Search Count: 62,096
All
Selected
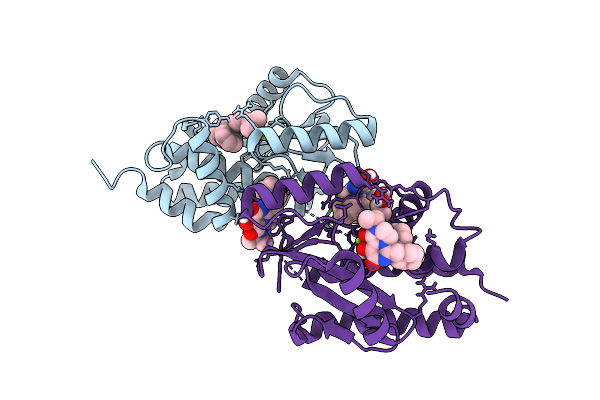 |
Three Viral Endonucleases Bound To The Same Inhibitor (9-Hydroxy-3,4-Dihydro-2H-Pyrazino[1,2-C]Pyrimidine-1,8-Dione Derivative). (1A) Influenza Virus A/Ph1N1 Pa Subunit Endonuclease With Magnesium Ions.
Organism: Influenza a virus
Method: X-RAY DIFFRACTION Resolution:1.86 Å Release Date: 2026-04-15 Classification: RNA BINDING PROTEIN Ligands: MG, A1J4F, GOL |
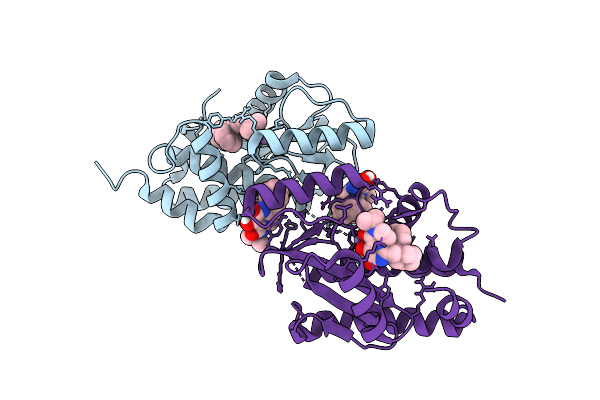 |
Three Viral Endonucleases Bound To The Same Inhibitor (9-Hydroxy-3,4-Dihydro-2H-Pyrazino[1,2-C]Pyrimidine-1,8-Dione Derivative). (1B) Influenza Virus A/Ph1N1 Polymerase Pa Subunit Endonuclease With Manganese Ions.
Organism: Influenza a virus
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-15 Classification: RNA BINDING PROTEIN Ligands: MN, A1J4F |
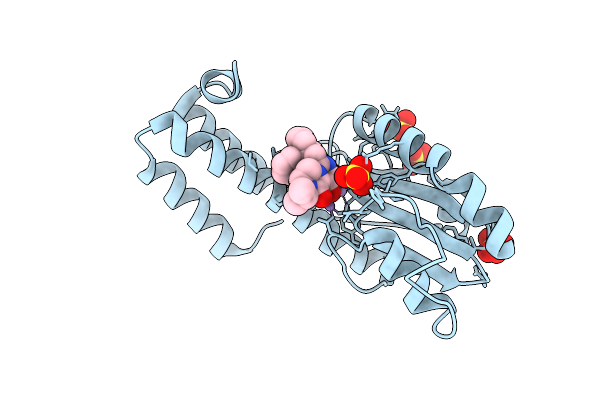 |
Three Viral Endonucleases Bound To The Same Inhibitor (9-Hydroxy-3,4-Dihydro-2H-Pyrazino[1,2-C]Pyrimidine-1,8-Dione Derivative). (2) La Crosse Virus L Protein Endonuclease With Manganese Ions.
