Search Count: 6,132
All
Selected
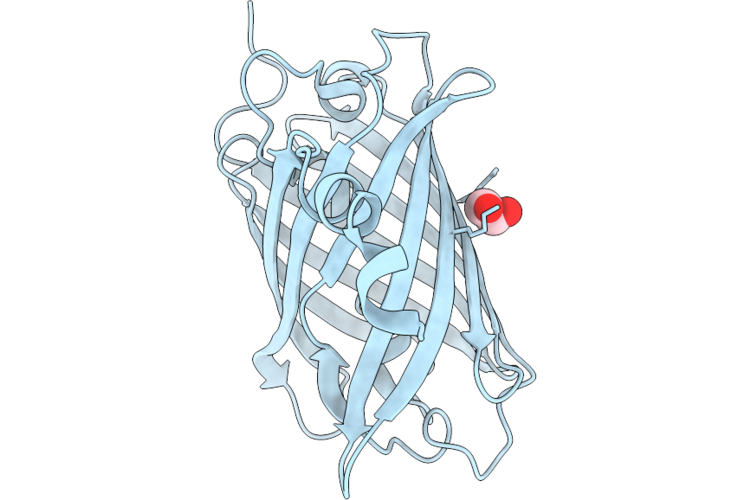 |
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:1.90 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN Ligands: EDO |
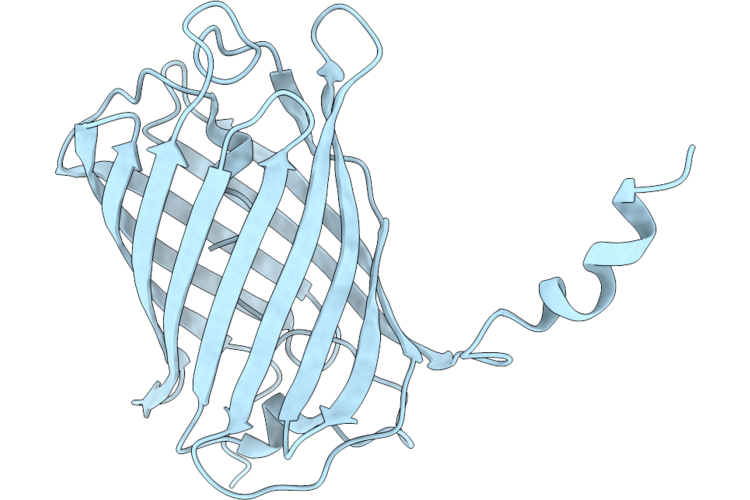 |
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:1.70 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN |
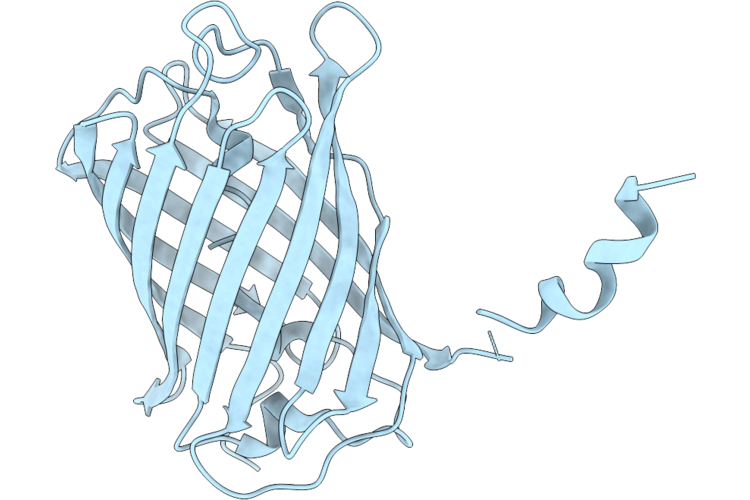 |
Crystal Structure Of Lssmorange Under Room Temperature At Ph 8.0, 250Ps After Pump Laser, Refined Against 8.33 Times Extrapolated Structure Factor
Organism: Discosoma sp.
