Search Count: 4,825
All
Selected
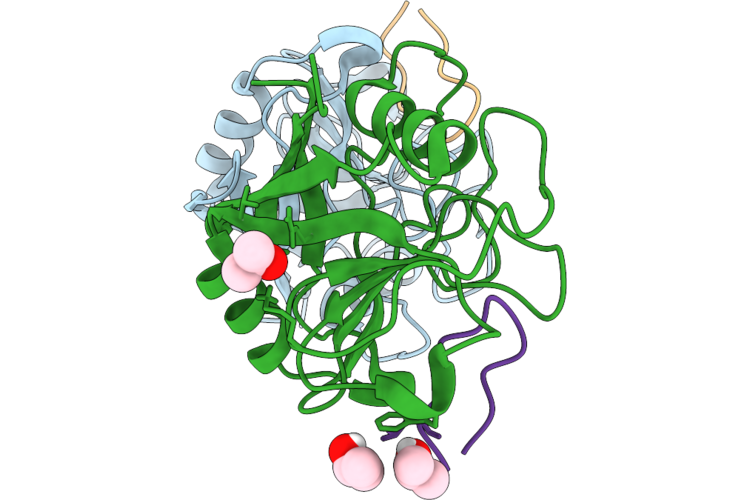 |
Organism: Entamoeba histolytica hm-1:imss, Tolypocladium inflatum
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-05-20 Classification: ISOMERASE Ligands: IPA |
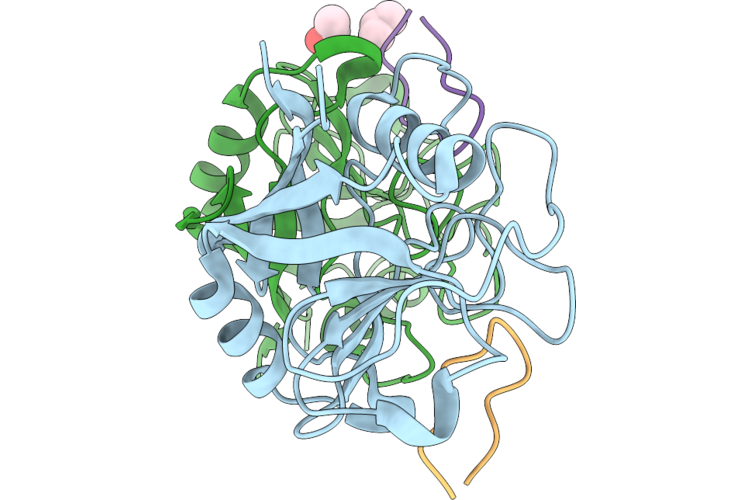 |
Organism: Entamoeba histolytica hm-1:imss, Tolypocladium inflatum
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-20 Classification: ISOMERASE Ligands: IPA |
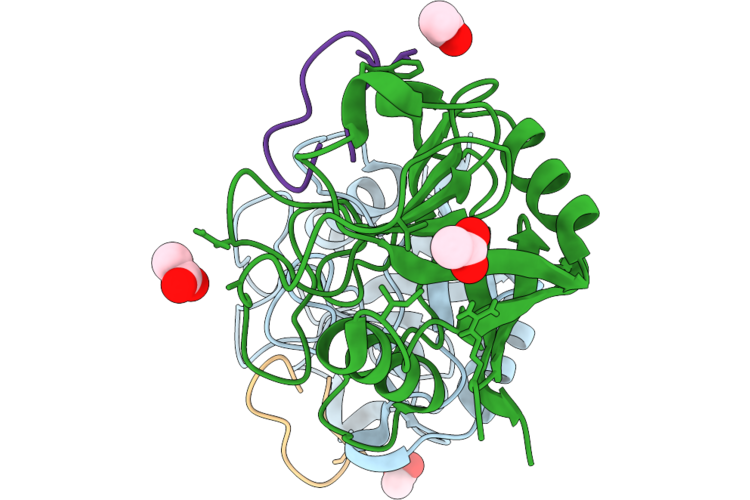 |
Organism: Entamoeba histolytica hm-1:imss, Tolypocladium inflatum
Method: X-RAY DIFFRACTION Resolution:1.60 Å Release Date: 2026-05-20 Classification: ISOMERASE Ligands: ACT |
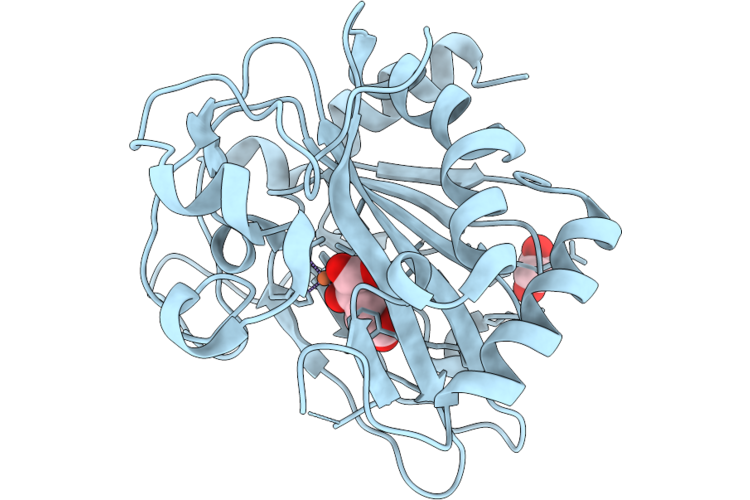 |
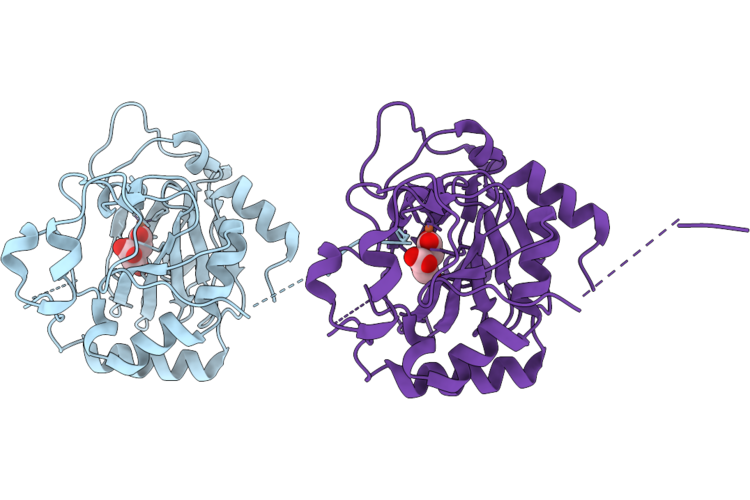
Crystal Structure Of Fe/2-Og Dependent Dioxygenase Mysh In Complex With Iron And 2-Oxoglutarate
Organism: Nostoc linckia
