Search Count: 20,143
All
Selected
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.03 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN,HYDROLASE/INHIBITOR Ligands: A1A5Z, CL |
 |
Organism: Severe acute respiratory syndrome coronavirus 2
Method: X-RAY DIFFRACTION Resolution:1.02 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN,HYDROLASE/INHIBITOR Ligands: A1A54, CL |
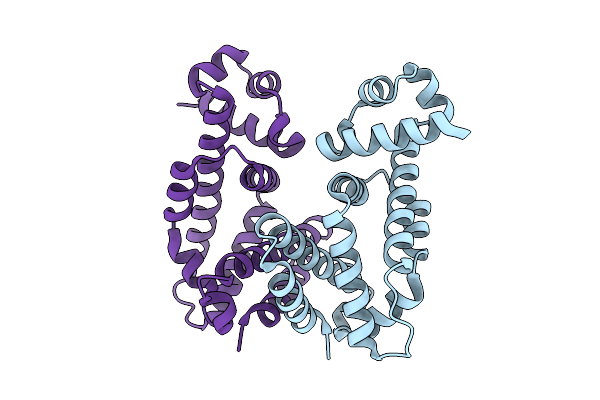 |
Organism: Salmonella enterica subsp. enterica serovar typhimurium
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-08 Classification: TRANSCRIPTION Ligands: CL |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: LDP, NA, CL |
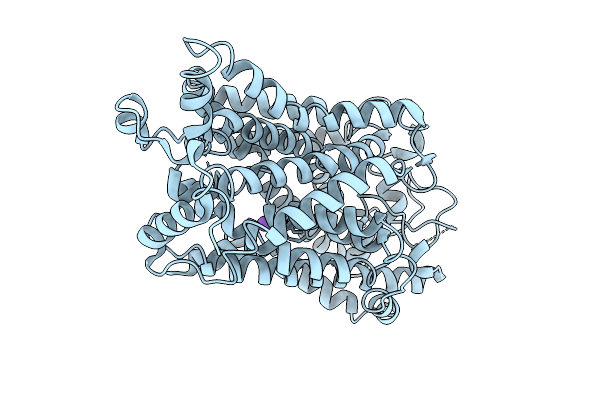 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: CL, K |
 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: MEMBRANE PROTEIN Ligands: A1D5S, CL |
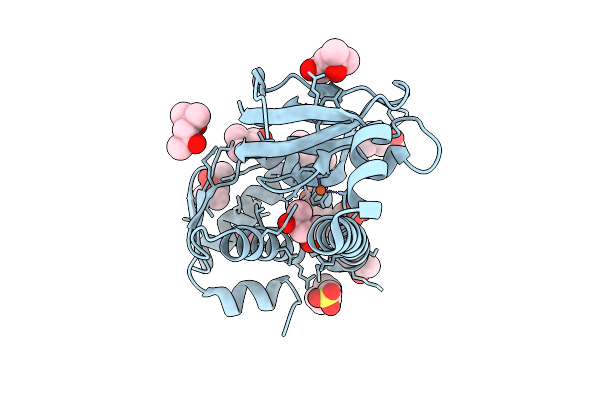 |
Organism: Aliivibrio fischeri
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-08 Classification: TRANSCRIPTION Ligands: FE, MPD, MES, CL |
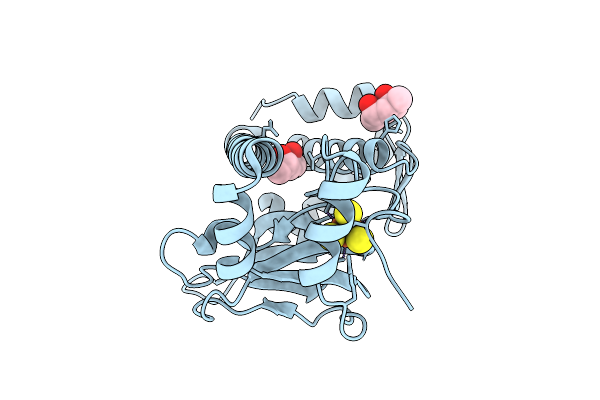 |
Organism: Aliivibrio fischeri
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-08 Classification: TRANSCRIPTION Ligands: SF4, MPD, CL |
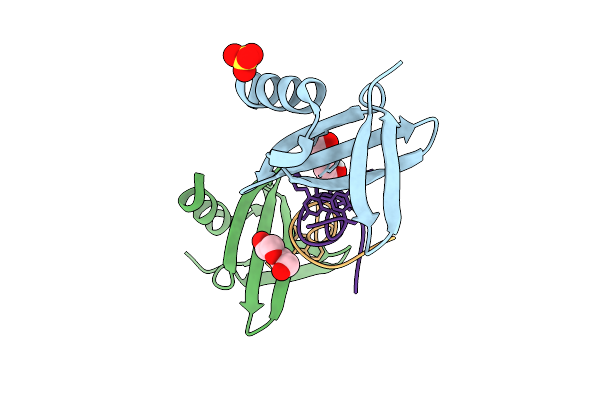 |
Crystal Structure Of Nanofitin C10 In Complex With A A Double-Helical Aromatic Oligoamide Foldamer