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-22 Classification: FLUORESCENT PROTEIN |
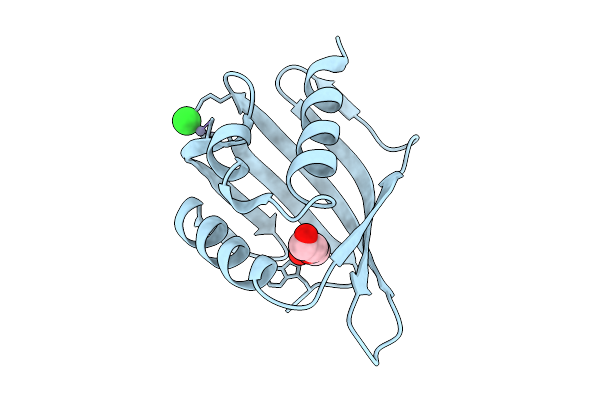 |
Crystal Structure Of A Snoal-Like Domain-Containing Protein From Mycobacterium Ulcerans (Zinc Bound)
Organism: Mycobacterium ulcerans agy99
Method: X-RAY DIFFRACTION Resolution:1.31 Å Release Date: 2026-04-15 Classification: ISOMERASE Ligands: PEG, ZN, CL |
 |
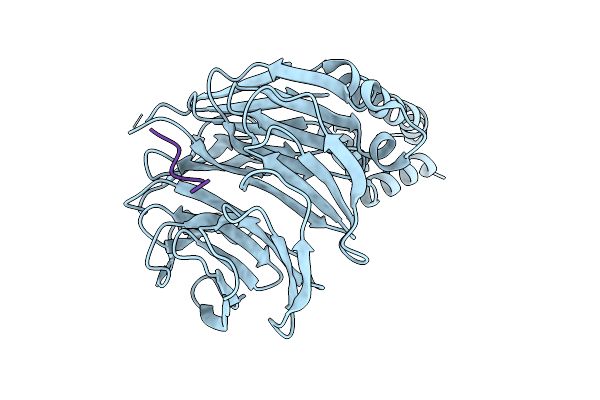
Crystal Structure Of Beta-Trcp Bound By Monophosphorylated Nrf2 Degron Peptide.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.80 Å Release Date: 2026-04-08 Classification: LIGASE |
 |
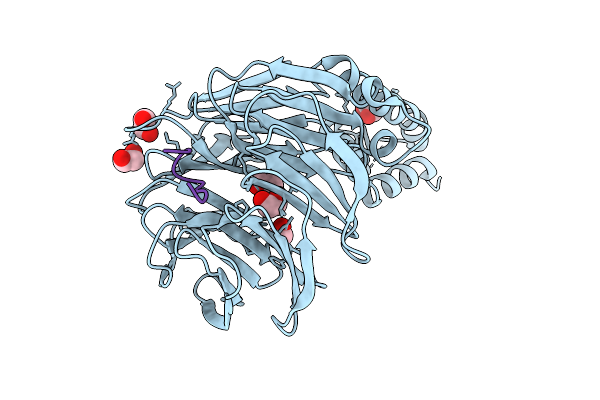
Crystal Structure Of Beta-Trcp Bound By Diphosphorylated Nrf2 Degron Peptide.
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.15 Å Release Date: 2026-04-08 Classification: LIGASE |
 |
Crystal Structure Of Beta-Trcp Bound By Diphosphorylated I-Kappa-B-Alpha Degron Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.16 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: EDO |
 |
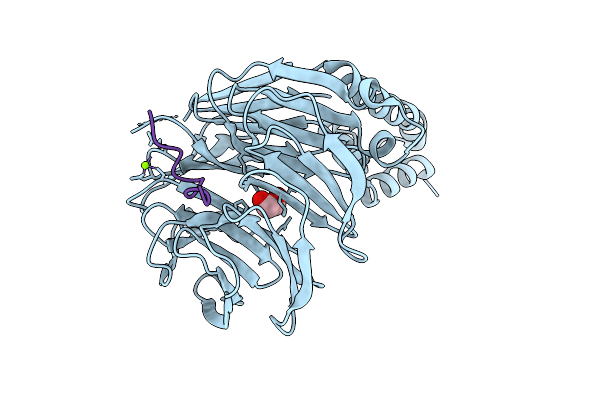
Crystal Structure Of Beta-Trcp Bound By Diphosphorylated Claspin Degron Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.35 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: EDO, MG |
 |
Crystal Structure Of Beta-Trcp Bound By Diphosphorylated Pdcd4 Degron Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.22 Å Release Date: 2026-04-08 Classification: TRANSFERASE Ligands: MG, EDO |
