Search Count: 35,830
All
Selected
 |
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme Sub-Conformation 1
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme Sub-Conformation 2
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme Sub-Conformation 3
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
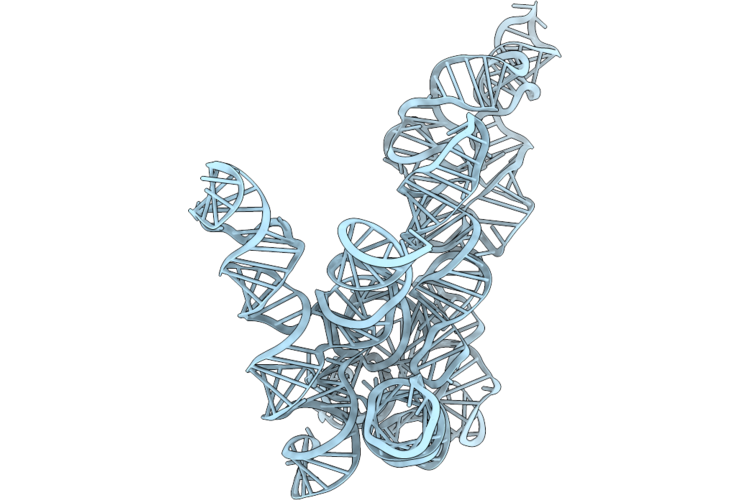 |
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: MG |
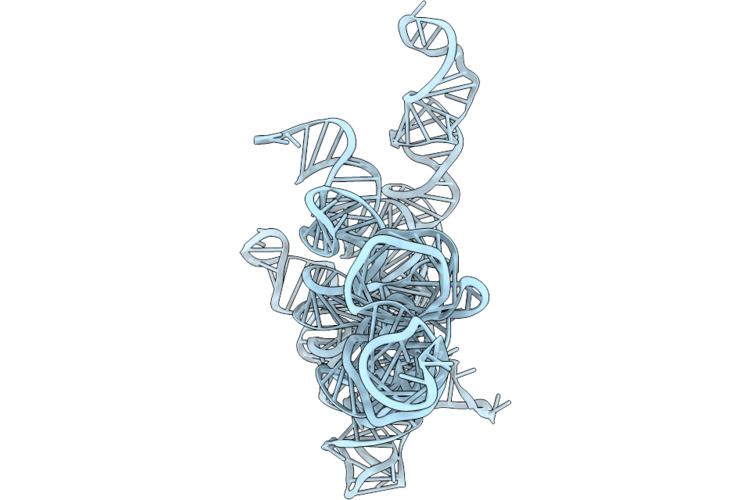 |
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: MG |
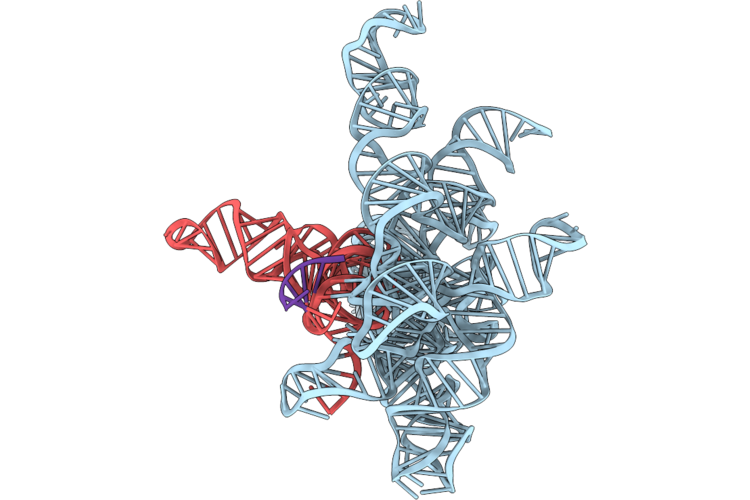 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme In Complex With Precursor Trna In 5 Mm Ca2+
