Search Count: 45,373
All
Selected
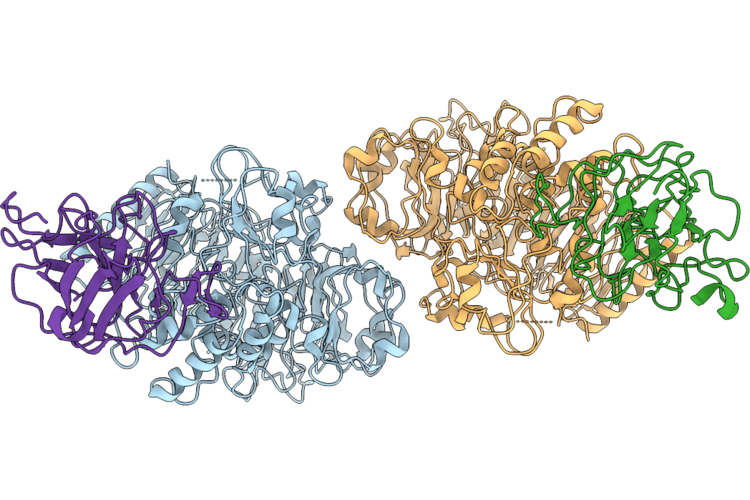 |
Cryo-Em Structure Of Plant Resistance Protein Nrc2 Dimer Bound To Nematode Effector Sprysec-15
Organism: Nicotiana benthamiana, Globodera rostochiensis
Method: ELECTRON MICROSCOPY Release Date: 2026-04-29 Classification: PLANT PROTEIN |
 |
Organism: Influenza b virus (b/brisbane/60/2008), Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.76 Å Release Date: 2026-04-29 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Influenza b virus (b/brisbane/60/2008), Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.76 Å Release Date: 2026-04-29 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Organism: Influenza b virus (b/brisbane/60/2008), Homo sapiens
Method: ELECTRON MICROSCOPY Resolution:2.65 Å Release Date: 2026-04-29 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
Crystal Structure Of A Cupin Protein (Tm1459, H52A/H54A/H92A/C106E Mutant) In Ruthenium(P-Cymene) Bound Form
Organism: Thermotoga maritima (strain atcc 43589 / dsm 3109 / jcm 10099 / nbrc 100826 / msb8)
Method: X-RAY DIFFRACTION Resolution:1.27 Å Release Date: 2026-04-29 Classification: METAL BINDING PROTEIN Ligands: MML, RU, MES |
 |
Crystal Structure Of A Cupin Protein (Tm1459, R39K/H52A/H54A/H92A/C106E Mutant) In Ruthenium(P-Cymene) Bound Form
Organism: Thermotoga maritima (strain atcc 43589 / dsm 3109 / jcm 10099 / nbrc 100826 / msb8)
Method: X-RAY DIFFRACTION Resolution:1.09 Å Release Date: 2026-04-29 Classification: METAL BINDING PROTEIN Ligands: MML, RU |
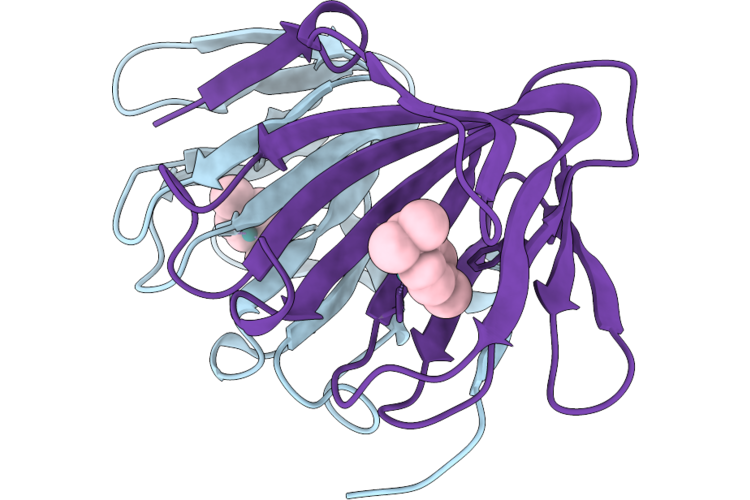 |