 |
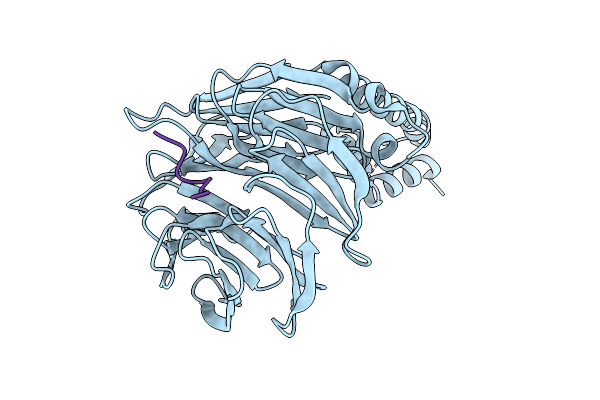
Crystal Structure Of Beta-Trcp Bound By Monophosphorylated Atf4 Degron Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:2.00 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
 |
Crystal Structure Of Beta-Trcp Bound By Monophosphorylated Wee1 Degron Peptide
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.68 Å Release Date: 2026-04-08 Classification: TRANSFERASE |
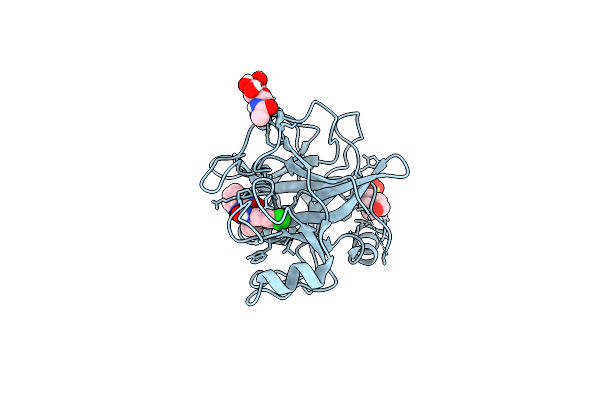 |
N-Alkyl &Amp; N-Aryl Aminopyrazole Spirocarbamates: A Two-Pronged Lead Optimization Strategy To Identify Orally Bioavailable Plasma Kallikrein Inhibitors Complex With Compound 15 ((3'R)-1'-(5-Amino-1-Phenyl-1H-Pyrazole-4-Carbonyl)-6-Chloro-5-Fluorospiro[[3,1]Benzoxazine-4,3'-Piperidin]-2(1H)-One)
Organism: Homo sapiens
Method: X-RAY DIFFRACTION Resolution:1.66 Å Release Date: 2026-04-01 Classification: HYDROLASE/INHIBITOR Ligands: MES, A1C6X, NAG |
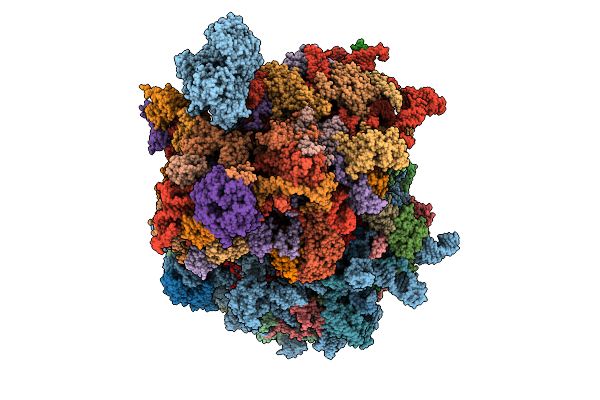 |
Organism: Homo sapiens
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: TRANSLATION Ligands: MG, ZN |
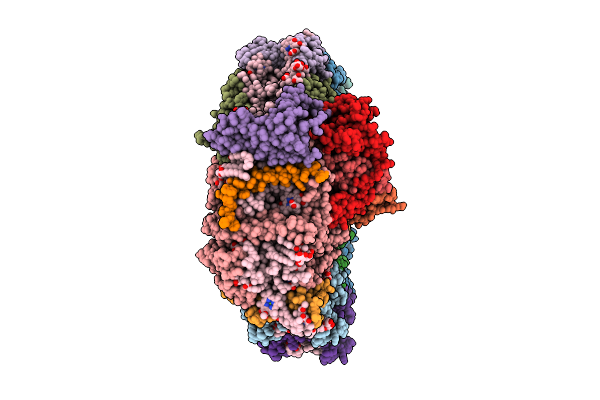 |
Structure Of Photosystem I From Chlamydomonas Reinhardtii At 1.83 A Resolution
Organism: Chlamydomonas reinhardtii
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: CLA, CHL, BCR, LUT, LMG, LMU, XAT, LHG, NEX, CL0, PQN, SF4, DGD |
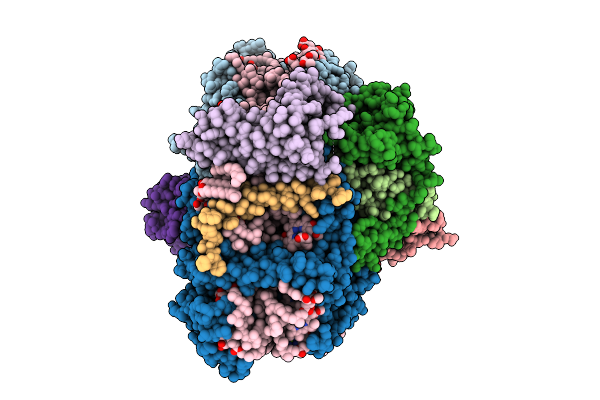 |