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
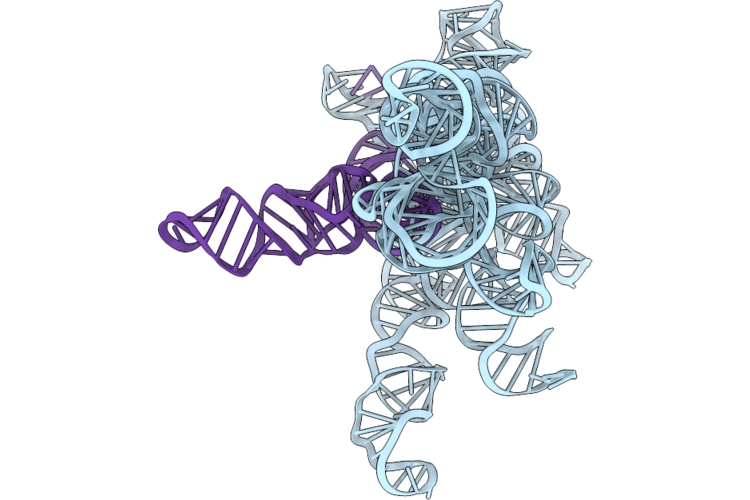 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme In Complex With Mature Trna In 5 Mm Ca2+
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
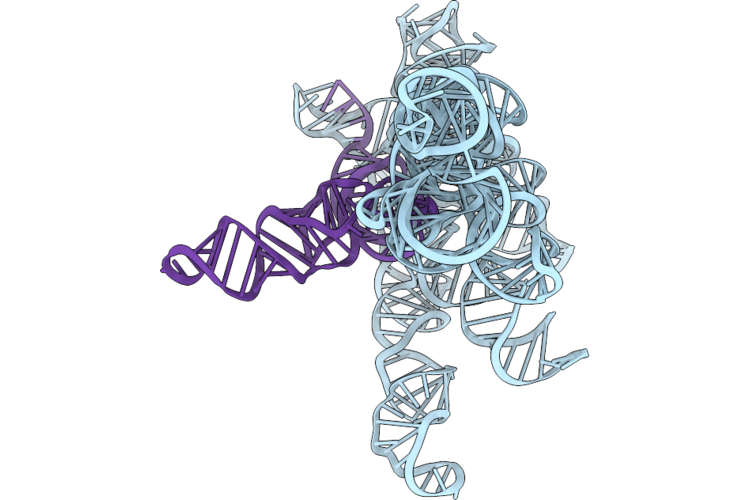 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme In Complex With Mature Trna In 10 Mm Ca2+
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
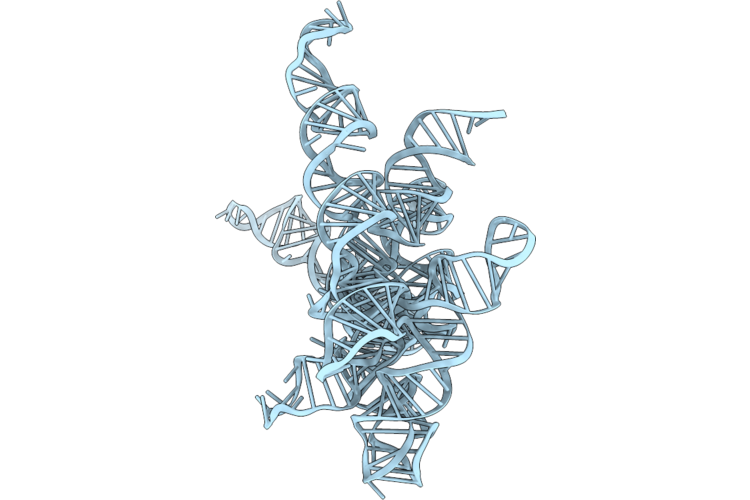 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme Tetraloop Mutant (Sub-Conformation 1)
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
