Å Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: A1EKK |
 |
Organism: Mus musculus
Method: ELECTRON MICROSCOPY Resolution:3.60 Å Release Date: 2026-01-21 Classification: MEMBRANE PROTEIN Ligands: A1EKK |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: RIBOSOME Ligands: MG, ZN, ANM |
 |
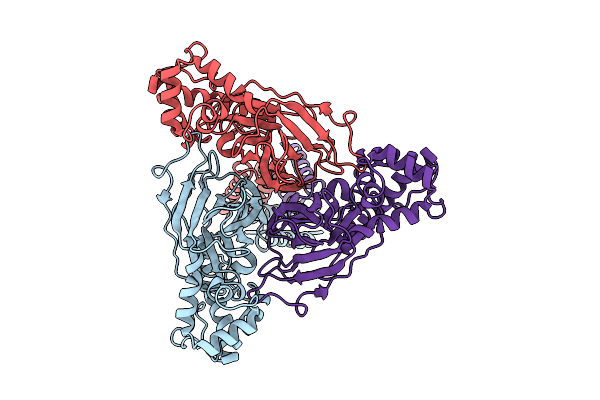
Structure Of Rack1 Bound To The C-Terminus Of Serbp1 And The Rih Region Of Zak
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2025-12-10 Classification: RIBOSOME |
 |
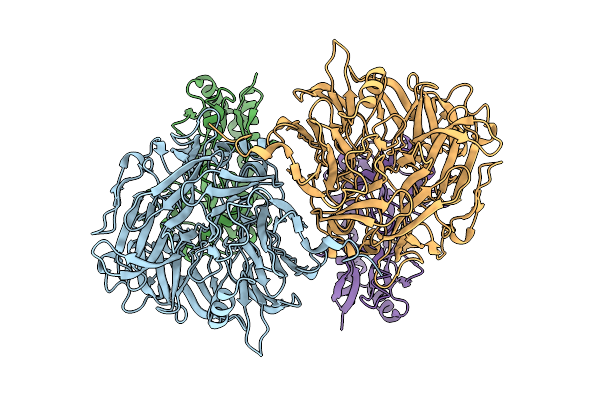
Organism: Scolopendra mutilans
Method: ELECTRON MICROSCOPY Release Date: 2025-08-20 Classification: MEMBRANE PROTEIN |
 |
Organism: Methylorubrum extorquens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: PROTEIN BINDING |
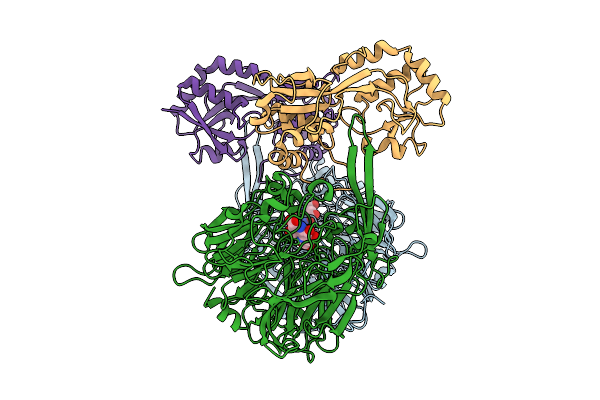 |
Organism: Methylorubrum extorquens
Method: ELECTRON MICROSCOPY Release Date: 2025-07-23 Classification: PROTEIN BINDING Ligands: PQQ |
 |
Organism: Lotus japonicus
Method: X-RAY DIFFRACTION Resolution:1.64 Å Release Date: 2025-07-16 Classification: PLANT PROTEIN Ligands: NAG, ACT, GOL |
 |
Organism: Medicago truncatula
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2025-07-16 Classification: PLANT PROTEIN Ligands: NAG, SO4, EDO |
 |
Organism: Medicago truncatula
Method: X-RAY DIFFRACTION Resolution:2.98 Å Release Date: 2025-07-16 Classification: PLANT PROTEIN Ligands: NAG |