Organism: Sulfolobus acidocaldarius, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-04-08 Classification: DE NOVO PROTEIN Ligands: SO4, PEG, CL |
 |
Organism: Pseudomonas aeruginosa
Method: X-RAY DIFFRACTION Resolution:1.79 Å Release Date: 2026-04-08 Classification: ANTIMICROBIAL PROTEIN Ligands: CL |
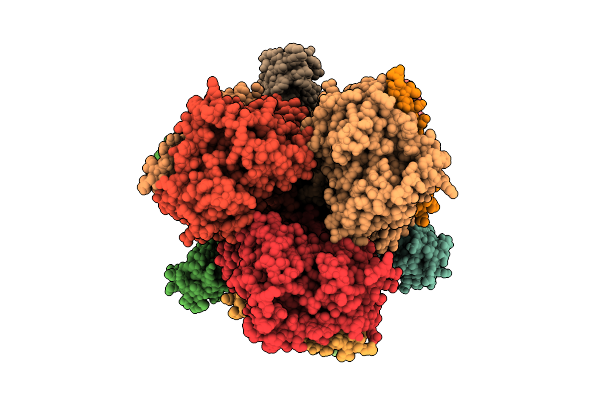 |
Organism: Escherichia coli
Method: ELECTRON MICROSCOPY Resolution:3.17 Å Release Date: 2026-04-08 Classification: TRANSPORT PROTEIN Ligands: CL |
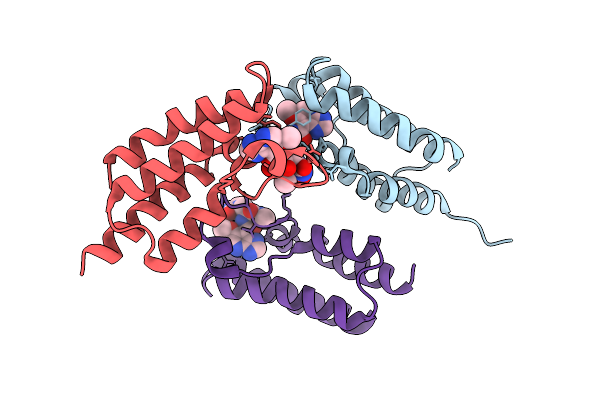 |
Crystal Structure Of Bromodomain From Plasmodium Falciparum Gcn5 Complexed With A Ligand
Organism: Plasmodium falciparum 3d7
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: A1JWP, CL |
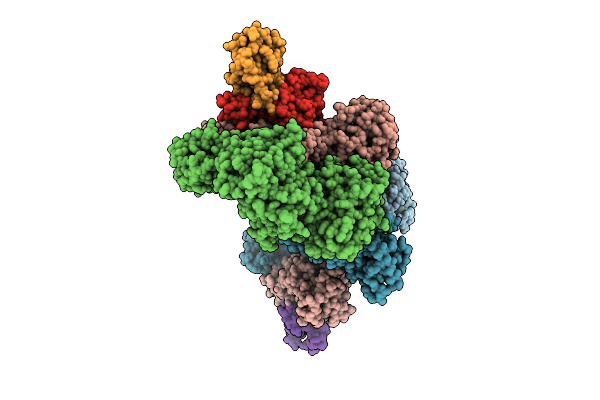 |
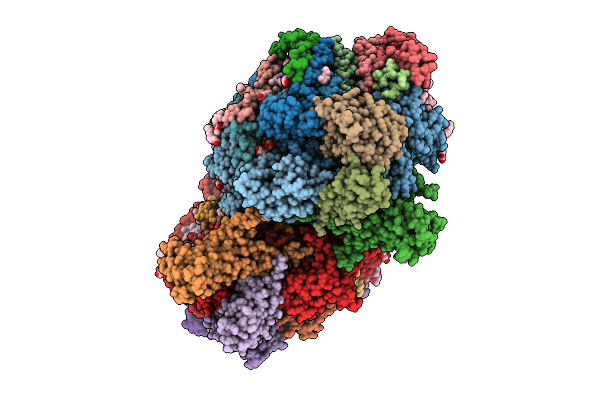
Complex Of N-Terminal Brxc Walker B, Brxb, And N-Termianl Pglz From The Acinetobacter Brex System
Organism: Acinetobacter sp. neb 394
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2026-04-08 Classification: ANTIMICROBIAL PROTEIN Ligands: ATP, CA, CL, MG |
 |
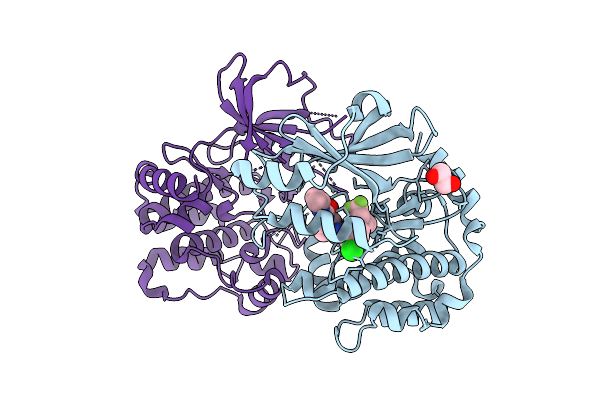
Single-Conformation Model Re-Refinement Of 2F/S3-Rich Psii Intermediate Structure At 2.09 Angstrom Resolution
Organism: Thermosynechococcus vestitus bp-1