Crystal Structure Of A Cupin Protein (Tm1459, R39M/H52A/H54A/H92A/C106E Mutant) In Ruthenium(P-Cymene) Bound Form
Organism: Thermotoga maritima (strain atcc 43589 / dsm 3109 / jcm 10099 / nbrc 100826 / msb8)
Method: X-RAY DIFFRACTION Resolution:1.42 Å Release Date: 2026-04-29 Classification: METAL BINDING PROTEIN Ligands: MML, RU |
 |
Structure Of Bacillus Subtilis Nitric Oxide Synthase In Complex With 7-((3-(((3-(6-Aminopyridin-2-Yl)Propyl)Amino)Methyl)Phenoxy)Methyl)Quinolin-2-Amine
Organism: Bacillus subtilis subsp. subtilis str. 168
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-22 Classification: OXIDOREDUCTASE Ligands: HEM, V5D, CL, PEG, GOL, POL |
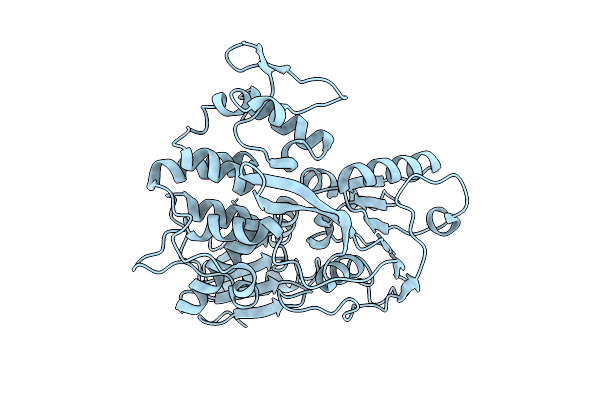 |
Organism: Cacipacore virus
Method: X-RAY DIFFRACTION Resolution:2.07 Å Release Date: 2026-04-15 Classification: HYDROLASE |
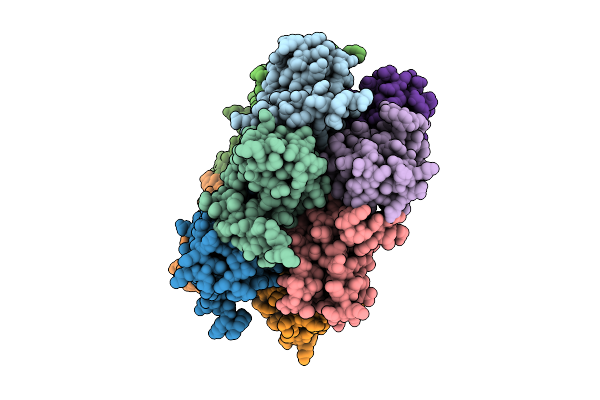 |
Organism: Trypanosoma brucei brucei (strain 927/4 gutat10.1)
Method: X-RAY DIFFRACTION Resolution:3.50 Å Release Date: 2026-04-15 Classification: TRANSCRIPTION |
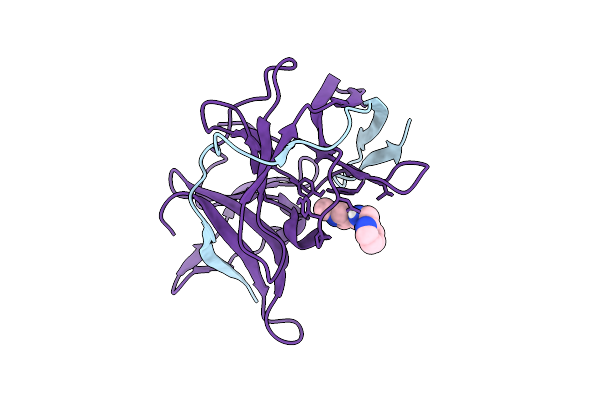 |
Crystal Structure Of Zika Virus Ns2B-Ns3 Protease In Complex With Z100582766
Organism: Zika virus
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN Ligands: A1I9X |
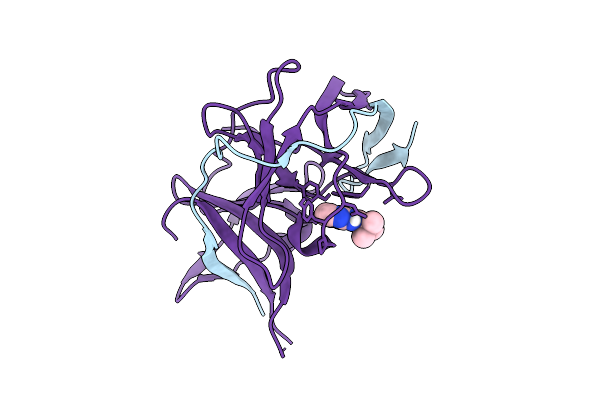 |
Crystal Structure Of Zika Virus Ns2B-Ns3 Protease In Complex With Z2242050769
Organism: Zika virus
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN Ligands: A1JAB |
 |
Organism: Zika virus
Method: X-RAY DIFFRACTION Resolution:2.40 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN Ligands: MG, EDO |
 |
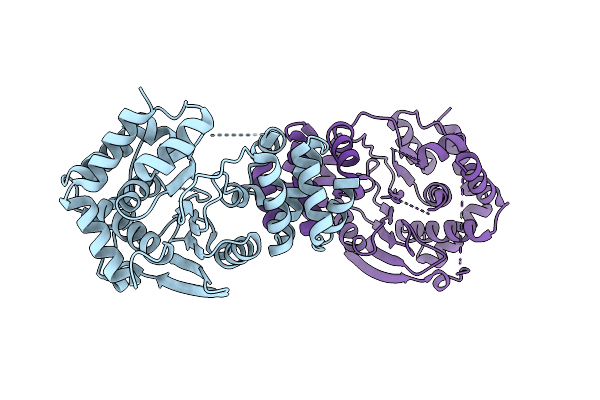