Organism: La crosse virus
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-15 Classification: RNA BINDING PROTEIN Ligands: MN, A1J4F, SO4 |
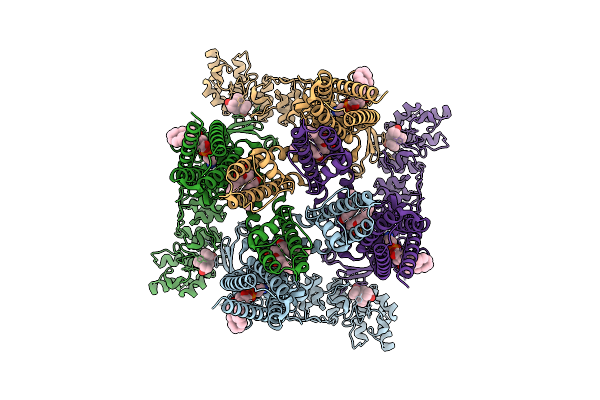 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: MEMBRANE PROTEIN Ligands: PEX, A1BMB |
 |
Crystal Structure Of Gp232 Tail Fiber Recognition Domain From Mycobacteriophage Bxz-1
Organism: Bixzunavirus
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-15 Classification: VIRAL PROTEIN |
 |
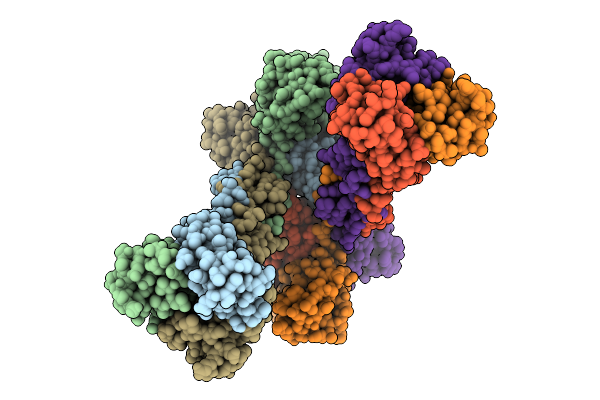
Structure Of The Measles Virus Fusion Glycoprotein Ectodomain In Complex With Neutralizing Antibody Y10F
Organism: Measles morbillivirus, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
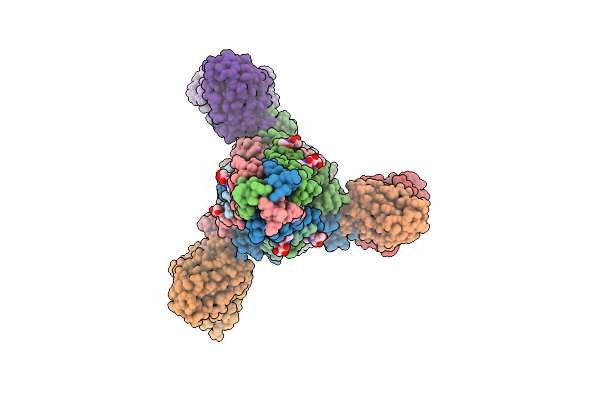
Structure Of The Measles Virus Fusion Glycoprotein Ectodomain In Complex With Two Neutralizing Antibodies Mab77 And Y10F
Organism: Measles morbillivirus, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
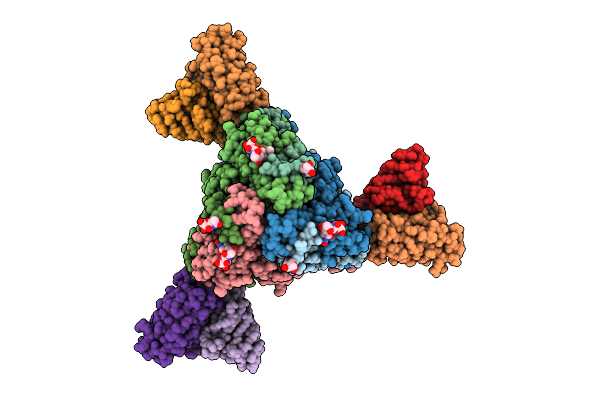
Structure Of The Measles Virus Fusion Glycoprotein Ectodomain In Postfusion Conformation Bound To Neutralizing Antibody H8
Organism: Measles morbillivirus, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
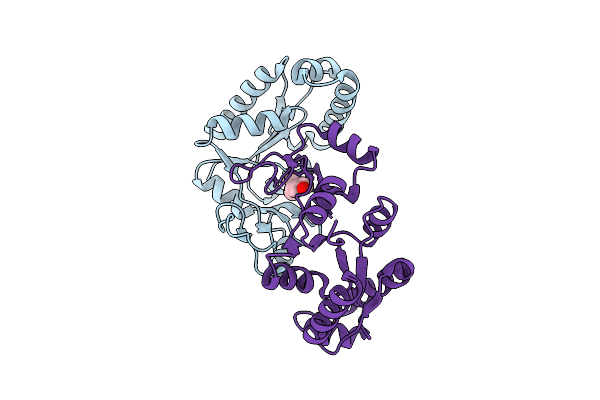