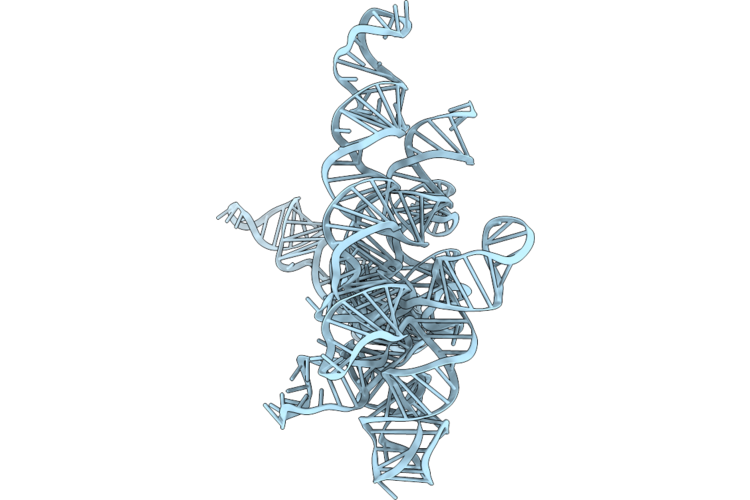 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme Tetraloop Mutant (Sub-Conformation 2)
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
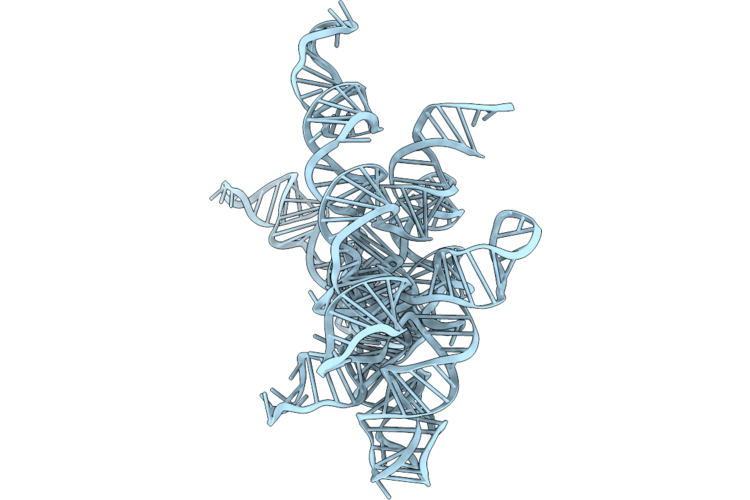 |
Structure Of Geobacillus Stearothermophilus Rnase P Ribozyme Tetraloop Mutant (Sub-Conformation 3)
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Release Date: 2026-05-20 Classification: RNA Ligands: CA |
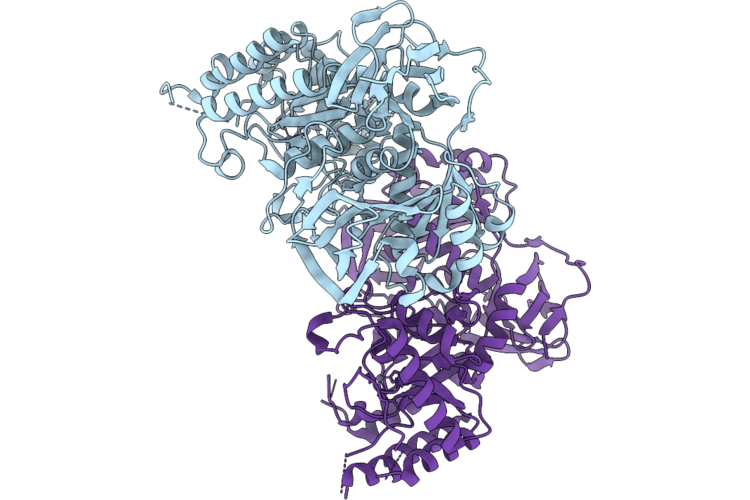 |
Organism: Bacillus cereus atcc 10987
Method: X-RAY DIFFRACTION Resolution:2.51 Å Release Date: 2026-05-20 Classification: MEMBRANE PROTEIN Ligands: ZN |
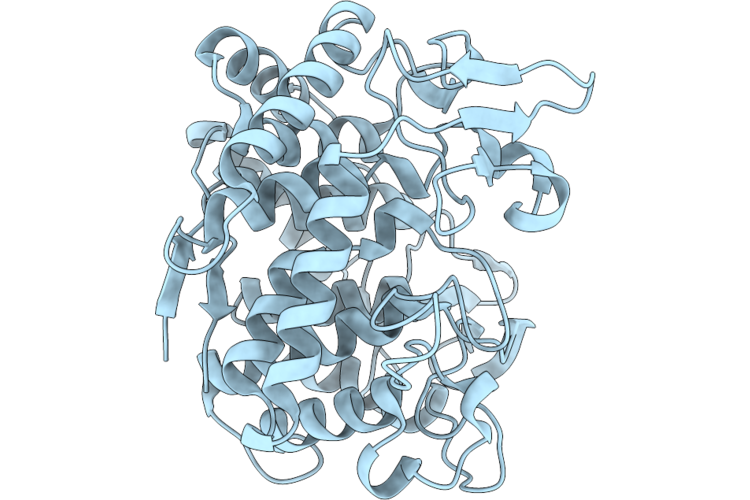 |
