Search Count: 2,070
All
Selected
 |
Organism: Streptomyces prunicolor
Method: X-RAY DIFFRACTION Resolution:1.96 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: GOL, PEG |
 |
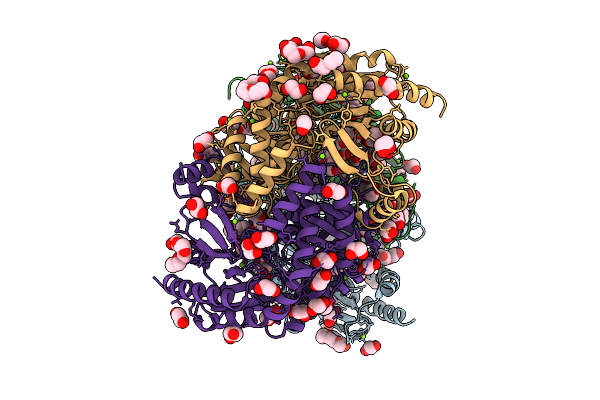
Aminoacyl-Trna-Dependent Peptide Synthase, Sbb17, Complexed With Streptothrisamine
Organism: Streptomyces prunicolor
Method: X-RAY DIFFRACTION Resolution:2.50 Å Release Date: 2026-04-01 Classification: TRANSFERASE Ligands: A1L3W |
 |
Organism: Streptomyces sp. bv333
Method: X-RAY DIFFRACTION Resolution:1.24 Å Release Date: 2026-03-18 Classification: TRANSFERASE Ligands: PEG, PGE, EDO, GOL, 1PG, CL, MG, PG4 |
 |
Organism: Streptomyces sp. bv333
Method: X-RAY DIFFRACTION Resolution:1.31 Å Release Date: 2026-03-18 Classification: TRANSFERASE |
 |
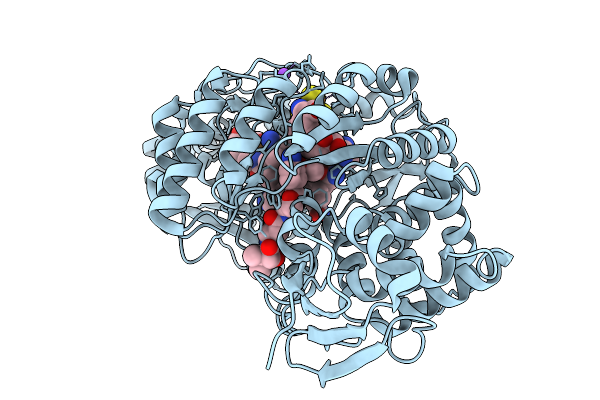
Crystal Structure Of Cyss From Corallococcus Sp. Ca054B With 5'-Deoxyadenosine, Methionine, And Pantetheinylated 3-Methoxyl-4-Amino Benzoic Acid Substrate Bound
Organism: Corallococcus sp. ca054b
Method: X-RAY DIFFRACTION Resolution:1.75 Å Release Date: 2026-01-28 Classification: OXIDOREDUCTASE/SUBSTRATE Ligands: B12, SF4, 5AD, MET, A1BUN, MG |
 |
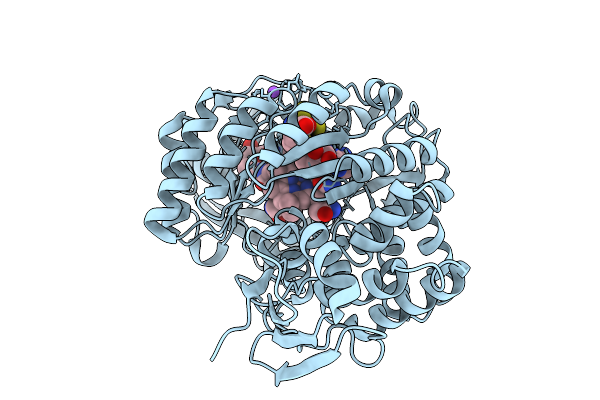
Crystal Structure Of Cyss From Corallococcus Sp. Ca054B With 5'-Deoxyadenosine, Methionine, And Pantetheinylated 3-Ethoxy-4-Amino Benzoic Acid Substrate Bound
Organism: Corallococcus sp. ca054b
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-01-28 Classification: OXIDOREDUCTASE/SUBSTRATE Ligands: B12, SF4, 5AD, MET, A1BUP, NA |
 |
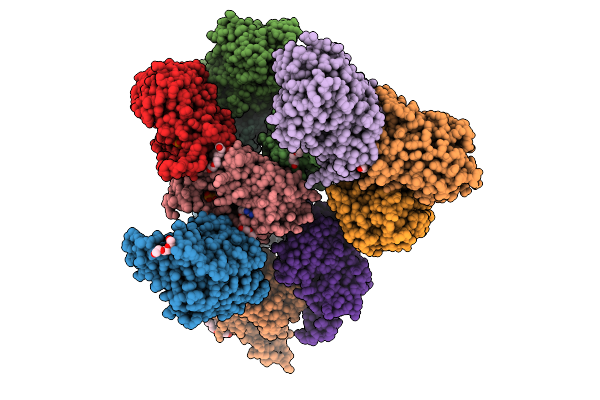