Method: X-RAY DIFFRACTION Resolution:1.50 Å Release Date: 2026-05-06 Classification: METAL BINDING PROTEIN Ligands: AKG, FE, EDO |
 |
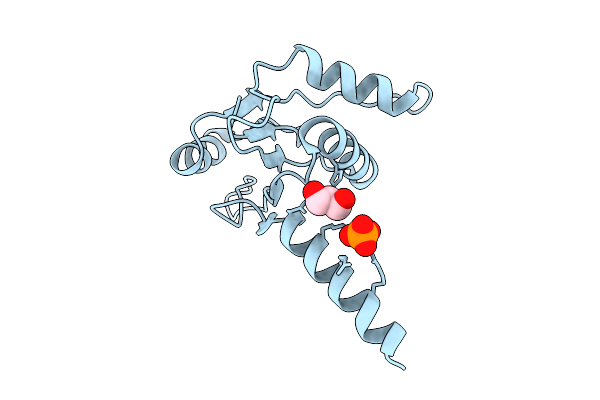
Crystal Structure Of Fe/2-Og Dependent Dioxygenase Mysh With Docked C-Terminus
Organism: Nostoc linckia
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-05-06 Classification: METAL BINDING PROTEIN Ligands: FLC, FE |
 |
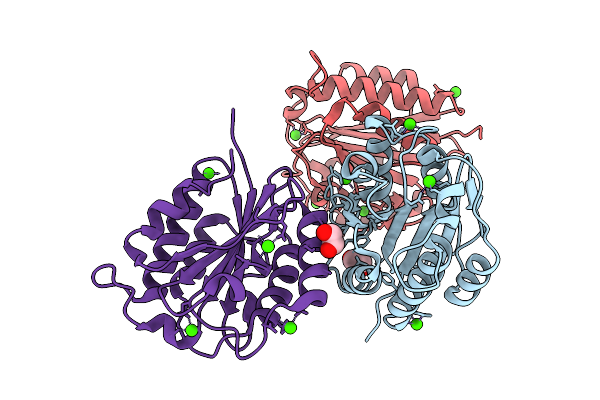
Oligomerisation Of Type Iii Crispr-Associated Csx15 To Regulate Antiviral Signalling
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.88 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN Ligands: GOL, PO4 |
 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:2.11 Å Release Date: 2026-04-01 Classification: OXYGEN BINDING Ligands: HEM, PG4, EDO, SO4, PEG, TRS, OXY, PGE |
 |
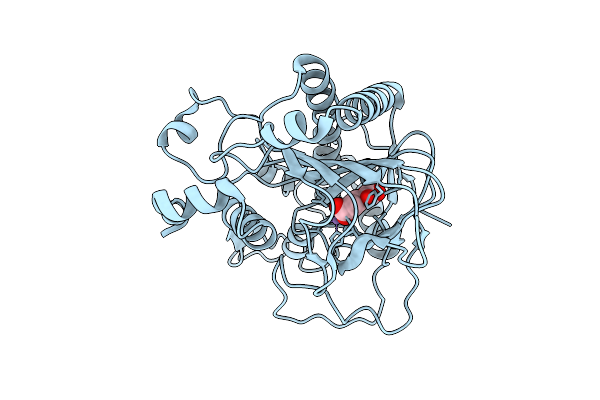