Structure Of The Measles Virus Fusion Glycoprotein Ectodomain In Prefusion Conformation Bound To Neutralizing Antibody H8
Organism: Measles morbillivirus, Mus musculus
Method: ELECTRON MICROSCOPY Release Date: 2026-04-15 Classification: VIRAL PROTEIN/IMMUNE SYSTEM Ligands: NAG |
 |
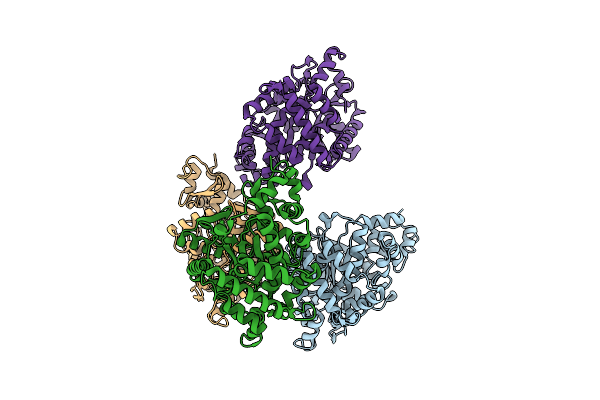
Organism: Rattus norvegicus
Method: X-RAY DIFFRACTION Resolution:2.44 Å Release Date: 2026-04-15 Classification: STRUCTURAL PROTEIN Ligands: A1EOW |
 |
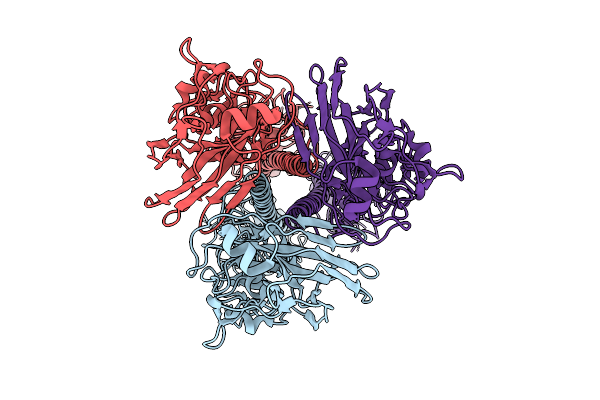
Organism: Solanum tuberosum
Method: X-RAY DIFFRACTION Resolution:2.53 Å Release Date: 2026-04-15 Classification: PLANT PROTEIN |
 |
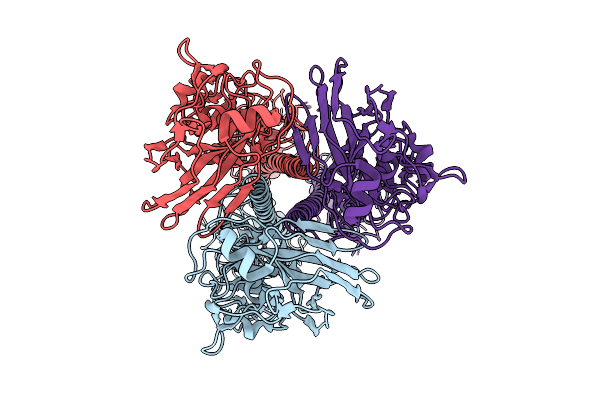
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.39 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.16 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
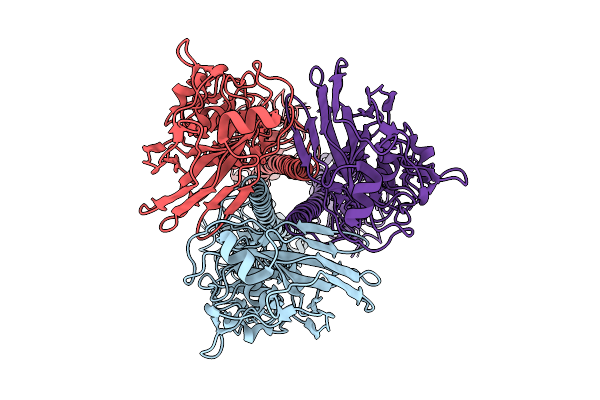
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.67 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
 |
Influenza A Virus Hemagglutinin (A/Darwin/6/2021 H3N2) (C3 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 1X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.47 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
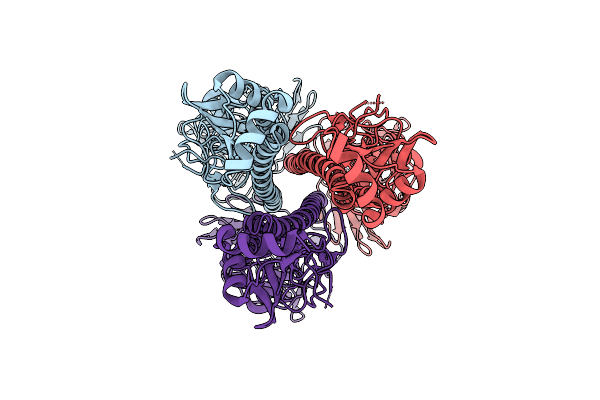 |