Crystal Structure Of Cyss From Corallococcus Sp. Ca054B With 5'-Deoxyadenosine And Methionine Bound
Organism: Corallococcus sp. ca054b
Method: X-RAY DIFFRACTION Resolution:1.94 Å Release Date: 2026-01-28 Classification: OXIDOREDUCTASE Ligands: B12, SF4, 5AD, MET, NA |
 |
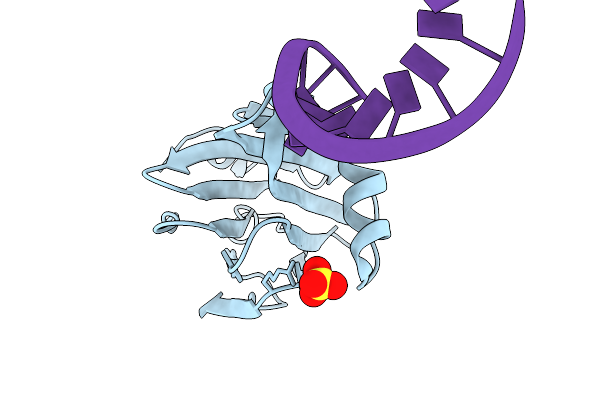
Sam-Dependent C6-Fpp Methytransferase From Streptomyces Varsoviensis In Complex With Sah And Fpp
Organism: Streptomyces varsoviensis
Method: X-RAY DIFFRACTION Resolution:2.20 Å Release Date: 2026-01-28 Classification: TRANSFERASE Ligands: FPP, SAH, PG4, PGE, PE8, PEG, EDO |
 |
Organism: Grimontia hollisae, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.42 Å Release Date: 2026-01-14 Classification: TOXIN Ligands: SO4 |
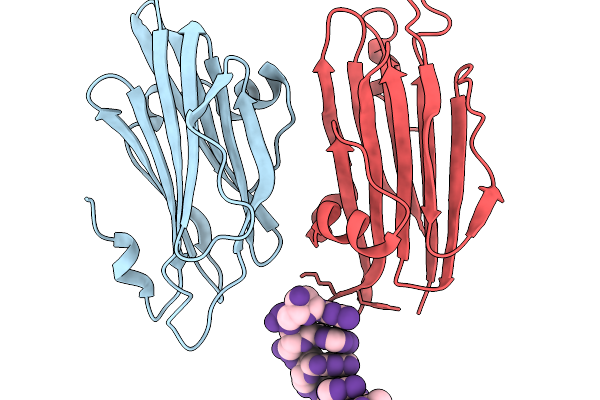 |
Grimontia Hollisae Thermostable Direct Hemolysin In Complex With 6-Nt Blunt-Ended Dsdna
Organism: Grimontia hollisae, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.82 Å Release Date: 2026-01-14 Classification: TOXIN |
 |
Grimontia Hollisae Thermostable Direct Hemolysin In Complex With 2-Nt Long 5'-Overhang Dsdna
Organism: Grimontia hollisae, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.58 Å Release Date: 2026-01-14 Classification: TOXIN |
 |
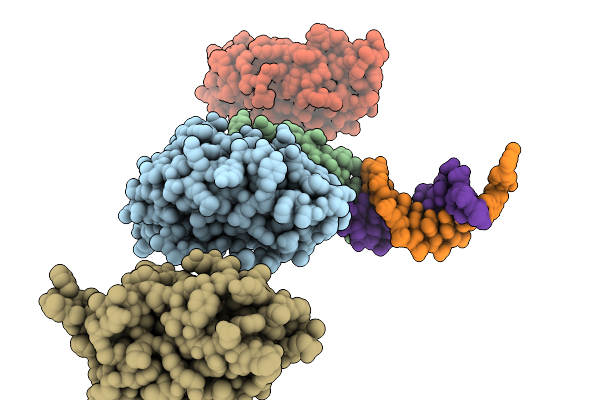
Grimontia Hollisae Thermostable Direct Hemolysin In Complex With 1-Nt Long 3'-Overhang Dsdna
Organism: Grimontia hollisae, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2026-01-14 Classification: TOXIN |
 |
Grimontia Hollisae Thermostable Direct Hemolysin K88A Mutant In Complex With 1-Nt Long 3'-Overhang Dsdna