Crystal Structure Of Vwfa Domain From Lapa Adhesin Of Pseudomonas Fluorescens
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.45 Å Release Date: 2026-04-01 Classification: CELL ADHESION Ligands: CA, EDO |
 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-01 Classification: ANTITOXIN |
 |
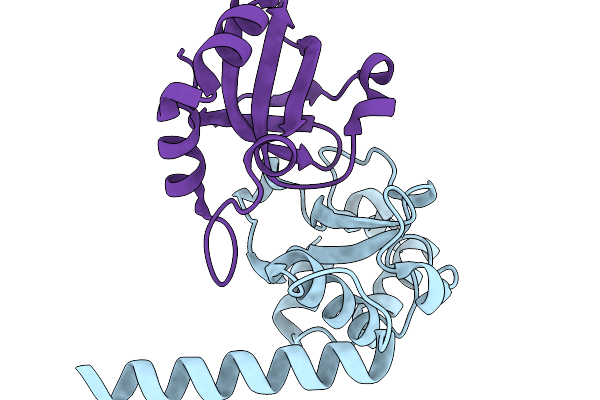
Fe(Ii)-2-Oxoglutarate-Dependent Pseudomonas Savastanoi Pv Phaseolicola 1449B In Complex With 2-Oxoglutarate
Organism: Pseudomonas savastanoi
Method: X-RAY DIFFRACTION Resolution:1.65 Å Release Date: 2026-03-18 Classification: OXIDOREDUCTASE Ligands: MN, AKG |
 |
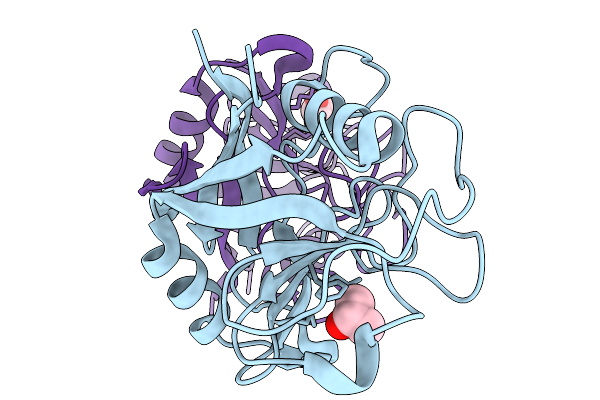
Organism: Neisseria meningitidis serogroup c (strain 8013), Veillonella
Method: X-RAY DIFFRACTION Resolution:2.08 Å Release Date: 2026-03-04 Classification: HYDROLASE |
 |
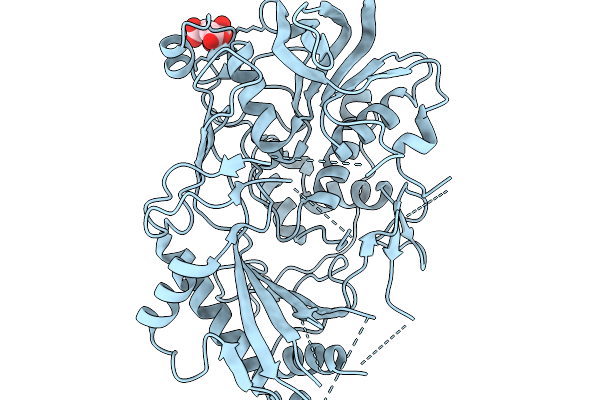