Organism: Bacillus glycinifermentans
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-05-13 Classification: HYDROLASE |
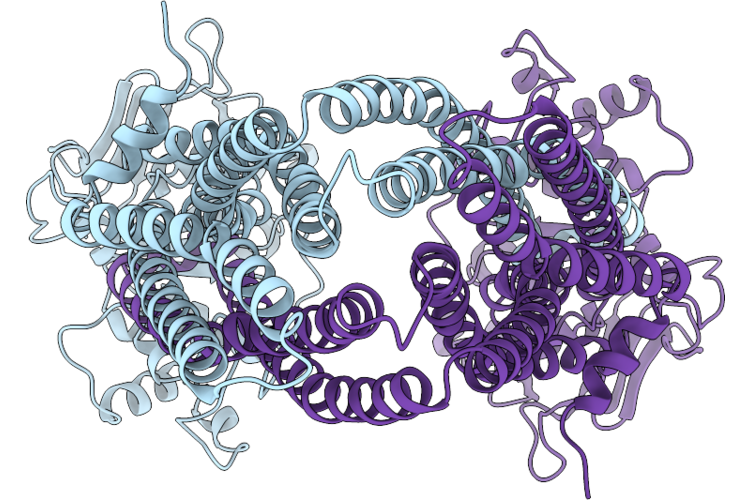 |
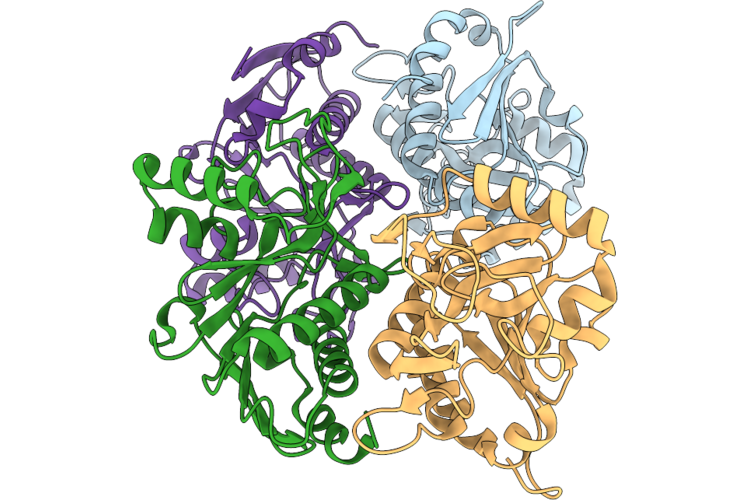
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: TRANSPORT PROTEIN |
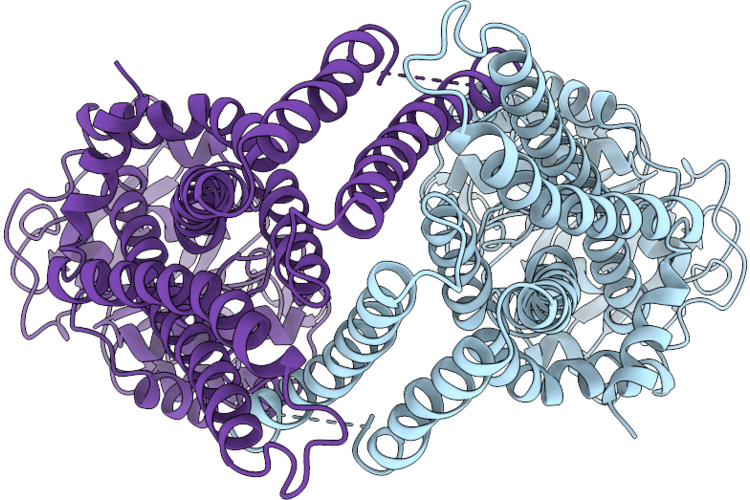 |
Organism: Bacillus subtilis
Method: ELECTRON MICROSCOPY Release Date: 2026-05-13 Classification: TRANSPORT PROTEIN |
 |
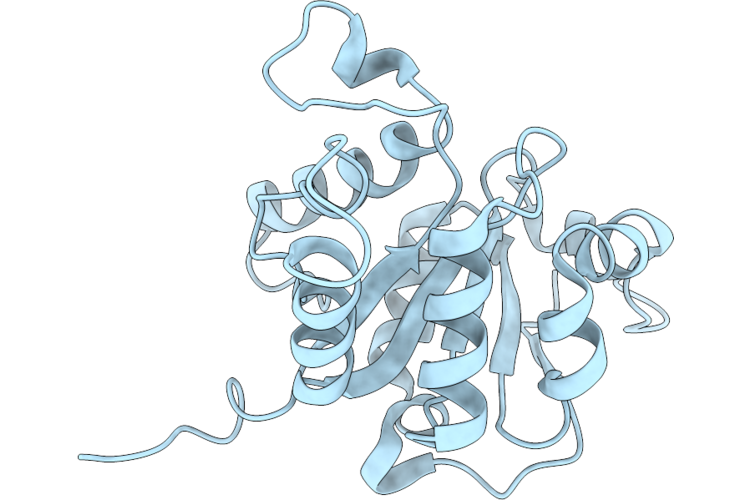
Organism: Bacillus velezensis yau b9601-y2
Method: X-RAY DIFFRACTION Resolution:2.59 Å Release Date: 2026-05-13 Classification: HYDROLASE |
 |