Organism: Grimontia hollisae, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:2.71 Å Release Date: 2026-01-14 Classification: TOXIN |
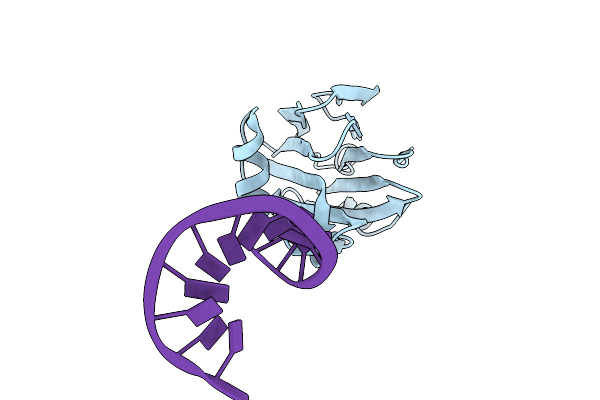 |
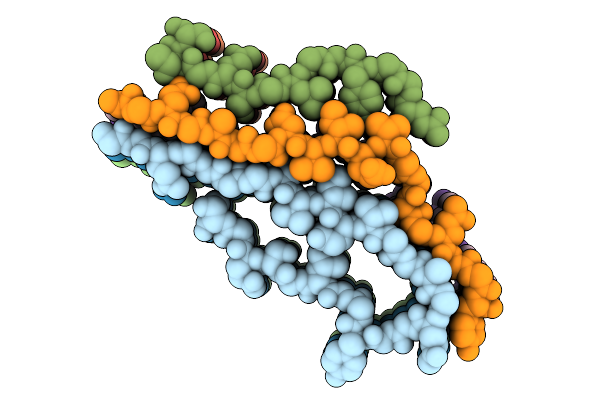
Structure Of Gh-Tdh With Additional N-Terminus In Complex With Double-Stranded Dna Containing A 2-Nucleotide 5' Overhang
Organism: Grimontia hollisae, Synthetic construct
Method: X-RAY DIFFRACTION Resolution:1.37 Å Release Date: 2025-12-31 Classification: TOXIN |
 |
Organism: Latrodectus hesperus
Method: ELECTRON MICROSCOPY Release Date: 2025-12-24 Classification: PROTEIN FIBRIL |
 |
Organism: Latrodectus hesperus
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: PROTEIN FIBRIL |
 |
Organism: Latrodectus hesperus
Method: ELECTRON MICROSCOPY Release Date: 2025-12-17 Classification: PROTEIN FIBRIL |
 |
Crystal Structure Of Cyua C289A Mutant From Methanococcus Maripaludis With [4Fe-4S] Clusters In Complex With Glycerol
Organism: Methanococcus maripaludis s2
Method: X-RAY DIFFRACTION Resolution:3.18 Å Release Date: 2025-11-05 Classification: LYASE Ligands: SF4, GOL |
 |
Crystal Structure Of Cyua C289A Mutant From Methanococcus Maripaludis With [4Fe-4S] Clusters In Complex With Ethylene Glycol
Organism: Methanococcus maripaludis s2
Method: X-RAY DIFFRACTION Resolution:3.74 Å Release Date: 2025-11-05 Classification: LYASE Ligands: SF4, EDO |
 |
Repetitive Domain (Rp) 2 Structure Of Aciniform Spidroin From Latrodectus Hesperus
Organism: Latrodectus hesperus
Method: X-RAY DIFFRACTION Resolution:1.06 Å Release Date: 2025-10-01 Classification: STRUCTURAL PROTEIN |