Organism: Entamoeba histolytica hm-1:imss
Method: X-RAY DIFFRACTION Resolution:1.40 Å Release Date: 2026-02-11 Classification: ISOMERASE Ligands: IPA |
 |
Crystal Structure Of Choline Dehydrogenase Beta From Pseudomonas Denitrificans
Organism: Pseudomonas sp. atcc 13867
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-02-04 Classification: OXIDOREDUCTASE Ligands: CIT |
 |
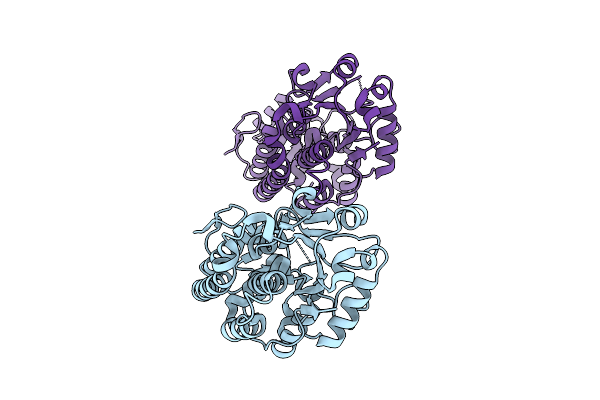
Structure And Mechanism Of The Broad Spectrum Crispr-Associated Ring Nuclease Crn4
Organism: Hyphomicrobiales bacterium
Method: X-RAY DIFFRACTION Resolution:1.10 Å Release Date: 2025-12-03 Classification: VIRAL PROTEIN |
 |
Organism: Yersinia ruckeri
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2025-11-12 Classification: METAL BINDING PROTEIN |
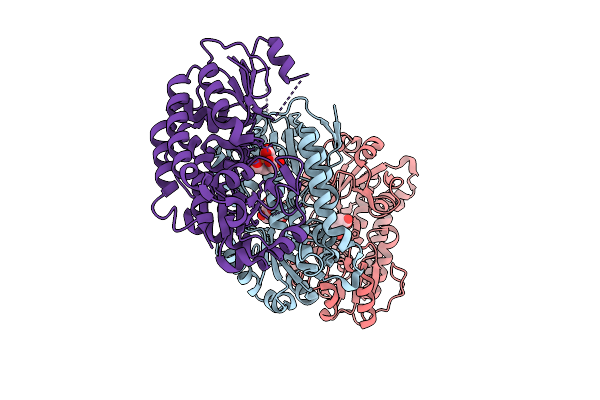 |
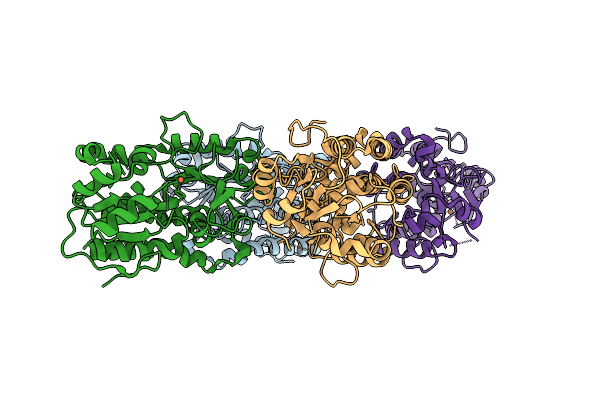