Organism: Bacillus velezensis yau b9601-y2
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-05-13 Classification: HYDROLASE |
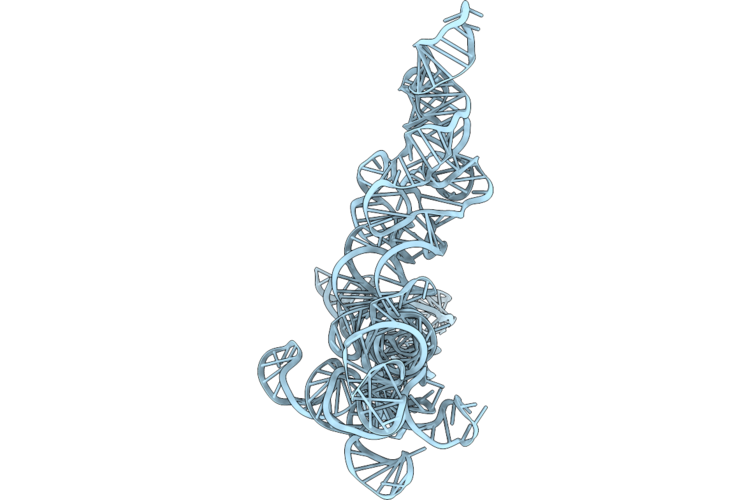 |
Cryo-Em Structure Of Rnase P Rna From Geobacillus Stearothermophilus, Conformer 51
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Resolution:3.30 Å Release Date: 2026-05-13 Classification: RNA Ligands: MG |
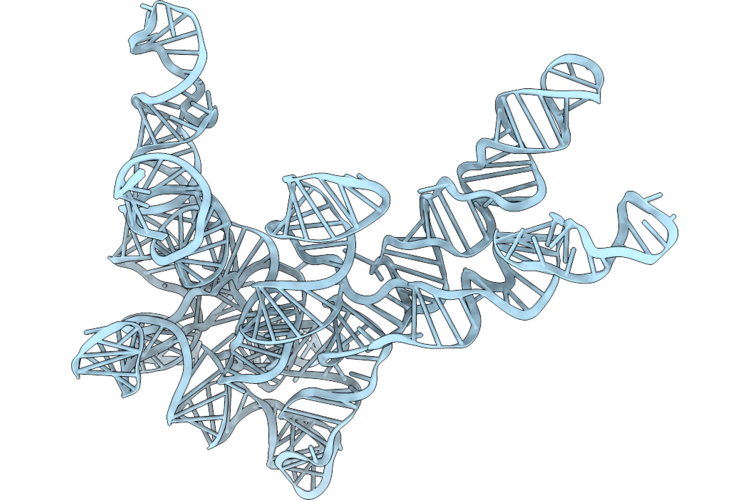 |
Cryo-Em Structure Of Rnase P Rna From Geobacillus Stearothermophilus, Conformer 1
Organism: Geobacillus stearothermophilus
Method: ELECTRON MICROSCOPY Resolution:2.90 Å Release Date: 2026-05-13 Classification: RNA Ligands: MG |