Influenza A Virus Hemagglutinin (A/California/04/2009 H1N1), E47K Ha2 Stabilizing Mutation (C1 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.94 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
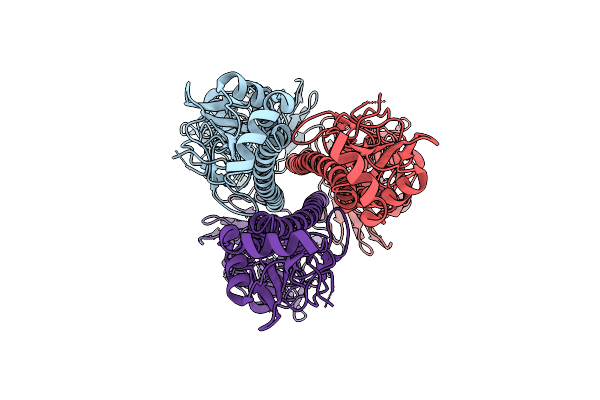 |
Influenza A Virus Hemagglutinin (A/California/04/2009 H1N1), E47K Ha2 Stabilizing Mutation (C3 Symmetry) Determined Using The Spt Labtech Chameleon In The Presence Of 0.25X Surfact
Organism: Influenza a virus
Method: ELECTRON MICROSCOPY Resolution:2.64 Å Release Date: 2026-04-08 Classification: VIRAL PROTEIN |
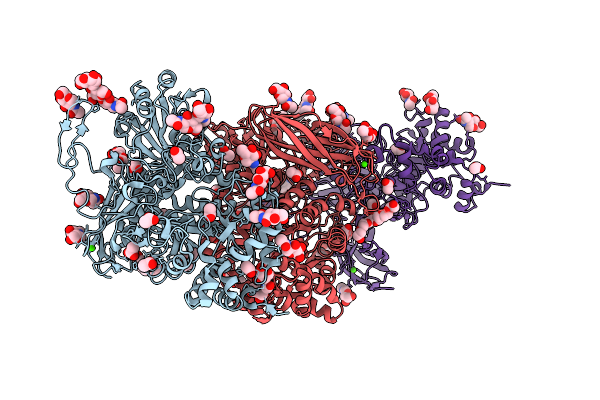 |
Crystal Structure Of Isoprimeverose-Producing Enzyme From Phaeoacremonium Minimum
Organism: Rattus rattus, Phaeoacremonium minimum
Method: X-RAY DIFFRACTION Resolution:2.80 Å Release Date: 2026-04-08 Classification: HYDROLASE Ligands: NAG, ACT, PG4, CA, PGE, P6G |
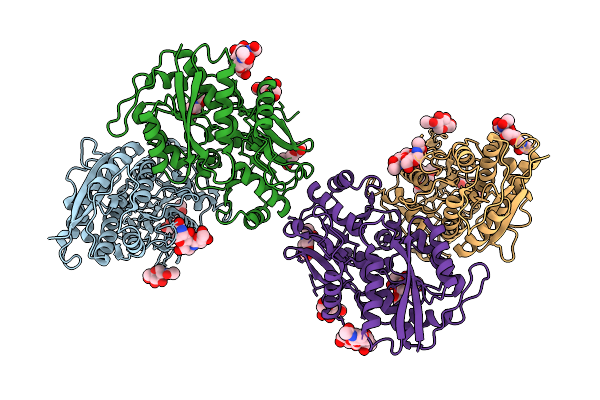 |
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.40 Å Release Date: 2026-04-08 Classification: STRUCTURAL PROTEIN Ligands: NAG |
 |
Cryoem Structure Of Gluk2 Lbd-Tmd Bound To Glutamate In The Shallow Desensitized State
Organism: Rattus norvegicus
Method: ELECTRON MICROSCOPY Resolution:3.86 Å Release Date: 2026-04-08 Classification: STRUCTURAL PROTEIN Ligands: NAG, GLU |