Method: X-RAY DIFFRACTION Resolution:2.09 Å Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: FE2, CLA, BCR, CL, PL9, SQD, DGD, STE, OEX, LMG, LHG, BCT, PHO, HEC |
 |
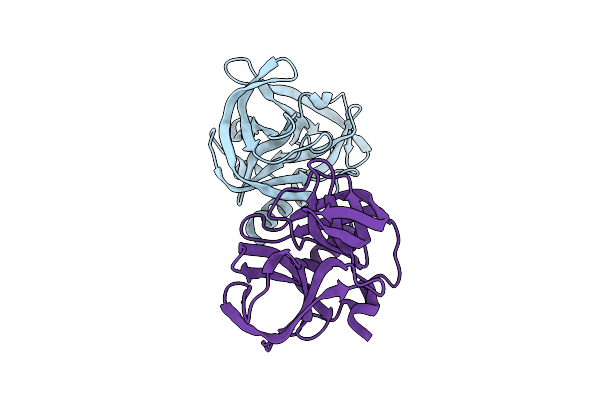
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.28 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: CL, A1JZR, GOL |
 |
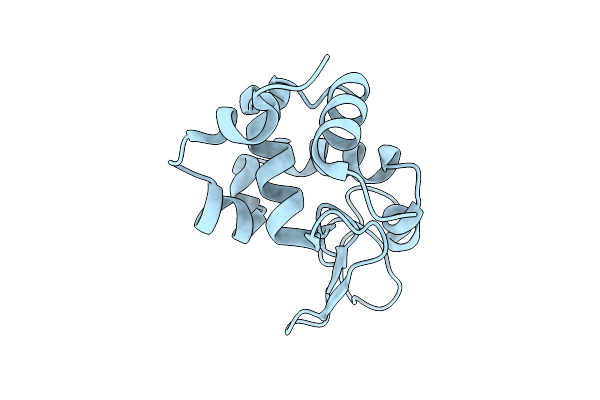
Crystal Structure Of Enterovirus D68 3C Protease Determined Via Sulfur Phasing
Organism: Enterovirus d68
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-01 Classification: VIRAL PROTEIN Ligands: CL |
 |
Organism: Gallus gallus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-01 Classification: HYDROLASE Ligands: CL |
 |
Organism: Leishmania donovani
Method: X-RAY DIFFRACTION Resolution:1.30 Å Release Date: 2026-04-01 Classification: CELL CYCLE Ligands: ZN, ACT, NA, CL, EDO |
 |
Crystal Structure Of Heme Binding Pas Domain From One Component Transcription Factor, Fg214
Organism: Fimbriimonas ginsengisoli gsoil 348
Method: X-RAY DIFFRACTION Resolution:1.47 Å Release Date: 2026-03-25 Classification: TRANSCRIPTION Ligands: HEM, IMD, PO4, CL, PG4, EDO, MG, FMT, POL |
 |
Crystal Structure Of Gdp-Bound Kras G12D/I55E: Suppressing G12D Oncogenicity Via Second-Site I55E Mutation
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.23 Å Release Date: 2026-03-25 Classification: ONCOPROTEIN Ligands: GDP, MG, CL |