Structure Of Yiua From Yersinia Ruckeri With Iron And Nitrilotriacetic Acid
Organism: Yersinia ruckeri
Method: X-RAY DIFFRACTION Resolution:2.14 Å Release Date: 2025-11-12 Classification: METAL BINDING PROTEIN Ligands: NTA, FE |
 |
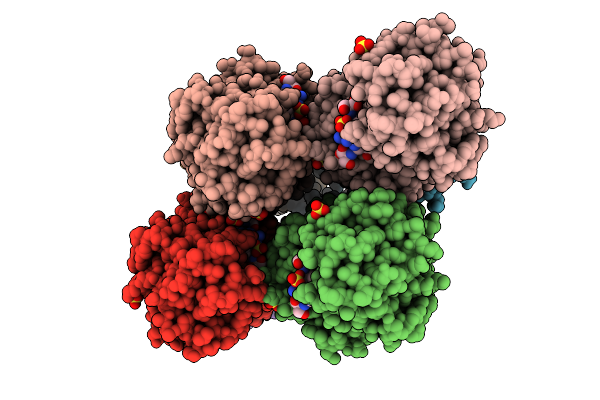
Organism: Yersinia ruckeri
Method: X-RAY DIFFRACTION Resolution:2.32 Å Release Date: 2025-11-05 Classification: METAL BINDING PROTEIN Ligands: FE |
 |
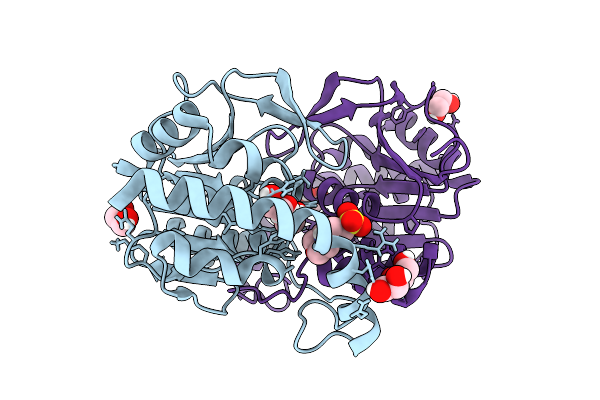
Organism: Yersinia ruckeri
Method: X-RAY DIFFRACTION Resolution:2.45 Å Release Date: 2025-11-05 Classification: METAL BINDING PROTEIN Ligands: FE, A1I15, SO4 |
 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:0.89 Å Release Date: 2025-10-29 Classification: LYASE Ligands: EDO, MES |
 |
Organism: Pseudomonas fluorescens
Method: X-RAY DIFFRACTION Resolution:1.20 Å Release Date: 2025-10-29 Classification: LYASE Ligands: MG, CL |