Structure Of Cytochrome C6 Bound Photosystem I From Chlamydomonas Reinhardtii At 2.07 A Resolution
Organism: Chlamydomonas reinhardtii
Method: ELECTRON MICROSCOPY Release Date: 2026-04-01 Classification: PHOTOSYNTHESIS Ligands: CLA, PQN, LHG, BCR, SF4, LMU, LMG, DGD, LUT, HEC |
 |
1H, 13C, And 15N Resonance Assignments And Solution Structure Of The Cid Domain Of Scaf8 (Rbm16)
Organism: Homo sapiens
Method: SOLUTION NMR Release Date: 2026-04-01 Classification: STRUCTURAL GENOMICS |
 |
Cryo-Em Structure Of The Vo Domain Of V/A-Atpase In Liposomes Under No Pmf Condition,State2
Organism: Thermus thermophilus hb8
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MOTOR PROTEIN |
 |
Cryo-Em Structure Of The Vo Domain Of V/A-Atpase In Liposomes Under No Pmf Condition,State3
Organism: Thermus thermophilus hb8
Method: ELECTRON MICROSCOPY Release Date: 2026-03-25 Classification: MOTOR PROTEIN |
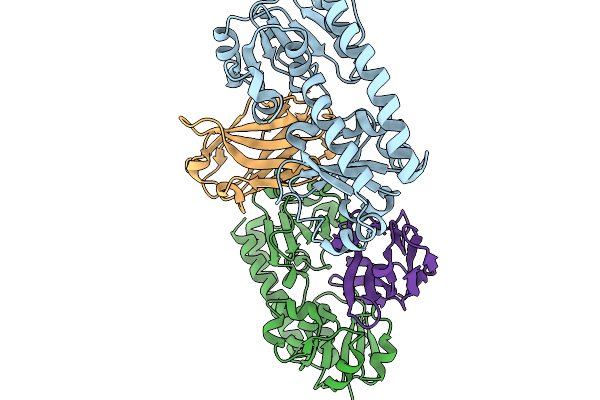 |
Crystal Structure Of Ftsb From Streptococcus Pyogenes In Complex With Nb1 Nanobody
Organism: Streptococcus pyogenes, Vicugna pacos
Method: X-RAY DIFFRACTION Resolution:2.55 Å Release Date: 2026-03-25 Classification: IMMUNE SYSTEM |
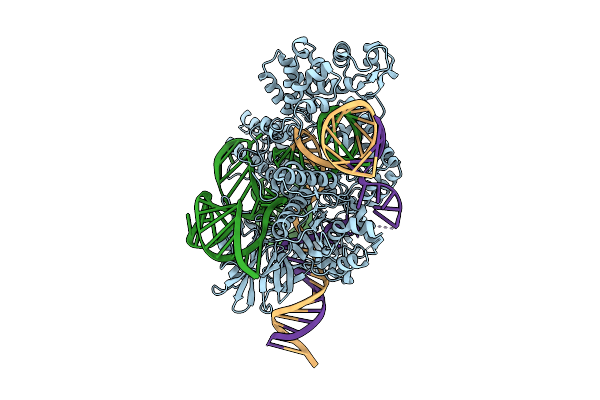 |
Cryo-Em Structure Of Wild-Type Sacas9-Guide Rna-Mismatched Target Dna Complex
Organism: Staphylococcus aureus
Method: ELECTRON MICROSCOPY Release Date: 2026-03-18 Classification: DNA BINDING PROTEIN |