Crystal Structure Of A Zikv E Glycoprotein Di-Diii Vaccine Candidate In Complex With Human Neutralizing Antibody Mz4
Organism: Homo sapiens, Zika virus zikv/h. sapiens/frenchpolynesia/10087pf/2013
Method: X-RAY DIFFRACTION Resolution:2.85 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN Ligands: NAG |
 |
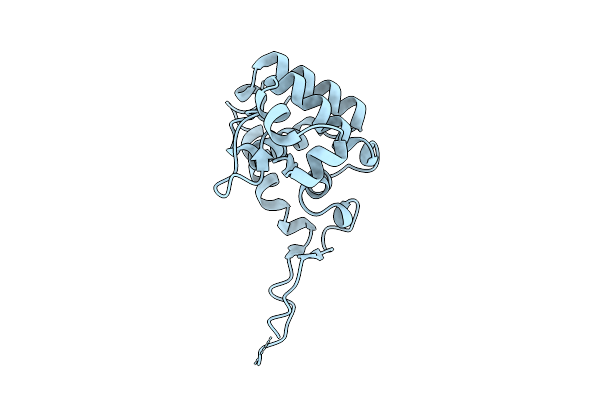
Organism: Methanocaldococcus jannaschii dsm 2661
Method: X-RAY DIFFRACTION Resolution:2.16 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
Organism: Klebsiella variicola
Method: X-RAY DIFFRACTION Resolution:1.85 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
Complex Of N-Terminal Brxc Walker B, Brxb, And N-Termianl Pglz From The Acinetobacter Brex System
Organism: Acinetobacter sp. neb 394
Method: X-RAY DIFFRACTION Resolution:2.74 Å Release Date: 2026-04-08 Classification: ANTIMICROBIAL PROTEIN Ligands: ATP, CA, CL, MG |
 |
Hybrid Model Of A Dimer Of Brxc-Brxb Fusion Complexed With Pglz From The Acinetobacter Brex System
Organism: Acinetobacter sp. neb 394
Method: ELECTRON MICROSCOPY Release Date: 2026-04-08 Classification: ANTIMICROBIAL PROTEIN |
 |
Crystal Structure Of Zika Virus Ns2B-Ns3 Protease Determined Via Sulfur Phasing
Organism: Zika virus
Method: X-RAY DIFFRACTION Resolution:1.77 Å Release Date: 2026-04-01 Classification: VIRAL PROTEIN |
 |
Crystal Structure Of Gh57 Family Amylopullulanase From Aquifex Aeolicus Mutant D352N In Complex With Maltoheptaose
Organism: Aquifex aeolicus vf5
Method: X-RAY DIFFRACTION Resolution:1.83 Å Release Date: 2026-03-25 Classification: HYDROLASE Ligands: GLC, GOL |

